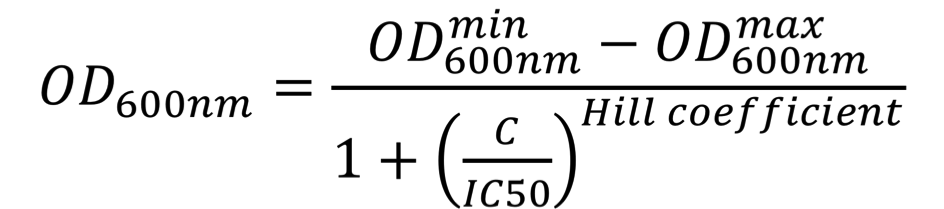Part:BBa_K3766054
This part is AZT-4, an AmpR variant evolved to confer resistance to aztreonam.
Usage and Biology
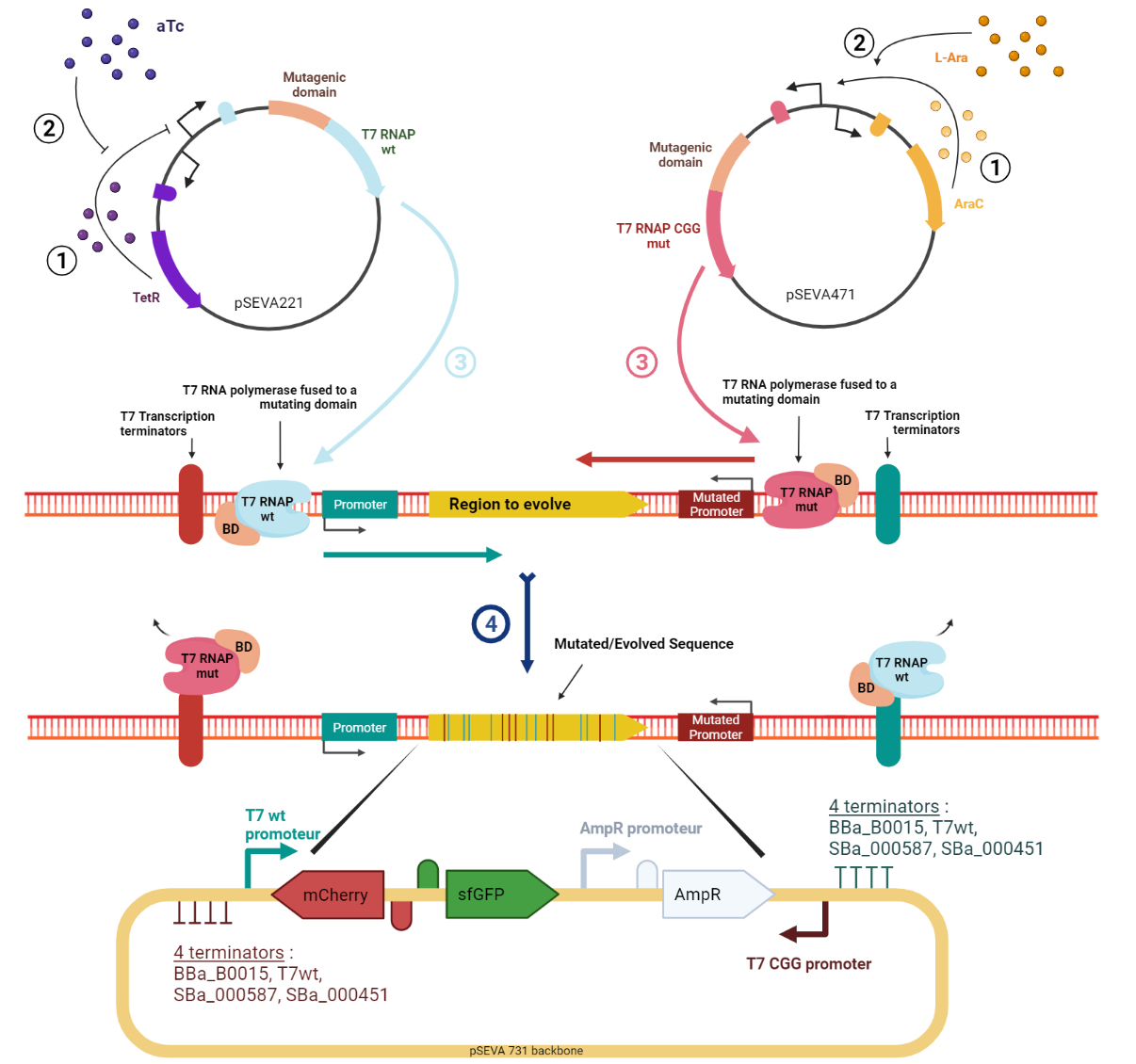
Evolution.T7 is a targeted directed evolution system based on the orthogonal RNA polymerases of phase T7 (T7RNAP) [1–4]. Mutagenesis is performed by a mutator protein (e.g. base deaminase) linked to the polymerase and, thanks to T7RNAP’s specificity towards the T7 promoter, it is possible to target the mutagenesis to a sequence flanked by T7 promoter and terminator. The high processivity of the T7RNAP allows for mutagenesis to occur over a long sequence of DNA without showing bias to the distance from the promoter. However, a bias was observed in the current tools [1–4] which was that mainly only one of the strands of DNA would be targeted for mutation at each mutation cycle, resulting in poorly diversified sequences. To address this, we designed the target sequence putting the T7 promoter upstream and downstream of the gene to have polymerases moving in both directions and generating mutations in both strands. This approach has been tried by Moore ‘’et al.’’ [1] but they found out that this would lead to the accumulation of A:T pairs along the gene. Also, we assumed that owing to the high processivity of T7RNAP, often there are active transcription machinery moving in both directions and thus, there is a high probability of collision resulting in the reduced processivity of both complexes. This phenomenon can lead to the accumulation of mutations on one DNA strand in one part of the gene and more mutations on the other strand in the other part. To bypass this, our innovative design takes advantage of a mutant T7RNAP that has higher