Assembly standard 25
< Back to Catalog
< RFC25 BioBrick collection
The 2007 Freiburg iGEM team proposed an extension of the original BioBrick assembly standard that enables in-frame assembly of protein domains. Two restriction sites are added within the original BioBrick prefix and suffix sites. These additional sites provide compatible ends, can be employed using the same cloning strategy as for the standard restriction sites, and code for amino acids suitable for linkers:
Prefix and suffix
Prefix Suffix
5' GAATTC GCGGCCGC T TCTAGA TG GCCGGC [part] ACCGGT TAAT ACTAGT A GCGGCCG CTGCAG 3'
EcoRI NotI XbaI Met NgoMIV AgeI * SpeI NotI PstI
(=NgoMI)
"N-part" prefix
5' GAATTC GCGGCCGC T TCTAG [ATG.part] EcoRI NotI XbaI
Scar
Assembling two parts based on the new Freiburg standard restriction sites leaves the following scar:
5' [part A] ACCGGC [part B] 3'
T G
See [http://openwetware.org/wiki/The_BioBricks_Foundation:Standards/Technical/Formats The BioBricks Foundation wiki] for a discussion and comparison of different technical standards.
| Plasmid backbones (?) | Protein domains (?) | Protein coding sequences (?) |
| Help: Want to know more about Assembly standard 25, often called the Freiburg standard? See the help pages for more information. |
Plasmid backbones
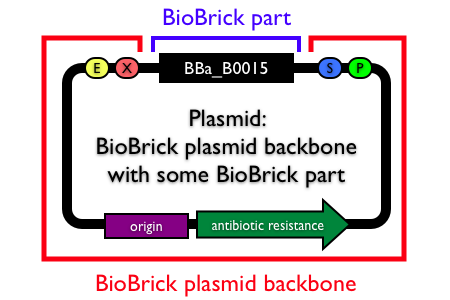
Plasmids are circular, double-stranded DNA molecules typically containing a few thousand base pairs that replicate within the cell independently of the chromosomal DNA. Plasmid DNA is easily purified from cells, manipulated using common lab techniques and incorporated into cells. Most BioBrick parts in the Registry are maintained and propagated on plasmids. Thus, construction of BioBrick parts, devices and systems usually requires working with plasmids.
Note: In the Registry, plasmids are made up of two distinct components:
- the BioBrick part, device or system that is located in the BioBrick cloning site, between (and excluding) the BioBrick prefix and suffix.
- the plasmid backbone which propagates the BioBrick part. The plasmid backbone is defined as the sequence beginning with the BioBrick suffix, including the replication origin and antibiotic resistance marker, and ending with the BioBrick prefix. [Note that the plasmid backbone itself can be composed of BioBrick parts.]
Many BioBrick parts in the Registry are maintained on more than one plasmid backbone!
| Raik Gruenberg developed the protein fusion vectors BBa_J18901, BBa_J18902 and BBa_J18903 that adhere to the Freiburg standard. |
> RFC25 BioBricks backbones collection
Protein domains
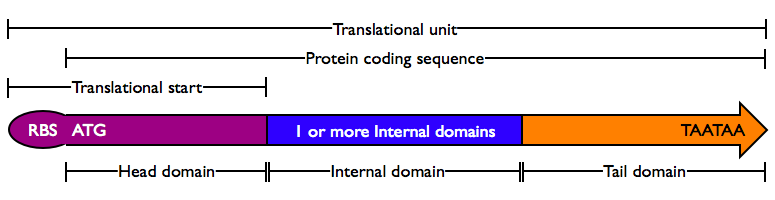
Protein domains encode portions of proteins and can be assembled together to form translational units, a genetic part spanning from translational initiation (the RBS) to translational termination (the stop codon).
There are several different types of protein domains.
