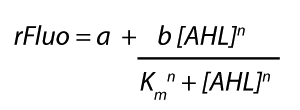Difference between revisions of "Part:BBa R0079:Experience"
(→Results) |
|||
| (30 intermediate revisions by 4 users not shown) | |||
| Line 27: | Line 27: | ||
|width='60%' valign='top'| | |width='60%' valign='top'| | ||
| − | = Characterization of | + | = Characterization of Crosstalk= |
| − | + | = Background information = | |
| − | + | We used an ''E. coli'' TOP10 strain transformed with two medium copy plasmids (about 15 to 20 copies per plasmid and cell). The first plasmid contained the commonly used p15A origin of replication, a kanamycin resistance gene, and promoter [https://parts.igem.org/Part:BBa_R0079 pLas (BBa_R0079)] followed by [https://parts.igem.org/Part:BBa_B0034 RBS (BBa_B0034)] and superfolder green fluorescent protein (sfGFP). In general, for spacer and terminator sequences the parts [https://parts.igem.org/Part:BBa_B0040 BBa_B0040] and [https://parts.igem.org/Part:BBa_B0015 BBa_B0015] were used, respectively. The second plasmid contained the pBR322 origin (pMB1), which yields a stable two-plasmid system together with p15A, an ampicillin resistance gene, and a strong promoter [https://parts.igem.org/Part:BBa_J23100 (BBa_J23100)] chosen from the [https://parts.igem.org/Promoters/Catalog/Anderson Anderson promoter collection] followed by one of the three different regulators ([https://parts.igem.org/Part:BBa_C0062 LuxR], [https://parts.igem.org/Part:BBa_C0179 LasR], and [https://parts.igem.org/Part:BBa_C0171 RhlR]) used in the experiments in order to quantify crosstalk with [https://parts.igem.org/Part:BBa_R0079 pLas]. The detailed regulator construct design and full sequences (piG0040, piG0041, piG0042, piG0050) are [http://2014.igem.org/Team:ETH_Zurich/lab/sequences available here]. In the following, we describe the experimental set-up and all the different levels of crosstalk we have assessed. | |
| − | == | + | = Experimental Set-Up = |
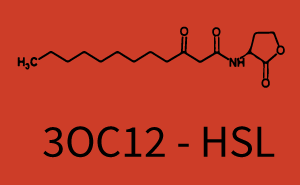
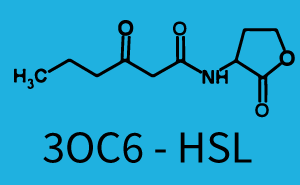
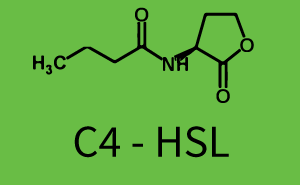
| + | The above described ''E. coli'' TOP10 strains were grown overnight in Lysogeny Broth (LB) containing kanamycin (50 μg/mL) and ampicillin (200 μg/mL) to an OD<sub>600</sub> of about 1.5 (37 °C, 220 rpm). As a reference, a preculture of the same strain lacking the sfGFP gene was included for each assay. The cultures were then diluted 1:40 in fresh LB containing the appropriate antibiotics and measured in triplicates