Part:BBa_K2675057
sfGFP-LVAtag expression cassette under the control of J23110 and pAimX(full) promoter of phage phi3T
This part is an sfGFP-LVAtag (BBa_K2675006) expression cassette under the control of a composite promoter: BBa_J23110 followed by th pAimX(full) promoter of phage phi3T (BBa_K2675020).
Usage and Biology
The switch from lytic-to-lysogenic cycle of the Bacillus phage phi3T is based on the expression of a single transcript, AimX [1]. The expression of AimX is controlled by AimR (BBa_K2279000) was described to be a transcriptional regulator of the pAimX promoter in B. subtilis [1]. The exact binding site of AimR to DNA was not precisely described, but it was mapped by ChIPseq in the intergenic region between AimP and AimX genes, a.k.a. the pAimX(full) promoter (BBa_K2675020).
pAimX(full) (BBa_K2675020) and pAimX(short) (BBa_K2675021) sequences, as well as their hybrid versions (BBa_K2675022, BBa_K2675023, BBa_K2675024, BBa_K2675025), proved not to be active promoters in E. coli. However, assuming that AimR retains its binding ability to its specific target sequence in E. coli, we considered converting the pAimX into an operator for regulation of an E. coli promoter. To do this, we needed to place a constitutive E. coli promoter upstream of pAimX and a reporter gene downstream of pAimX. In this design, transcription will be initiated at the upstream constitutive E. coli promoter and, in the absence of AimR, the RNA polymerase should be able to transcribe the pAimX region, the RBS and the reporter sequence up to the terminator. In the presence of AimR, AimR will bind to its specific recognition sequence mapped in pAimX(full) region and thus it should prevent the RNA polymerase from continuing the transcription. The expected results are: - in the absence of AimR: expression of the reporter gene - in the presence of AimR: no expression of the reporter gene Thus, AimR will be converted from an activator in Bacillus to a repressor in E. coli.
We chose the BBa_J23110 constitutive promoter from the Anderson Library that is reported to have a medium activity compared to the other members of this promoter family. A medium strength promoter allows more easily to detect variation in its activity (inhibition or activation) than a low or high strength promoter.