specificity to an altered T7 promoter sequence [5]. The canonical and mutant T7 promoters are installed upstream and downstream of the gene, respectively (Figure 1). The wild-type T7RNAP expression is induced by tetracycline, while the mutant T7RNAP is induced by arabinose. In addition. to enhance the mutation rate and diversity, our tool expands the panoply of base base deaminases by including, apart from the ones used in previously described methods (AID, pmCDA1, rAPOBEC1, TadA*), three of their potent variants ABE8.20m [6], evoAPOBEC1-BE4max [7], evoCDA1-BE4max [7]. In addition to higher efficiency, these variants generate mutations without being affected by the base context of the substrate, resulting in higher target compatibility.
Figure 1. Schematic representation of the Evolution.T7 system. T7RNAP is the natural T7 RNA polymerase that transcribes from the T7wt promoter. T7RNAP-CGG-R12-KIRV is a mutated T7 RNA polymerase version that transcribes from the mutant T7CGG promoter. Image created in BioRender.com.
As a proof of concept of our targeted directed evolution system, Evolution.T7, we evolved the ampicillin resistance gene, AmpR, in a way that it would confer resistance against aztreonam, which is a powerful antibiotic against ampicillin resistant bacteria [8,9].
AmpR is an E. coli gene coding for a β-lactamase enzyme. It is a derivative of the TEM-1 β-lactamase (Uniprot P62593) with two mutations V82I and V182A. This protein is associated with the resistance mechanism against the large family of β-lactams, antibiotics which include the well known penicillin and ampicillin.
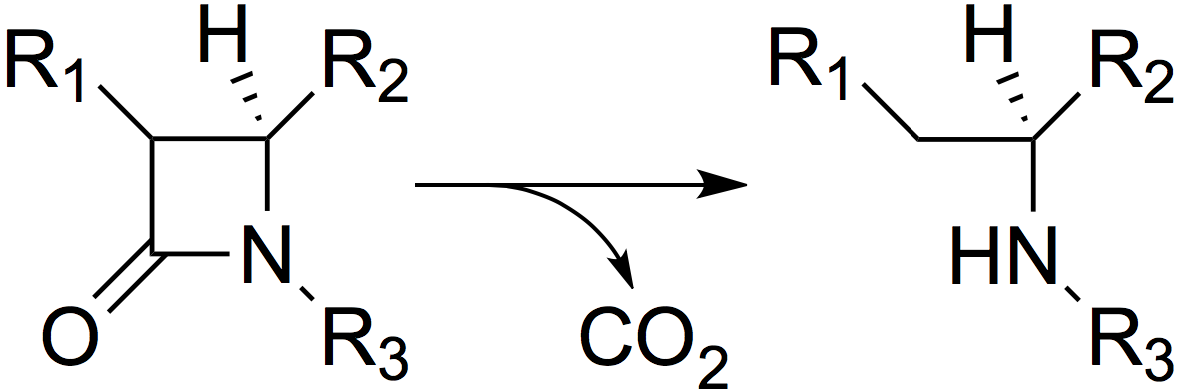
β-Lactamases act by hydrolysing the β-lactam ring (Figure 2), thus inactivating the antibiotic.