in microtiter plate format on 96-well plates (200 μL culture volume) for 10 h at 37 °C with a Tecan infinite M200 PRO plate reader (optical density measured at 600 nm; fluorescence with an excitation wavelength of 488 nm and an emission wavelength of 530 nm). After 200 min we added the following concentrations of inducers ([[3OC6HSL|3OC6-HSL]], [[AHL|3OC12-HSL]], and [[AHL|C4-HSL]]): 10<sup>-4</sup> nM and 10<sup>4</sup> nM (from 100 mM stocks in DMSO). Attention: All the dilutions of [[AHL|3OC12-HSL]] should be made in DMSO in order to avoid precipitation. In addition, in one triplicate only H<sub>2</sub>O was added as a control. From the the obtained kinetic data, we calculated mean values and plotted the dose-response-curves for 200 min past induction. | ||
== First-order crosstalk == | == First-order crosstalk == | ||
| + | |||
| + | In the first order crosstalk section we describe crosstalk of [https://parts.igem.org/Part:BBa_R0079 pLas] due to [https://parts.igem.org/Part:BBa_C0179 LasR] binding to inducers different from [[AHL|3OC12-HSL]] or [https://parts.igem.org/Part:BBa_R0079 pLas] itself binding to a regulator-inducer pair different from [https://parts.igem.org/Part:BBa_C0179 LasR]-[[AHL|3OC12-HSL]]. | ||
=== First Level crosstalk: LasR binds to different HSL and activates the promoter === | === First Level crosstalk: LasR binds to different HSL and activates the promoter === | ||
| + | In the conventional system [[AHL|3OC12-HSL]] binds to its corresponding regulator, [https://parts.igem.org/Part:BBa_C0179 LasR], and activates the pLas promoter (figure 2, red). However, LasR can potentially also bind other AHLs and then activate pLas (Figure 1, [[3OC6HSL|3OC6-HSL]] in light blue and [[AHL|C4-HSL]] in green). This leads then to unwanted gene expression (crosstalk). | ||
| − | [[File:ETH Zurich 1crosstalkPlas.png|400px|center]] | + | [[File:ETH Zurich 1crosstalkPlas.png|400px|thumb|center| '''Figure 1 Overview of possible crosstalk of the [https://parts.igem.org/Part:BBa_C0179 LasR]/[https://parts.igem.org/Part:BBa_R0079 pLas] system with three different [[AHL|AHLs]].''' Usually, [[AHL|3OC12-HSL]] binds to its corresponding regulator, [https://parts.igem.org/Part:BBa_C0179 LasR], and activates the [https://parts.igem.org/Part:BBa_C0179 pLas] promoter (red). However, [https://parts.igem.org/Part:BBa_R0079 LasR] may also bind [[3OC6HSL|3OC6-HSL]] (light blue) or [[AHL|C4-HSL]] (green) and then unintentionally activate [https://parts.igem.org/Part:BBa_R0079 pLas].]] |