Then we placed the pAimX(full) and pAimX(short) sequences (BBa_K2675020 and BBa_K2675021) between BBa_J23110 and sfGFP-LVAtag (BBa_K2675006), and we thus constructed 2 composite parts: BBa_K2675057 and BBa_K2675058 respectively. During the cloning process, we selected a colony that, upon sequencing, revealed to have a truncated version of the J23110 with only the -10 box (BBa_K2675026) upstream of pAimX(short). As this colony was green under blue light (with amber filter) due to sfGFP expression, we continued characterising this cloning artefact for its interesting promoter activity (BBa_K2675059).
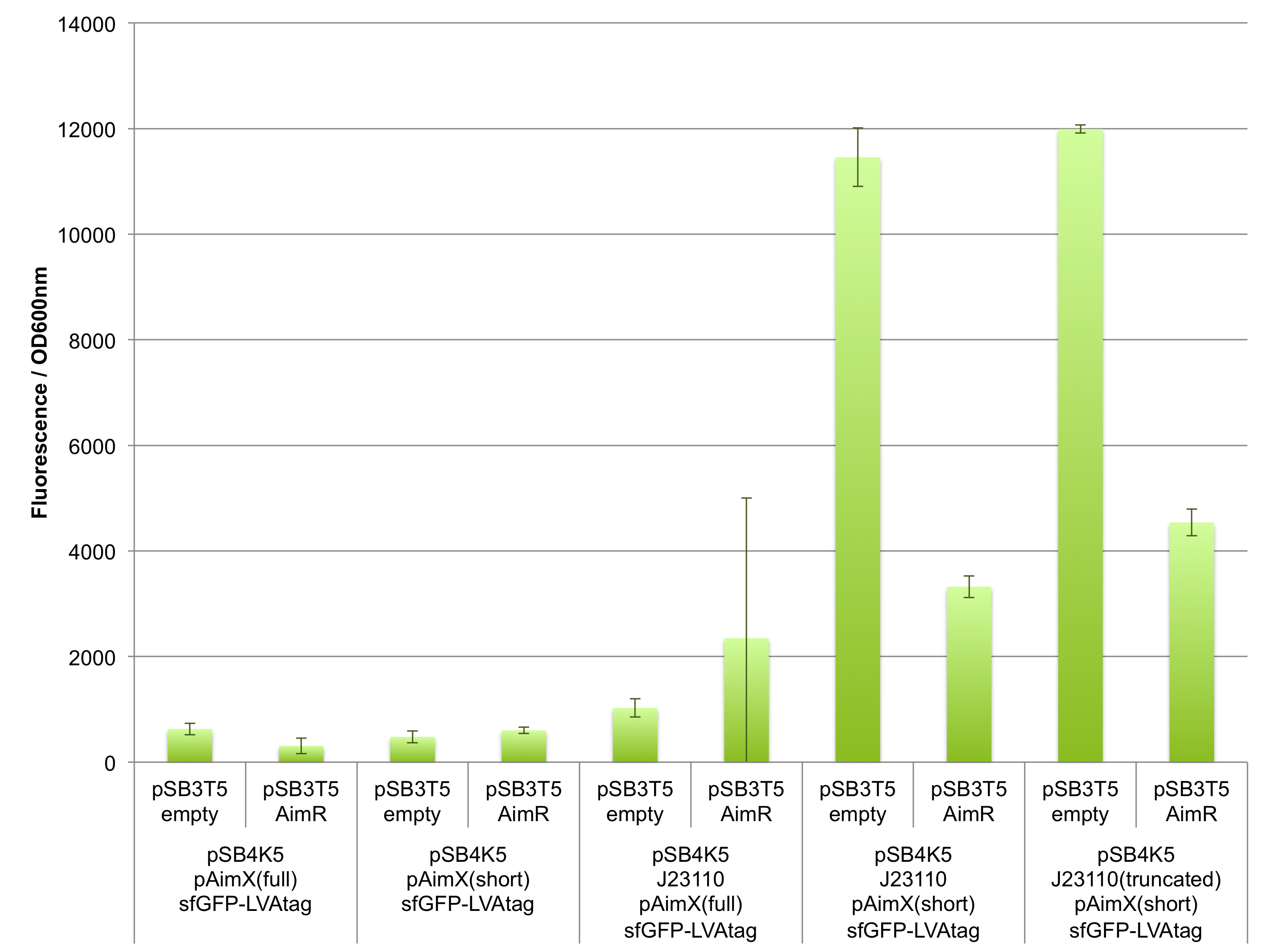
The results presented in Figures 1 and 2 show that the pAimX(short), but not pAimX(full) acts as expected as an operator for J23110 promoter in E. coli. Indeed, we can observe the fluorescent sfGFP expressed when this reporter gene is placed downstream of the composite promoter J23110 + pAimX(short) (BBa_K2675058). Moreover, the fluorescence decreases in the presence of AimR suggesting that AimR acts as predicted for this design: as a repressor. The same trend is observed also when the truncated version of J23110 in present upstream of pAimX(short) (BBa_K2675059).
However, no significant sfGFP expression was observed when the pAimX(full) sequence was inserted between between the J23110 promoter and sfGFP (BBa_K2675059). One plausible explanation is that transcription is initiated from the upstream J23110 promoter, but it ends prematurely at an internal terminator predicted to exist in the pAimX(full) sequence (see arrows marking a hairpin loop in Figure 1). In addition to proving that AimR can act as an effective repressor in E. coli, these experiments help identify the precise site of AimR binding to DNA: the pAimX(short) promoter (BBa_K2675021).
Figure 1: In vivo characterisation of sfGFP expression by E. coli cells harbouring the expression cassettes of sfGFP-LVAtag under the control of pAimX(full) promoter (BBa_K2675050), pAimX(short) promoter (BBa_K2675051), J23110 + pAimX(full) promoter (BBa_K2675057), J23110 + pAimX(short) promoter (BBa_K2675058), J23110 truncated promoter (-10 only) + pAimX(short) promoter (BBa_K2675059) in pSB4K5 backbone. The AimR was expressed as BBa_K2675040 on pSB3T5. An empty pSB3T5 served as negative control. Fluorescence values were normalised by OD600nm.
