Part:BBa_K2656014
mRFP1 Coding Sequence
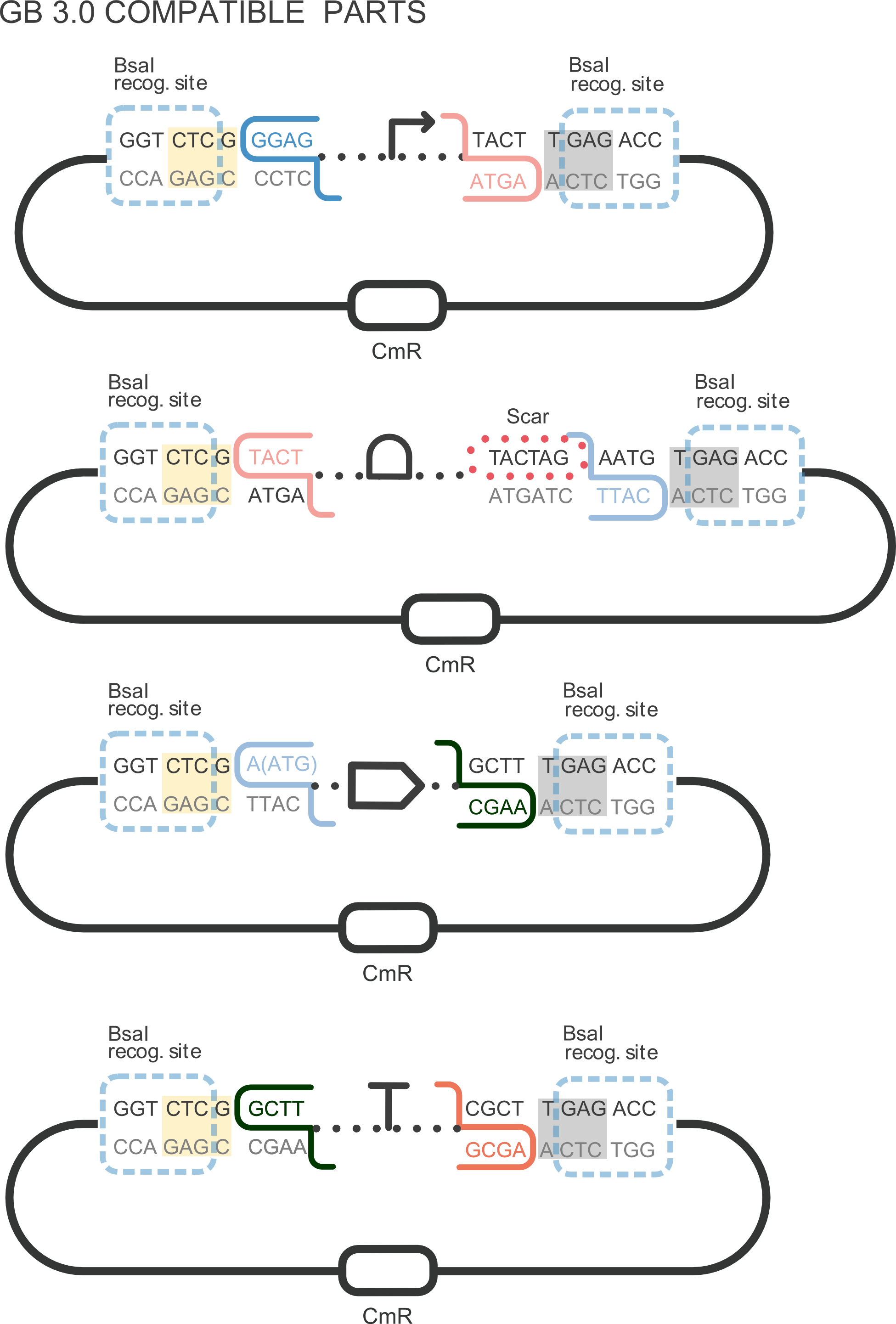
Part BBa_K2656014 is the monomeric Red Fluorescent Protein 1 coding sequence BBa_E1010 adapted into the [http://2018.igem.org/Team:Valencia_UPV/Design Golden Braid assembly method]. Thus, this sequence is both compatible with the BioBrick and GoldenBraid 3.0. grammar.
This coding sequence can be combined with other Golden Braid compatible parts from our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection Valencia UPV IGEM 2018 Printeria Collection] to assemble transcriptional units in a one-step BsaI reaction with the Golden Gate assembly protocol .