| − | === Second Level crosstalk: other regulatory proteins, like LuxR, bind to their natural | + | ==== Second Level crosstalk: other regulatory proteins, like LuxR and RhlR, bind to their natural AHL substrate and activate the pLas promoter ==== |
| − | [[File:ETH_Zurich_2crosstalkPlas.png|400px|center]] | + | In the conventional system [[AHL|3OC12-HSL]] binds to its corresponding regulator, [https://parts.igem.org/Part:BBa_C0179 LasR], and activates the [https://parts.igem.org/Part:BBa_R0079 pLas] promoter (Figure 2, red). However, [https://parts.igem.org/Part:BBa_R0079 pLas] can potentially be activate by other regulators ([https://parts.igem.org/Part:BBa_R0062 LuxR], [https://parts.igem.org/Part:BBa_R0062 RhlR]), binding their corresponding regulator (Figure 2, [[3OC6HSL|3OC6-HSL]] in light blue, [[AHL|C4-HSL]] in green). This leads then to unwanted gene expression (crosstalk). |
| + | |||
| + | [[File:ETH_Zurich_2crosstalkPlas.png|thumb|400px|center|'''Figure 2 Overview of possible crosstalk of the [https://parts.igem.org/Part:BBa_C0179 LasR]/[https://parts.igem.org/Part:BBa_R0079 pLas] system with two additional regulators ([https://parts.igem.org/Part:BBa_R0062 LuxR] and [https://parts.igem.org/Part:BBa_C0171 RhlR]).''' Usually, [https://parts.igem.org/Part:BBa_C0179 LasR] together with inducer [[AHL|3OC12-HSL]] activate their corresponding promoter [https://parts.igem.org/Part:BBa_R0079 pLas] (red). However, [https://parts.igem.org/Part:BBa_R0079 pLas] may also be activated by the [https://parts.igem.org/Part:BBa_R0062 LuxR] regulator together with [[3OC6AHL|3OC6-AHL]] (light blue) or by the [https://parts.igem.org/Part:BBa_C0171 RhlR] regulator together with [[AHL|C4-AHL]] (green).]] | ||
== Second order crosstalk: Combination of both cross-talk levels == | == Second order crosstalk: Combination of both cross-talk levels == | ||
| − | + | The second order crosstalk describes unintended activation of [https://parts.igem.org/Part:BBa_R0079 pLas] by a mixture of both the levels described above. The regulator and inducer are being different from [https://parts.igem.org/Part:BBa_C0179 LasR] and [[AHL|3OC12-HSL]], respectively, and at the same time they do not belong to the same module. For example, the inducer [[AHL|C4-HSL]] (green), usually binding to the regulator [https://parts.igem.org/Part:BBa_C0171 RhlR], could potentially interact with [https://parts.igem.org/Part:BBa_R0062 LuxR] regulator (light blue) and together activate [https://parts.igem.org/Part:BBa_R0079 pLas] (red). This kind of crosstalk is explained in Figure 3. | |