- Head Domain: The Head domain consists of the start codon followed immediately by zero or more triplets specifiying an N-terminal tag, such as a protein export tag or lipoprotein binding tag. Head domains should begin with an
ATGstart codon and include codons 2 and 3 of the protein at a minimum. Examples of head domains includeATGstart codonATGstart codon and codons 2-3ATGstart codon and signal sequenceATGstart codon and affinity tag
- Internal Domains: Protein domains consist of a series of codon triplets coding for an amino acid sequence without a start codon or stop codon. Multiple Internal Domains can be fused. Examples of internal domains include
- DNA binding domains
- Dimerization domains
- Kinase domains
- Special Internal Domains: Short Domains with specific function may be separately categorized, but obey the same composition rules as normal internal domains. Examples of special internal domains include
- Linkers
- Cleavage sites
- Inteins
- Tail Domain: The C-terminus of a coding region consists of zero or more triplet codons, followed by a pair of TAA stop codons. In the simplest case, the stop codons terminate the protein with an Stop. More complex Tails may include degradation tags appropriate to the organism (i.e., with different degradation rates). Examples of Tail domain include
TAATAAstop codons- A degradation tag followed by
TAATAAstop codons - An affinity tag followed by
TAATAAstop codon
Unfortunately, the original BioBrick assembly standard, Assembly standard 10, does not support in-frame assembly of protein domains. (Assembly standard 10 creates an 8 bp scar between adjacent parts.) Therefore, it is recommended that you use an alternate approach to assemble protein domains together to make a translational unit. There are several possible approaches to assembling protein domains including direct synthesis (preferred because it creates no scars) as well as various assembly standards. Regardless of which standard you choose, we suggest that the resulting protein coding sequence or translational unit comply with the original BioBrick assembly standard so that your parts can be assembled with most of the parts in the Registry.
Protein coding sequences should be as follows
GAATTC GCGGCCGC T TCTAG [ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG
Note: Although most RBSs are currently specified as separate parts in the Registry, we are now moving to a new design in which the RBS and Head domain are combined into a single part termed a Translational start. The new design has the advantage of encapsulating both ribosome binding and translational initiation within a single part. Our working hypothesis is that the new design will reduce the likelihood of unexpected functional composition problems between the RBS and coding sequence.
> RFC25 protein domain BioBricks collection

|
The 2008 Albert-Ludwigs-Universitat Freiburg iGEM team designed and constructed these protein tags and modifiers that adhere to the Freiburg assembly standard. |
Protein coding sequences
Protein coding sequences are DNA sequences that are transcribed into mRNA and in which the corresponding mRNA molecules are translated into a polypeptide chain. Every three nucleotides, termed a codon, in a protein coding sequence encodes 1 amino acid in the polypeptide chain. In some cases, different chassis may either map a given codon to a different sequence or may use different codons more or less frequently. Therefore some protein coding sequences may be optimized for use in a particular chassis.
In the Registry, protein coding sequences begin with a start codon (usually ATG) and end with a stop codon (usually with a double stop codon TAA TAA). Protein coding sequences are often abbreviated with the acronym CDS.
Although protein coding sequences are often considered to be basic parts, in fact proteins coding sequences can themselves be composed of one or more regions, called protein domains. Thus, a protein coding sequence could either be entered as a basic part or as a composite part of two or more protein domains.
- The N-terminal domain of a protein coding sequence is special in a number of ways. First, it always contains a start codon, spaced at an appropriate distance from a ribosomal binding site. Second, many coding regions have special features at the N terminus, such as protein export tags and lipoprotein cleavage and attachment tags. These occur at the beginning of a coding region, and therefore are termed Head domains.
- A protein domain is a sequence of amino acids which fold relatively independently and which are evolutionarily shuffled as a unit among different protein coding regions. The DNA sequence of such domains must maintain in-frame translation, and thus is a multiple of three bases. Since these protein domains are within a protein coding sequence, they are called Internal domains. Certain Internal domains have particular functions in protein cleavage or splicing and are termed Special Internal domains.
- Similarly, the C-terminal domain of a protein is special, containing at least a stop codon. Other special features, such as degradation tags, are also required to be at the extreme C-terminus. Again, these domains cannot function when internal to a coding region, and are termed Tail domains.
For more details on protein domains including how to assemble protein domains into protein coding sequences, please see Protein domains.
Protein coding sequences should be as follows
GAATTC GCGGCCGC T TCTAG [ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG
> RFC25 protein coding sequence BioBricks collection

|
The 2008 Albert-Ludwigs-Universitat Freiburg iGEM team designed and constructed these protein coding sequences that adhere to the Freiburg assembly standard. |
References
- [http://hdl.handle.net/1721.1/45140 BBF RFC 25: Fusion Protein (Freiburg) Biobrick assembly standard] by Kristian M. Müller, Katja M. Arndt, the 2007 Freiburg iGEM team, and Raik Grünberg
<biblio>
- Nishikubo pmid=16140408
- Pfleger pmid=16247774
- Varshavsky pmid=8901547
</biblio>
- [http://www.neb.com/nebecomm/products_Intl/productR0564.asp NgoMIV at NEB]