Difference between revisions of "Plasmid backbones/Other standards"
(Replacing Berkeley standard template redirecting link with direct link) |
(Replacing redirecting link with direct link) |
||
| Line 27: | Line 27: | ||
==Lim assembly standard plasmid backbones== | ==Lim assembly standard plasmid backbones== | ||
| − | {{: | + | {{:Assembly standard 28/Overview}} |
While any vector can be adapted to adhere to the Lim standard, most of the available vectors from the Registry that adhere to the Lim standard are derived from the yeast pRS__ series of vectors. The Registry has several types of acceptor vectors built in the pRS315 or 305 backbone. When necessary, markers can be exchanged by one-piece subclones of the completed cassettes into alternative pRS vectors, using the Kpn1/PspOMI and SacI sites. | While any vector can be adapted to adhere to the Lim standard, most of the available vectors from the Registry that adhere to the Lim standard are derived from the yeast pRS__ series of vectors. The Registry has several types of acceptor vectors built in the pRS315 or 305 backbone. When necessary, markers can be exchanged by one-piece subclones of the completed cassettes into alternative pRS vectors, using the Kpn1/PspOMI and SacI sites. | ||
Latest revision as of 21:29, 28 June 2017
| Part assembly | System operation | Protein expression | Assembly of protein fusions | Part measurement | Screening of part libraries | Building BioBrick vectors | DNA synthesis | Other standards | Archive |
| Or get some help on plasmid backbones. |
Silver assembly standard plasmid backbones
Please see plasmid backbones for protein fusions
Freiburg assembly standard plasmid backbones
Please see plasmid backbones for protein fusions
Berkeley assembly standard plasmid backbones
Researchers at UC Berkeley have developed the BglBrick assembly standard, or Assembly standard 21, based on idempotent assembly with BamHI and BglII restriction enzymes. In a nutshell, most parts look like this:
Prefix Suffix 5' GAATTC atg AGATCT ...part... GGATCC taa CTCGAG 3' EcoRI BglII BamHI * XhoI
Fusing two parts leaves the following scar:
5' [part A] GGATCT [part B] 3'
G S
Note, however, that Assembly standard 20 is intended as a minimal physical assembly standard, and only those features needed for interconversion of BglBrick assembly standard plasmids are formally defined. Therefore, atg and taa spacers are not core definitions of the standard.
See [http://openwetware.org/wiki/The_BioBricks_Foundation:Standards/Technical/Formats The BioBricks Foundation wiki] for a discussion and comparison of different technical standards.
There are no parts for this table
Lim assembly standard plasmid backbones
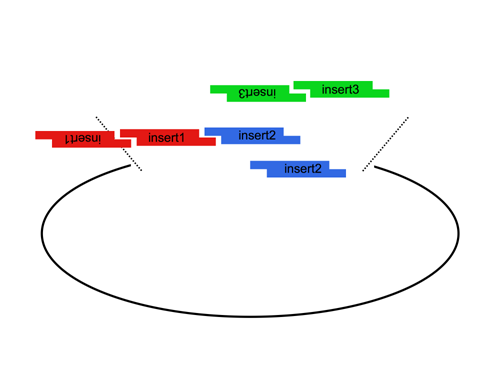
| The Lim standard is based on a multi-part/combinatorial cloning technique that is particularly well suited to shuffling protein domains. The key to this approach is the Type IIS restriction enzyme, AarI, a rare (7-cutter) that cuts 4bp offset from its binding site. Thus, AarI can generate four base overhangs of any sequence. | 
|
|
Since the user can specify the overhangs, this method can be used to "stitch-together" fragments without a scar, which is sometimes necessary to preserve protein function. More importantly, these overhangs can be non-palindromic thereby avoiding a key problem of trying to do multipart ligations using standard restriction enzymes: the self ligation of a part (which blocks proper assembly of parts). |
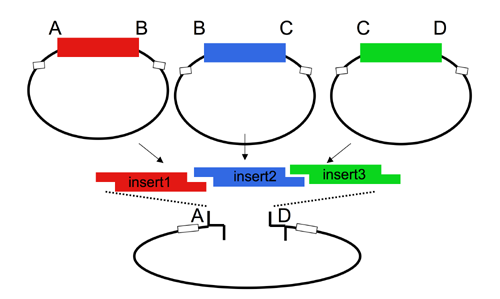
| The Lim standard enables high efficiency ligations of up to 4 parts simultaneously (vector plus 3 inserts). While parts can be made with any 4 base overhangs, we chose a standard set, termed A, B, C, and D to facilitate exchange of parts between researchers.
Note that most parts that adhere to the Lim standard are primarily intended for use when working with yeast. |
|
While any vector can be adapted to adhere to the Lim standard, most of the available vectors from the Registry that adhere to the Lim standard are derived from the yeast pRS__ series of vectors. The Registry has several types of acceptor vectors built in the pRS315 or 305 backbone. When necessary, markers can be exchanged by one-piece subclones of the completed cassettes into alternative pRS vectors, using the Kpn1/PspOMI and SacI sites.
There are no parts for this table

|

|
Sergio Peisajovich and Andrew Horowitz, from Wendell Lim's lab, developed several of the yeast plasmid backbones as an instructor of the 2008 UCSF iGEM team. |
References
<biblio>
- Sikorski pmid=2659436
</biblio>