| − | [[File:ETh Zurich 3crosstalkPlas.png|400px|center]] | + | [[File:ETh Zurich 3crosstalkPlas.png|400px|thumb|center| '''Figure 3 Overview of possible crosstalk of the [https://parts.igem.org/Part:BBa_R0079 pLas] promoter with both the regulator and inducer being unrelated to the promoter and each other.''' Usually, [https://parts.igem.org/Part:BBa_C0179 LasR] together with inducer [[AHL|3OC12-HSL]] activate their corresponding promoter [https://parts.igem.org/Part:BBa_R0079 pLas] (red). However, [https://parts.igem.org/Part:BBa_R0079 pLas] may also be activated by another regulator together with an unrelated inducer. For example, the inducer [[AHL|C4-HSL]] (green) may interact with the [https://parts.igem.org/Part:BBa_R0062 LuxR] regulator (light blue) and together activate [https://parts.igem.org/Part:BBa_R0079 pLas] (red).]] |
== Results == | == Results == | ||
{|class="wikitable" style="background-color: white; text-align:center; width:auto; margin: auto;" | {|class="wikitable" style="background-color: white; text-align:center; width:auto; margin: auto;" | ||
| − | |+'''Table | + | |+'''Table 1''' Crosstalk matrix for the promoter plas ([https://parts.igem.org/Part:BBa_R0079:Experience BBa_R0079]) |
|colspan="4" style='font-size:10pt';text-align:left| | |colspan="4" style='font-size:10pt';text-align:left| | ||
| Line 66: | Line 72: | ||
|- | |- | ||
|[[File:ETH_Zurich_2014_qs-table_LasR.png|100px|link=https://parts.igem.org/Part:BBa_C0179:Experience]] | |[[File:ETH_Zurich_2014_qs-table_LasR.png|100px|link=https://parts.igem.org/Part:BBa_C0179:Experience]] | ||
| − | |[[File:ETH_Zurich_2014_qs-table_PlasRef.png| | + | |[[File:ETH_Zurich_2014_qs-table_PlasRef.png|200px]] |
| − | |[[File:ETH_Zurich_2014_qs-table_PlasLasRLuxAHL.png| | + | |[[File:ETH_Zurich_2014_qs-table_PlasLasRLuxAHL.png|200px]] |
| − | |[[File:ETH_Zurich_2014_qs-table_PlasLasRRhlAHL.png| | + | |[[File:ETH_Zurich_2014_qs-table_PlasLasRRhlAHL.png|200px]] |
|- | |- | ||
|[[File:ETH_Zurich_2014_qs-table_LuxR.png|100px|link=https://parts.igem.org/Part:BBa_C0062:Experience]] | |[[File:ETH_Zurich_2014_qs-table_LuxR.png|100px|link=https://parts.igem.org/Part:BBa_C0062:Experience]] | ||
| − | |[[File:ETH_Zurich_2014_qs-table_PlasLuxRLasAHL.png| | + | |[[File:ETH_Zurich_2014_qs-table_PlasLuxRLasAHL.png|200px]] |