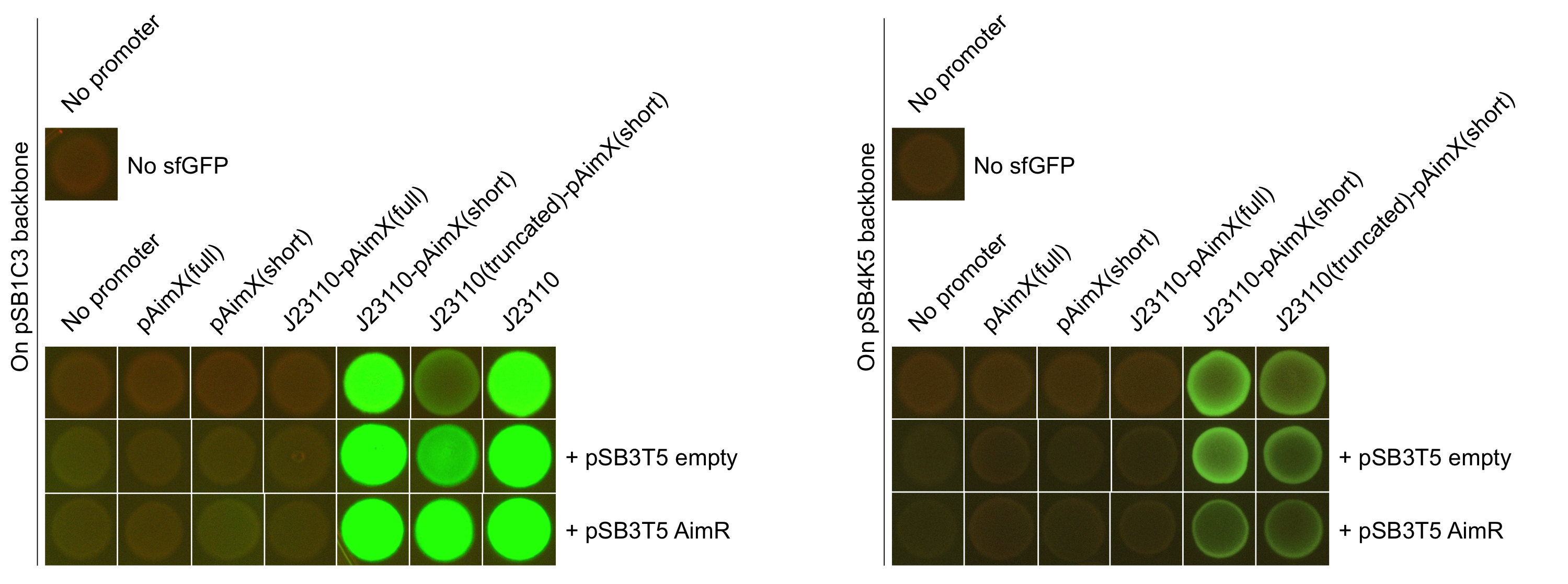
Figure 2: Pictures of E. coli cells harbouring the expression cassettes of sfGFP-LVAtag under the control of pAimX(full) promoter (BBa_K2675050), pAimX(short) promoter (BBa_K2675051), J23110 + pAimX(full) promoter (BBa_K2675057), J23110 + pAimX(short) promoter (BBa_K2675058), J23110 truncated promoter (-10 only) + pAimX(short) promoter (BBa_K2675059) in pSB1C3 or pSB4K5 backbones. The negative control has been performed with an empty pSB1C3 or pSB3K4 and the positive one with BBa_K2675056.
Sequence analysis of the pAimX(full) promoter (BBa_K2675020) revealed the presence of an inverted repeat sequence potentially forming the stem-loop structure of a terminator (Figure 1). Disrupting the predicted terminator lead to sfGFP expression initiated at the upstream J23110 promoter (or further details, visit the BBa_K2675190, BBa_K2675191, BBa_K2675192, BBa_K2675193, BBa_K2675194, BBa_K2675195, BBa_K2675196 and BBa_K2675197 page in the registry).
References
[1] Erez Z, Steinberger-Levy I, Shamir M, Doron S, Stokar-Avihail A, Peleg Y, Melamed S, Leavitt A, Savidor A, Albeck S, Amitai G, Sorek R. Communication between viruses guides lysis-lysogeny decisions. Nature (2017) 541, 488-493.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 227
- 1000COMPATIBLE WITH RFC[1000]
| None |