Figure 2. The mode of action of β-lactamases: hydrolysis of the β-lactam ring.
Fortunately, among all the subgroups of β-lactams, some have less affinity with the β-lactamase(s) and overcome this resistance. For example, in monobactams, Aztreonam is known to have a bactericidal effect even on bacteria carrying the AmpR gene [10]. We confirmed this experimentally by testing the resistance of the bacteria carrying AmpR to different β-lactam antibiotics (see results on the Parts Registry page of BBa_K3766012).
For AmpR evolution, we decided to use E. coli MG1655 cells harboring two mutations Δung and Δnfi that were shown to be important for enhancing the mutation rate of the mutagenic system [2]. ung is a gene encoding the uracil DNA N-glycosilase that is involved in the elimination of uracil from DNA molecules [11], while nif is a gene encoding the endonuclease V of E. coli which eliminates inosines [12,13]. Uracil and inosine are nucleobases produced by the action of cytosine and adenine deaminases, respectively, which are the mutator domains of our Evolution.T7 system. E. coli MG1655 Δung Δnfi [2] were first transformed with a plasmid carrying the AmpR resistance gene (BBa_K3766012) within the Evolution.T7 system mutational region, BBa_K3766111 in pSEVA73 backbone [14], together with different combinations of forward and reverse mutators in pSEVA22 and pSEVA47 backbones, respectively (Figure 1). Transformed bacteria were first grown overnight in 96-deep-well plates containing 1 mL of LB medium supplemented with the relevant antibiotics (5 µg/mL trimethoprim, 12.5 µg/mL kanamycin and 12.5 µg/mL spectinomycin), then diluted by 40x into similar media. Upon reaching early log-phase, cells were further diluted 20x in 100 µL of LB medium supplemented with the antibiotics (5 µg/mL trimethoprim, 12.5 µg/mL kanamycin, 12.5 µg/mL spectinomycin), and inducers (200 ng/mL anhydrotetracycline and 1.5 mM L-arabinose) in a 96-well microplate. The plate was incubated at 37 °C at 200 rpm and 5 µL of cell suspensions were spotted, 1, 3 and 24 hours later, onto LB agar plate containing increasing concentrations of aztreonam.
This part is one of the AmpR variants evolved to confer resistance to aztreonam (BBa_K3766051 to BBa_K3766084).
AZT-4 was isolated on a plate containing 5 µg/mL aztreonam. The Evolution.T7 system contained only the cytosine deaminase AID-CGG-R12-KIRV [2] (BBa_K3766101).
Sequence analysis of AZT-4 revealed the presence of 1 mutation (G485A) which lead to 1 mutation at protein level (R162H).
To further understand the impact of each mutation on aztreonam resistance, the IC50 values were estimated. For this, E. coli NEB5ɑ cells were first transformed with AZT-4 containing plasmids (in pSEVA73 backbone), then grown overnight in 96-deep-well plates containing 1 mL of LB medium supplemented with 10 µg/mL trimethoprim, then diluted by 40x into similar media. Upon reaching early log-phase, cells were further diluted 20x in 100 µL of LB medium supplemented with increasing concentrations of aztreonam (0 to 50 µg/mL) in a transparent 96-well polystyrene microplate (Sarstedt). The plate was incubated at 37 °C at 200 rpm and OD600nm was measured every 10 min for 24 hours in a CLARIOstar (BMGLabtech) or Infinite 500 (Tecan) plate reader. The OD600nm values at the time-point when the cultures performed in the absence of aztreonam reached the stationary phase were plotted as a function of the antibiotic concentration. These dose-response curves were fitted using the AAT Bioquest, Inc., Quest Graph™ IC50 Calculator to the sigmoidal équation:
where OD600nmmin represents the minimal OD600nm, OD600nmmax represents the maximal OD600nm, C is the concentration of the antibiotic, IC50 is the antibiotic concentration that inhibits 50% of the bacterial growth, Hill coefficient is describing the steepness of the curve. The OD600nm values were converted into equivalent particles using the calibration curves presented on the "Measurement" page of our wiki. Similarly, IC50 values for aztreonam were estimated for the chassis cells, E. coli NEB5ɑ with or without the parental AmpR (BBa_K3766012). In this case, the aztreonam range was lower (0.005 to 10 µg/mL).