| − | |[[File:ETH_Zurich_2014_qs-table_PlasLuxRLuxAHL.png| | + | |[[File:ETH_Zurich_2014_qs-table_PlasLuxRLuxAHL.png|200px]] |
| − | |[[File:ETH_Zurich_2014_qs-table_PlasLuxRRhlAHL.png| | + | |[[File:ETH_Zurich_2014_qs-table_PlasLuxRRhlAHL.png|200px]] |
|- | |- | ||
|[[File:ETH_Zurich_2014_qs-table_RhlR.png|100px|link=https://parts.igem.org/Part:BBa_C0171:Experience]] | |[[File:ETH_Zurich_2014_qs-table_RhlR.png|100px|link=https://parts.igem.org/Part:BBa_C0171:Experience]] | ||
| − | |[[File:ETH_Zurich_2014_qs-table_PlasRhlRLasAHL.png| | + | |[[File:ETH_Zurich_2014_qs-table_PlasRhlRLasAHL.png|200px]] |
| − | |[[File:ETH_Zurich_2014_qs-table_PlasRhlRLuxAHL.png| | + | |[[File:ETH_Zurich_2014_qs-table_PlasRhlRLuxAHL.png|200px]] |
| − | |[[File:ETH_Zurich_2014_qs-table_PlasRhlRRhlAHL.png| | + | |[[File:ETH_Zurich_2014_qs-table_PlasRhlRRhlAHL.png|200px]] |
|} | |} | ||
| + | === Modeling crosstalk === | ||
| + | |||
| + | Each experimental data set was fitted to an Hill function using the Least Absolute Residual method. | ||
| + | |||
| + | [[File:ETHZ_HillEq.png|center|200px]] | ||
| + | <p>The fitting of the graphs was performed using the following equation :<br><br> | ||
| + | rFluo = the relative fluorescence (absolute measured fluorescence value over OD)[a.u.]<br> | ||
| + | a = basal expression rate [a.u.](“leakiness”)<br> | ||
| + | b = maximum expression rate [a.u.]("full induction")<br> | ||
| + | n = Hill coefficient (“cooperativity”)<br> | ||
| + | K<sub>m</sub> = Half-maximal effective concentration (“sensitivity”)<br> | ||
| + | [AHL] = AHL concentration [nM]</p> | ||
| + | <br clear="all"/> | ||
| + | |||
| + | {| border="1" class="wikitable" style="margin: 1em auto 1em auto;text-align:center;" | ||
| + | |+ Parameters of HillFunction for crosstalk with Plas (with 95% confidence bounds) | ||
| + | ! | ||
| + | ! [[AHL|3OC12-HSL]] | ||
| + | ! [[3OC6HSL|3OC6-HSL]] | ||
| + | ! [[AHL|C4-HSL]] | ||
| + | |- | ||
| + | ! [https://parts.igem.org/Part:BBa_C0179 LasR] | ||
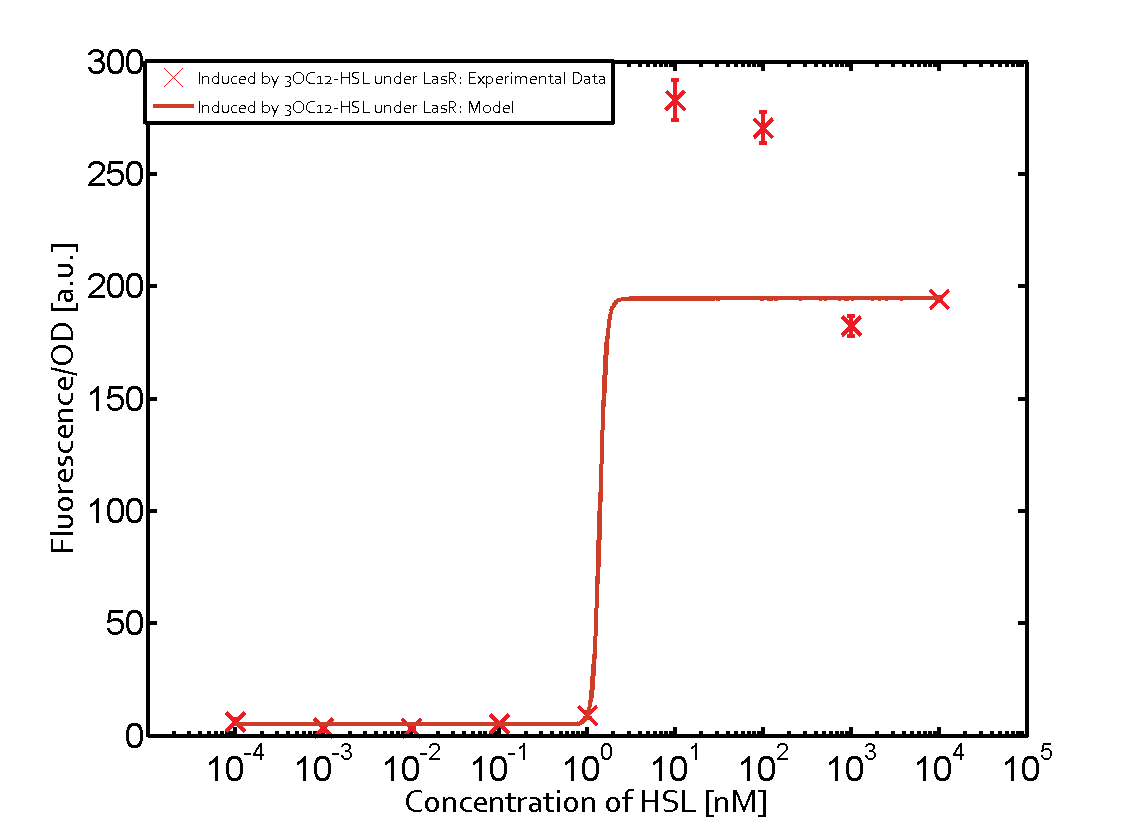
| + | | a = 5.274 (0, 15.51) [a.u.]<br> n = 11.67 (0, 3.763e8)<br> Km = 1.396 (0, 1.504e7) [nM]<br>b = 189 (162.5, 215.6) [a.u.]<br> | ||
| + | | No crosstalk | ||
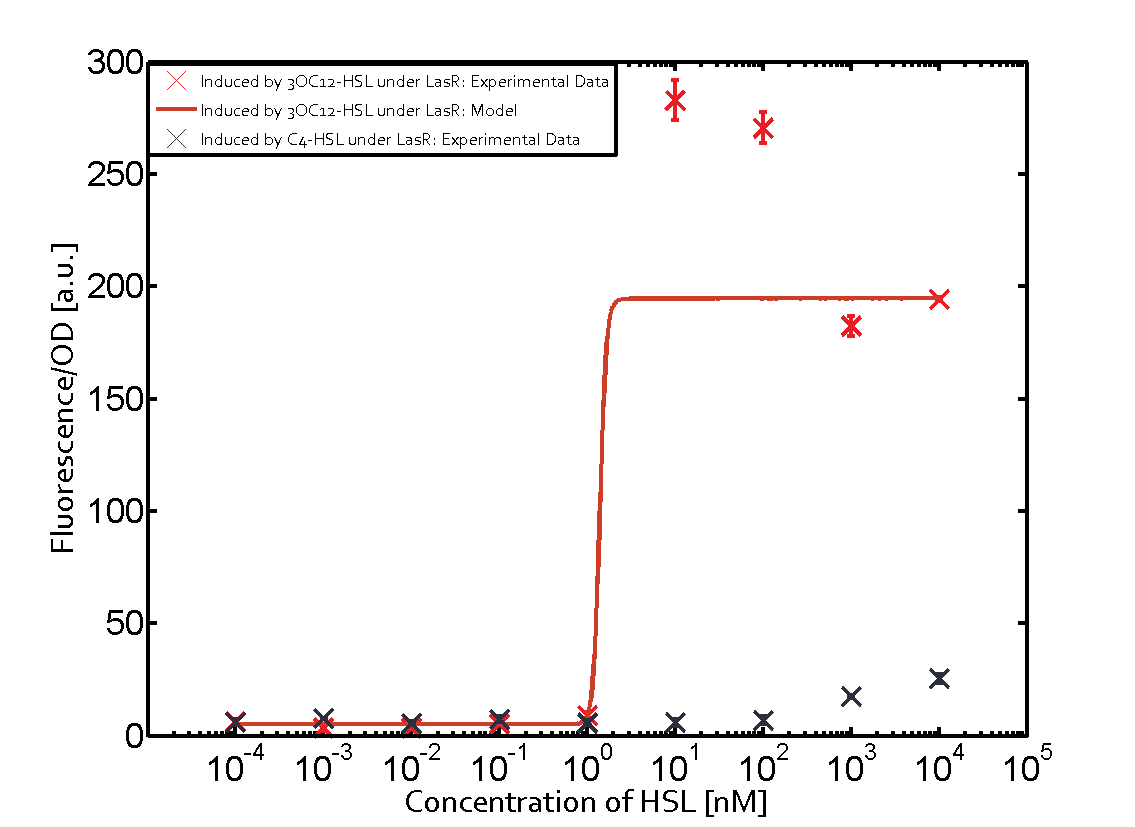
| + | | No crosstalk | ||
| + | |- | ||
| + | ! [https://parts.igem.org/Part:BBa_C0062 LuxR] | ||
| + | | No crosstalk | ||
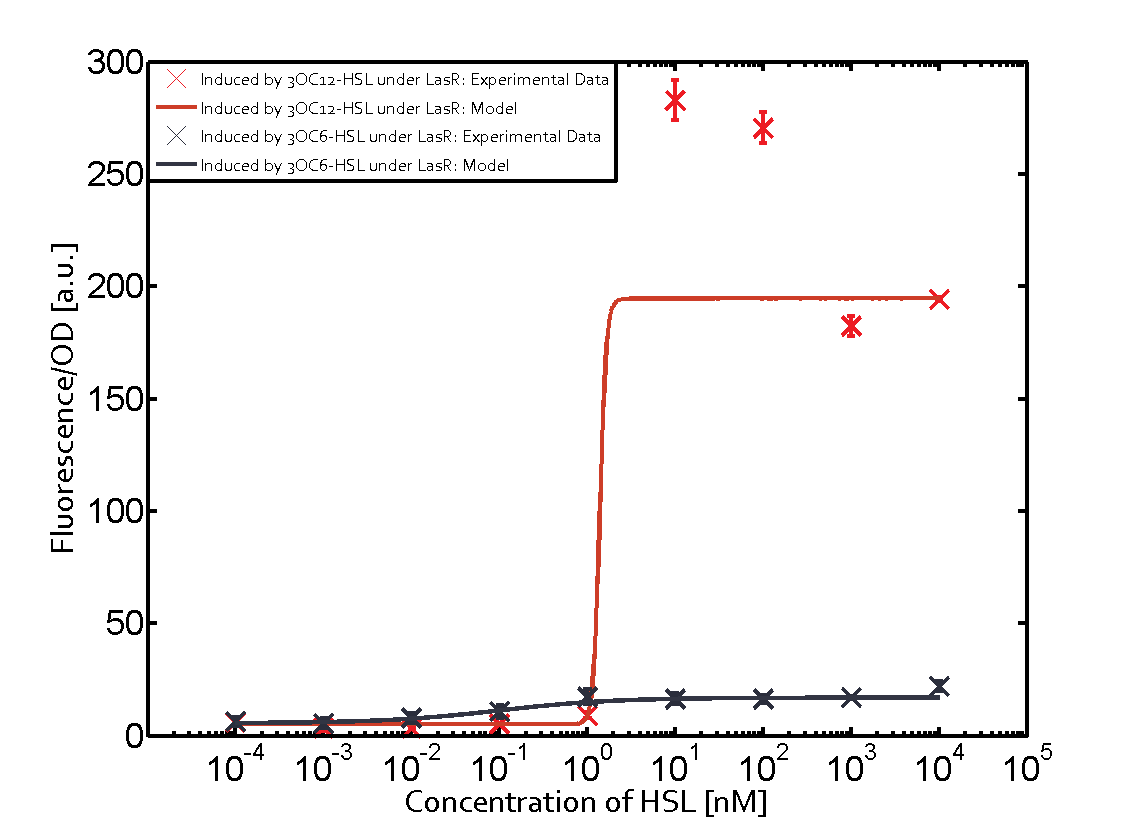
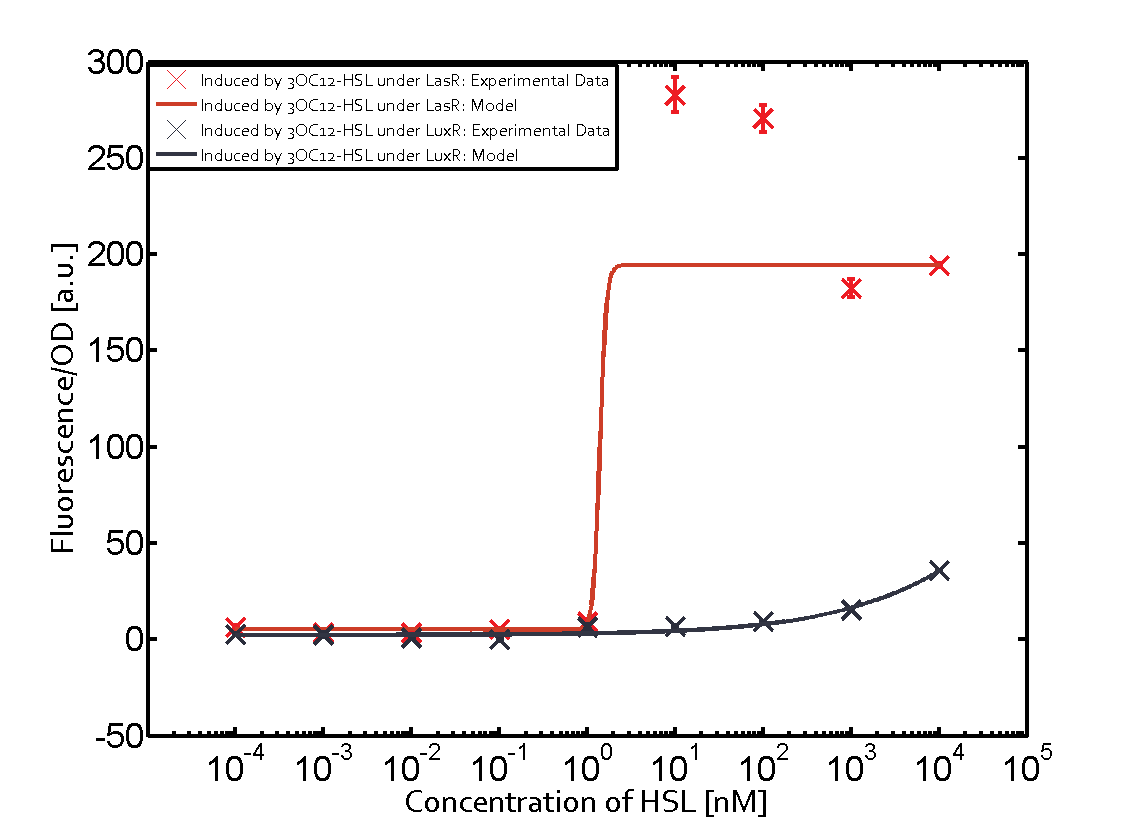
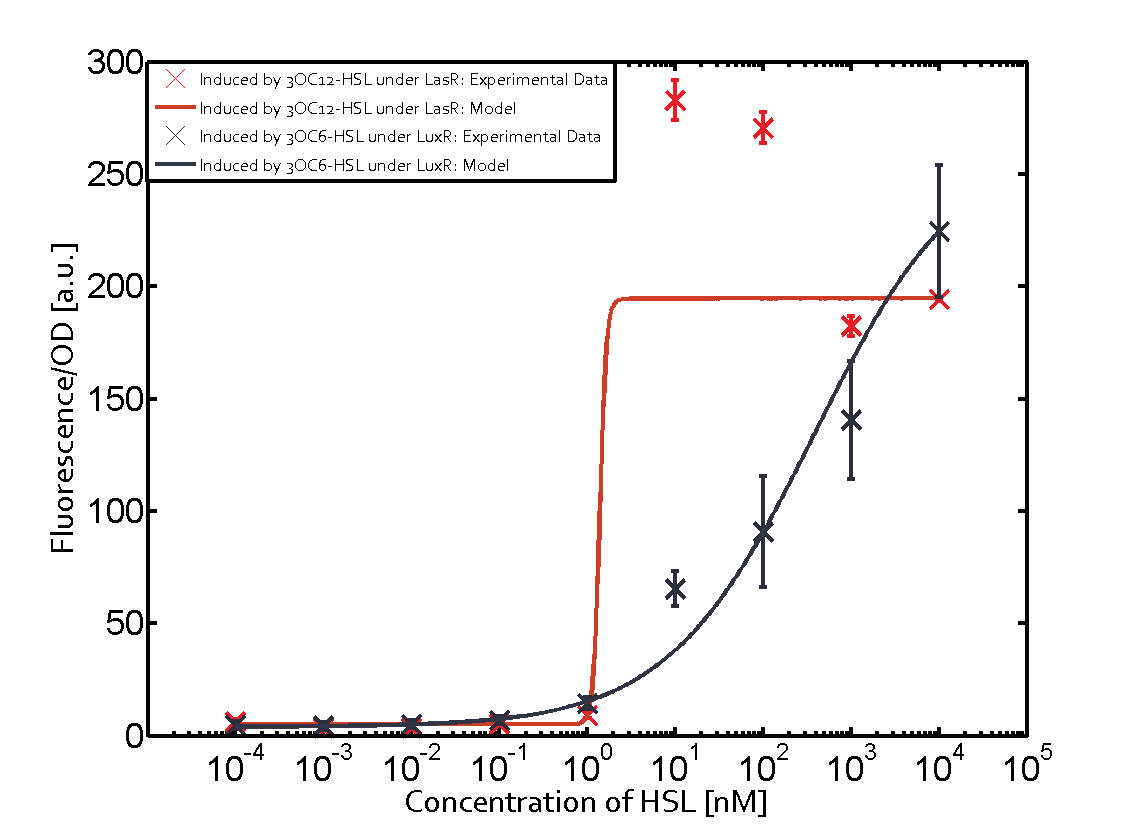