The characterization of this protein (and by extension of all the other part that codify for the mRFP1) was performed with our transcriptional unit BBa_K2656109. This transcriptional unit was assembled in a GoldenBraid alpha 1 plasmid including the following parts:
- BBa_K2656004: the J23106 promoter in its Golden Braid compatible version from our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection Part Collection]
- BBa_K2656009: the B0030 ribosome biding site in its Golden Braid compatible version from our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection Part Collection]
- BBa_K2656014: coding sequence
- BBa_K2656026: the B0015 transcriptional terminator in its Golden Braid compatible version from our [http://2018.igem.org/Team:Valencia_UPV/Part_Collection Part Collection]
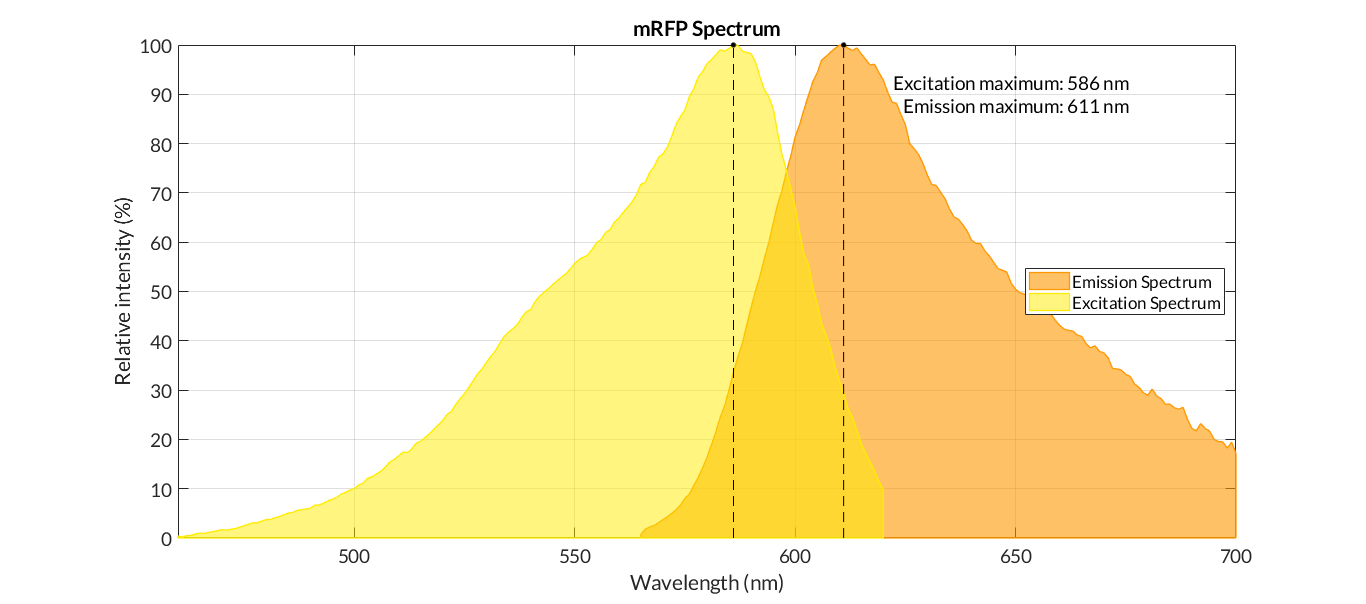
In order to carry out a correct characterization of the protein and to be able to use it to make measurements of the different transcriptional units that we assembled with it, we obtained the emission and excitation spectra in the conditions of our equipment. By using this [http://2018.igem.org/Team:Valencia_UPV/Experiments#spectra protocol] with the parameters of Table 1, Figure 1 was obtained.
| Parameter | Value | ||
| Number of samples | 3 | ||
| Excitation Wavelength measurement range 1 (nm) | [450-620] | ||
| Excitation Wavelength measurement range 2 (nm) | [620-700] | ||
| Emission wavelenght 1 (nm) | 650 | ||
| Emission wavelenght 2 (nm) | 590 | ||
| Emission Wavelength measurement range (nm) | [565-700] | ||
| Excitation wavelenght (nm) | 540 | ||
| Gain (G) | 70 | ||
| Table 1. Parameters used to obtain the spectra | |||
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 556
Illegal AgeI site found at 668 - 1000COMPATIBLE WITH RFC[1000]
//function/reporter
//function/reporter/fluorescence
//test
| None |