For AZT-4, the aztreonam IC50 value was estimated at 7,3927 µg/mL (Figure 3), while the Aztreonam IC50 of the parental AmpR (BBa_K3766012) was of only 0.034 µg/mL aztreonam.
Figure 3. Dose-response curve used to calculate this part’s IC50 value of aztreonam.
In addition, we evaluated the capacity of this part to still confer resistance to ampicillin. As can be observed in Figure 4, the capacity of AZT-4 to confer resistance to ampicillin is similar to AmpR (BBa_K3766012), but the E. coli cells are still able to grow at 100 µg/ml ampicillin, the application concentration of this antibiotic in molecular biology for selection in the laboratory.
Figure 4. Pictures of E. coli cells carrying the AmpR resistance gene (BBa_K3766012) or this part plated on different concentrations of ampicillin.
References
[1] Moore CL, Papa LJ, Shoulders MD. A processive protein chimera introduces mutations across defined DNA regions in vivo. Journal of the American Chemical Society (2018) 140: 11560–11564.
[2] Álvarez B, Mencía M, de Lorenzo V, Fernández LÁ. In vivo diversification of target genomic sites using processive base deaminase fusions blocked by dCas9. Nature Communications (2020) 11: 6436.
[3] Park H, Kim S. Gene-specific mutagenesis enables rapid continuous evolution of enzymes in vivo. Nucleic Acids Research (2021) 49: e32.
[4] Cravens A, Jamil OK, Kong D, Sockolosky JT, Smolke CD. Polymerase-guided base editing enables in vivo mutagenesis and rapid protein engineering. Nature Communications (2021) 12: 1579.
[5] Meyer AJ, Ellefson JW, Ellington AD. Directed evolution of a panel of orthogonal T7 RNA polymerase variants for in vivo or in vitro synthetic circuitry. ACS synthetic biology (2015) 4: 1070–1076.
[6] Gaudelli NM, Lam DK, Rees HA, Solá-Esteves NM, Barrera LA, Born DA, Edwards A, Gehrke JM, Lee S-J, Liquori AJ, Murray R, Packer MS, Rinaldi C, Slaymaker IM, Yen J, Young LE, Ciaramella G. Directed evolution of adenine base editors with increased activity and therapeutic application. Nature Biotechnology (2020) 38: 892–900.
[7] Thuronyi BW, Koblan LW, Levy JM, Yeh W-H, Zheng C, Newby GA, Wilson C, Bhaumik M, Shubina-Oleinik O, Holt JR, Liu DR. Continuous evolution of base editors with expanded target compatibility and improved activity. Nature Biotechnology (2019) 37: 1070–1079.
[8] Brogden RN, Heel RC. Aztreonam. A review of its antibacterial activity, pharmacokinetic properties and therapeutic use. Drugs (1986) 31: 96–130.
[9] Ramsey C, MacGowan AP. A review of the pharmacokinetics and pharmacodynamics of aztreonam. The Journal of Antimicrobial Chemotherapy (2016) 71: 2704–2712.
[10] Cantu C, Huang W, Palzkill T. Selection and characterization of amino acid substitutions at residues 237-240 of TEM-1 beta-lactamase with altered substrate specificity for aztreonam and ceftazidime. The Journal of Biological Chemistry (1996) 271: 22538–22545.
[11] Pearl LH. Structure and function in the uracil-DNA glycosylase superfamily. Mutation Research (2000) 460: 165–181.
[12] Guo G, Ding Y, Weiss B. nfi, the gene for endonuclease V in Escherichia coli K-12. Journal of Bacteriology (1997) 179: 310–316.
[13] Vik ES, Nawaz MS, Strøm Andersen P, Fladeby C, Bjørås M, Dalhus B, Alseth I. Endonuclease V cleaves at inosines in RNA. Nature Communications (2013) 4: 2271.
[14] Martínez-García E, Goñi-Moreno A, Bartley B, McLaughlin J, Sánchez-Sampedro L, Pascual Del Pozo H, Prieto Hernández C, Marletta AS, De Lucrezia D, Sánchez-Fernández G, Fraile S, de Lorenzo V. SEVA 3.0: an update of the Standard European Vector Architecture for enabling portability of genetic constructs among diverse bacterial hosts. Nucleic Acids Research (2020) 48: D1164–D1170.
| None |