| + | | a = 3.996 (0, 9.185) [a.u.]<br> n = 0.5198 (0.2879, 0.7517)<br> Km = 382.3 (0, 1206) [nM]<br>b = 261 (155.5, 366.5) [a.u.]<br> | ||
| + | | No crosstalk | ||
| + | |- | ||
| + | ! [https://parts.igem.org/Part:BBa_C0171 RhlR] | ||
| + | | No crosstalk | ||
| + | | No crosstalk | ||
| + | | No crosstalk | ||
| + | |} | ||
|} | |} | ||
Latest revision as of 16:10, 26 October 2014
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_R0079
User Reviews
UNIQ68bdbfdfaffc3103-partinfo-00000000-QINU UNIQ68bdbfdfaffc3103-partinfo-00000001-QINU
|
••••
ETH Zurich 2014 |
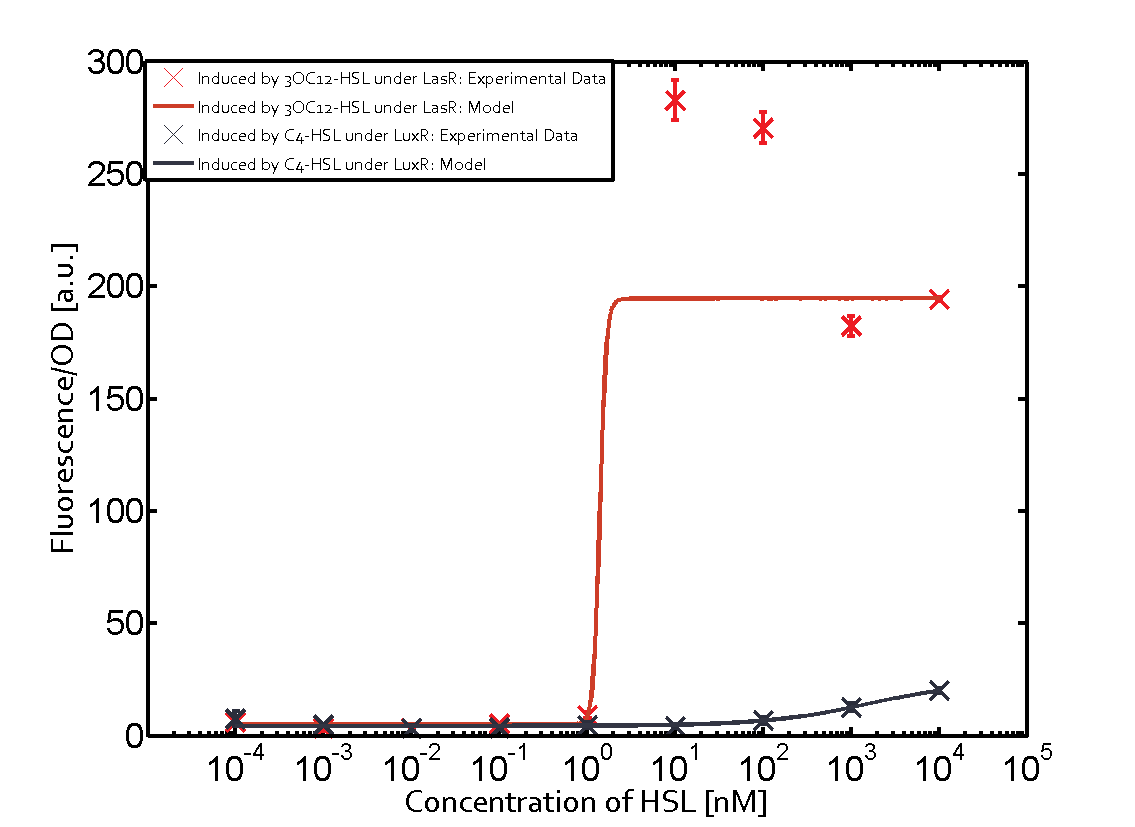
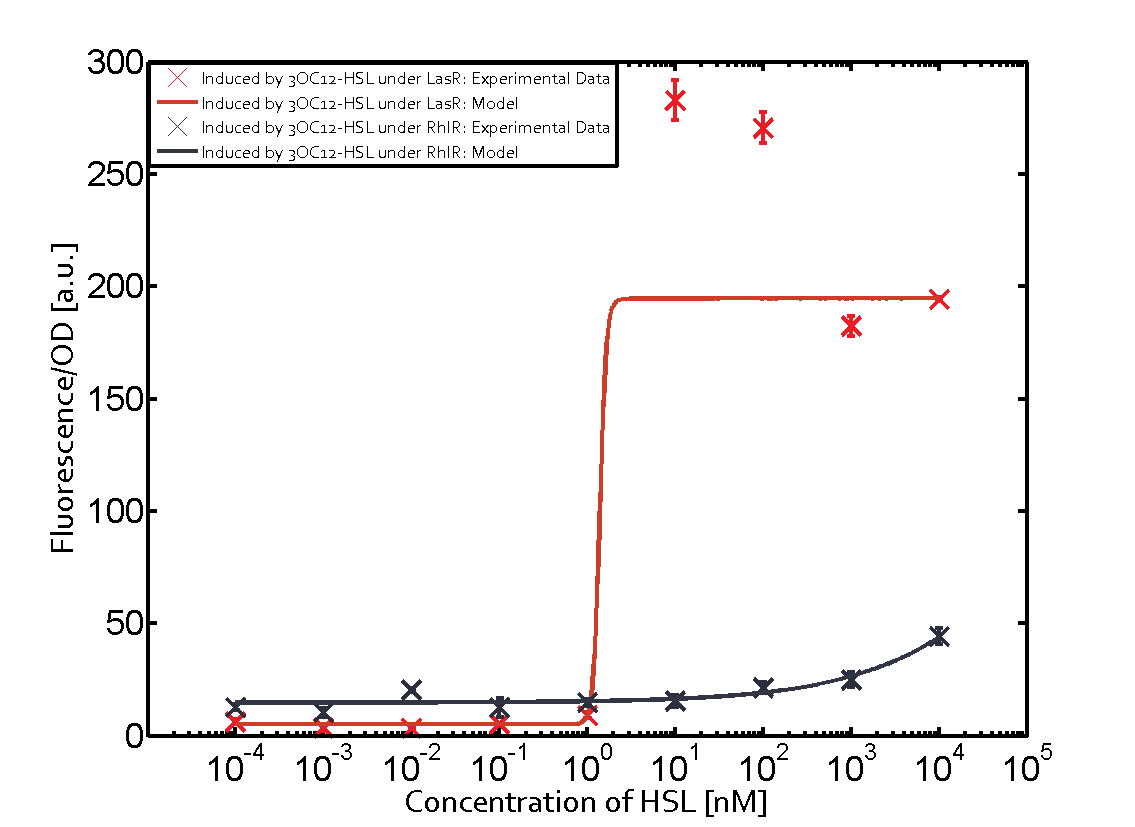
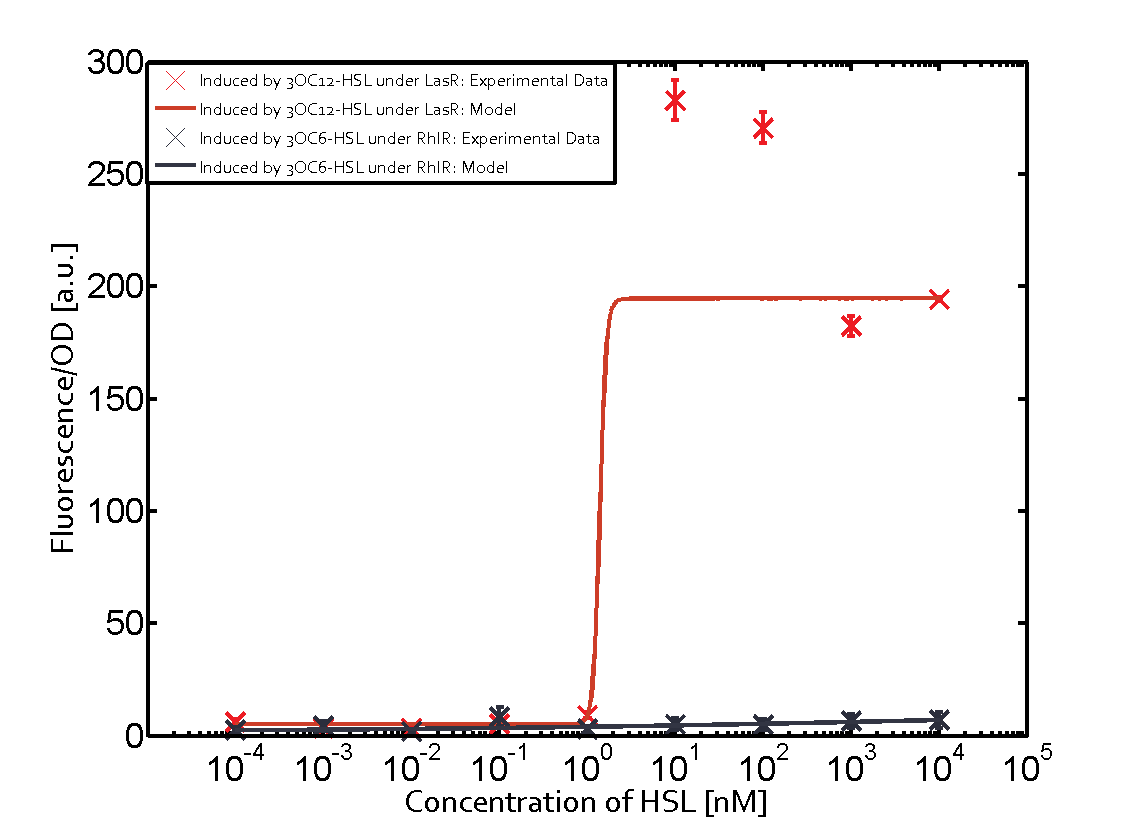
Characterization of CrosstalkBackground informationWe used an E. coli TOP10 strain transformed with two medium copy plasmids (about 15 to 20 copies per plasmid and cell). The first plasmid contained the commonly used p15A origin of replication, a kanamycin resistance gene, and promoter pLas (BBa_R0079) followed by RBS (BBa_B0034) and superfolder green fluorescent protein (sfGFP). In general, for spacer and terminator sequences the parts BBa_B0040 and BBa_B0015 were used, respectively. The second plasmid contained the pBR322 origin (pMB1), which yields a stable two-plasmid system together with p15A, an ampicillin resistance gene, and a strong promoter (BBa_J23100) chosen from the Anderson promoter collection followed by one of the three different regulators (LuxR, LasR, and RhlR) used in the experiments in order to quantify crosstalk with pLas. The detailed regulator construct design and full sequences (piG0040, piG0041, piG0042, piG0050) are [http://2014.igem.org/Team:ETH_Zurich/lab/sequences available here]. In the following, we describe the experimental set-up and all the different levels of crosstalk we have assessed. Experimental Set-UpThe above described E. coli TOP10 strains were grown overnight in Lysogeny Broth (LB) containing kanamycin (50 μg/mL) and ampicillin (200 μg/mL) to an OD600 of about 1.5 (37 °C, 220 rpm). As a reference, a preculture of the same strain lacking the sfGFP gene was included for each assay. The cultures were then diluted 1:40 in fresh LB containing the appropriate antibiotics and measured in triplicates in microtiter plate format on 96-well plates (200 μL culture volume) for 10 h at 37 °C with a Tecan infinite M200 PRO plate reader (optical density measured at 600 nm; fluorescence with an excitation wavelength of 488 nm and an emission wavelength of 530 nm). After 200 min we added the following concentrations of inducers (3OC6-HSL, 3OC12-HSL, and C4-HSL): 10-4 nM and 104 nM (from 100 mM stocks in DMSO). Attention: All the dilutions of 3OC12-HSL should be made in DMSO in order to avoid precipitation. In addition, in one triplicate only H2O was added as a control. From the the obtained kinetic data, we calculated mean values and plotted the dose-response-curves for 200 min past induction. First-order crosstalkIn the first order crosstalk section we describe crosstalk of pLas due to LasR binding to inducers different from 3OC12-HSL or pLas itself binding to a regulator-inducer pair different from LasR-3OC12-HSL. First Level crosstalk: LasR binds to different HSL and activates the promoterIn the conventional system 3OC12-HSL binds to its corresponding regulator, LasR, and activates the pLas promoter (figure 2, red). However, LasR can potentially also bind other AHLs and then activate pLas (Figure 1, 3OC6-HSL in light blue and C4-HSL in green). This leads then to unwanted gene expression (crosstalk). 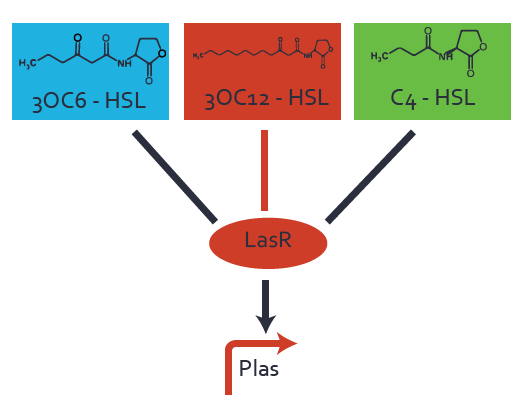 Figure 1 Overview of possible crosstalk of the LasR/pLas system with three different AHLs. Usually, 3OC12-HSL binds to its corresponding regulator, LasR, and activates the pLas promoter (red). However, LasR may also bind 3OC6-HSL (light blue) or C4-HSL (green) and then unintentionally activate pLas. Second Level crosstalk: other regulatory proteins, like LuxR and RhlR, bind to their natural AHL substrate and activate the pLas promoterIn the conventional system 3OC12-HSL binds to its corresponding regulator, LasR, and activates the pLas promoter (Figure 2, red). However, pLas can potentially be activate by other regulators (LuxR, RhlR), binding their corresponding regulator (Figure 2, 3OC6-HSL in light blue, C4-HSL in green). This leads then to unwanted gene expression (crosstalk). 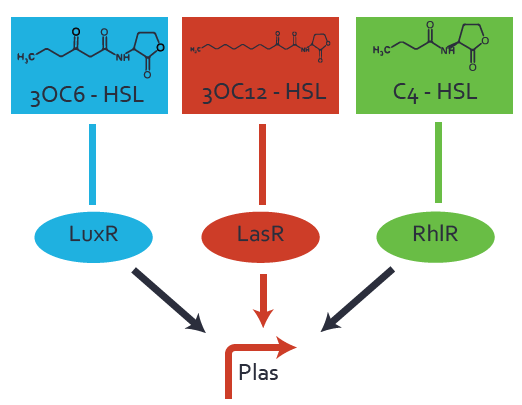 Figure 2 Overview of possible crosstalk of the LasR/pLas system with two additional regulators (LuxR and RhlR). Usually, LasR together with inducer 3OC12-HSL activate their corresponding promoter pLas (red). However, pLas may also be activated by the LuxR regulator together with 3OC6-AHL (light blue) or by the RhlR regulator together with C4-AHL (green). Second order crosstalk: Combination of both cross-talk levelsThe second order crosstalk describes unintended activation of pLas by a mixture of both the levels described above. The regulator and inducer are being different from LasR and 3OC12-HSL, respectively, and at the same time they do not belong to the same module. For example, the inducer C4-HSL (green), usually binding to the regulator RhlR, could potentially interact with LuxR regulator (light blue) and together activate pLas (red). This kind of crosstalk is explained in Figure 3. 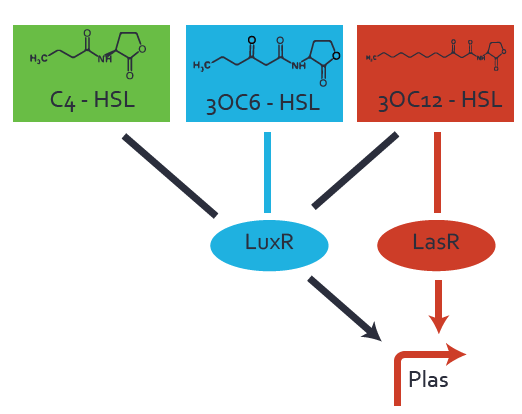 Figure 3 Overview of possible crosstalk of the pLas promoter with both the regulator and inducer being unrelated to the promoter and each other. Usually, LasR together with inducer 3OC12-HSL activate their corresponding promoter pLas (red). However, pLas may also be activated by another regulator together with an unrelated inducer. For example, the inducer C4-HSL (green) may interact with the LuxR regulator (light blue) and together activate pLas (red). Results
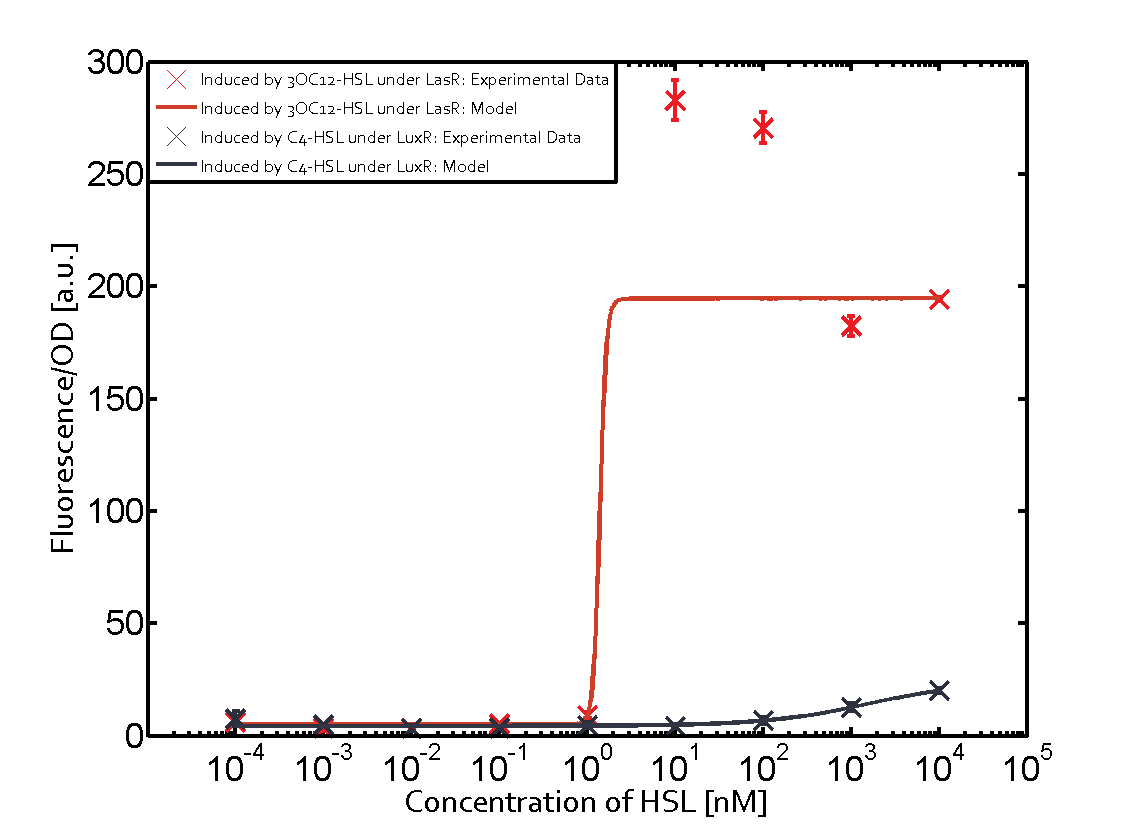
Modeling crosstalkEach experimental data set was fitted to an Hill function using the Least Absolute Residual method. The fitting of the graphs was performed using the following equation :
| ||||||||||||||||||||||||||||||||||||
|
No review score entered. NYMU-Taipei 2009 |
|
|
No review score entered. Northwestern 2011 |
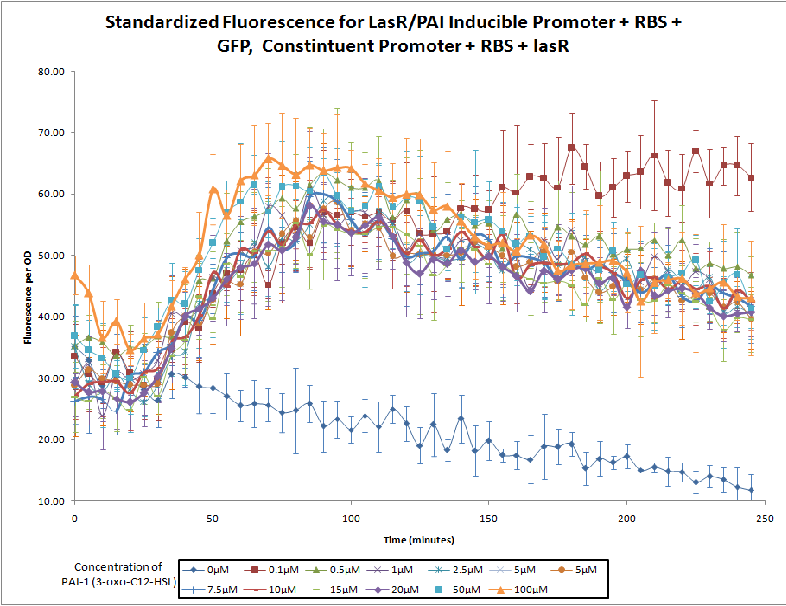
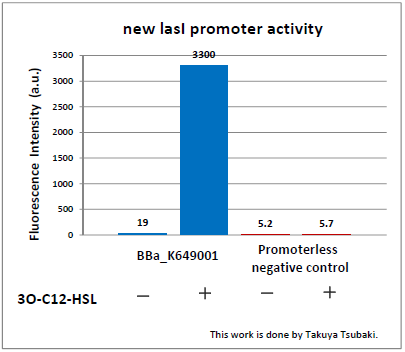
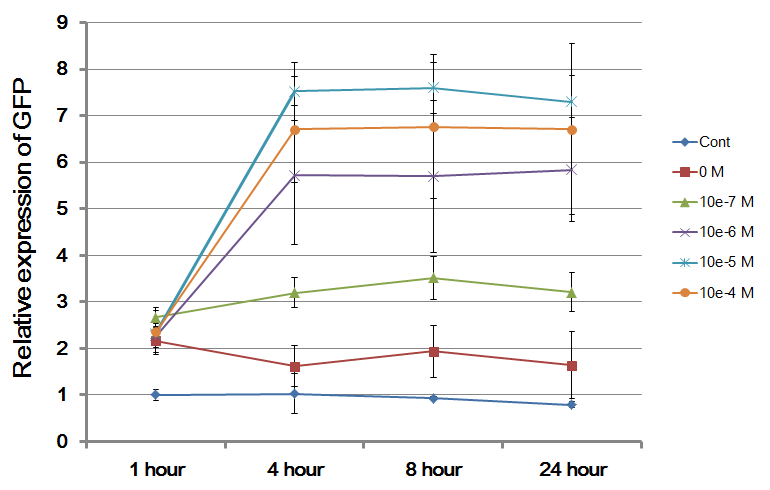
The Northwestern iGEM team used this part as a unit within our Pseudomonas Aeruginosa biosensor. When this LasR/PAI regulated promoter is induced at varying concentrations of PAI in the presence of excess LasR, we observed GFP fluorescence in accordance with the graph below. |
|};
|
•••
Tokyo-tech iGEM 2011 |
Judging from Northwestern iGEM 2011 team's data, in the presence of 3OC12-HSL, fluorescence intensity was about 3-fold higher than that in the absence of 3OC12-HSL.
|
|};
|
No review score entered. Tsinghua-A 2011 |
Tsinghua-A 2011 assembled E0840 (BBa_E0840) under the pLas promoter (BBa_R0079) that was contained into K574009 (BBa_K574009). We kept pSB1A2 as the scaffold vector.
|
|
•••••
iGEM Dundee 2014 |
Dundee iGEM 2014 used this lasB promoter region to build a composite part BBa_K1315009. This was designed as a biosensor for Pseudomonas aeruginosa AutoInducer-1 (PAI-1), which was to be used in a bio-electronic device to improve diagnostics for Cystic Fibrosis patients. Details of experimental work are logged on the experience page of BBa_K1315009. |