Part:BBa_K592009
amilCP, blue chromoprotein
This chromoprotein from the coral Acropora millepora, amilCP, naturally exhibits strong color when expressed. The protein has an absorbance maximum at 588 nm giving it a blue/purple color visible to the naked eye, thereby requiring no instruments to observe. The strong color is readily observed in both LB or agar culture, in less than 24 hours of incubation.
This part has been adaptated to B. subtilis with a LVA tail. by the UPMC-Paris Team in 2016. (see part BBa K2180019)
Usage and Biology
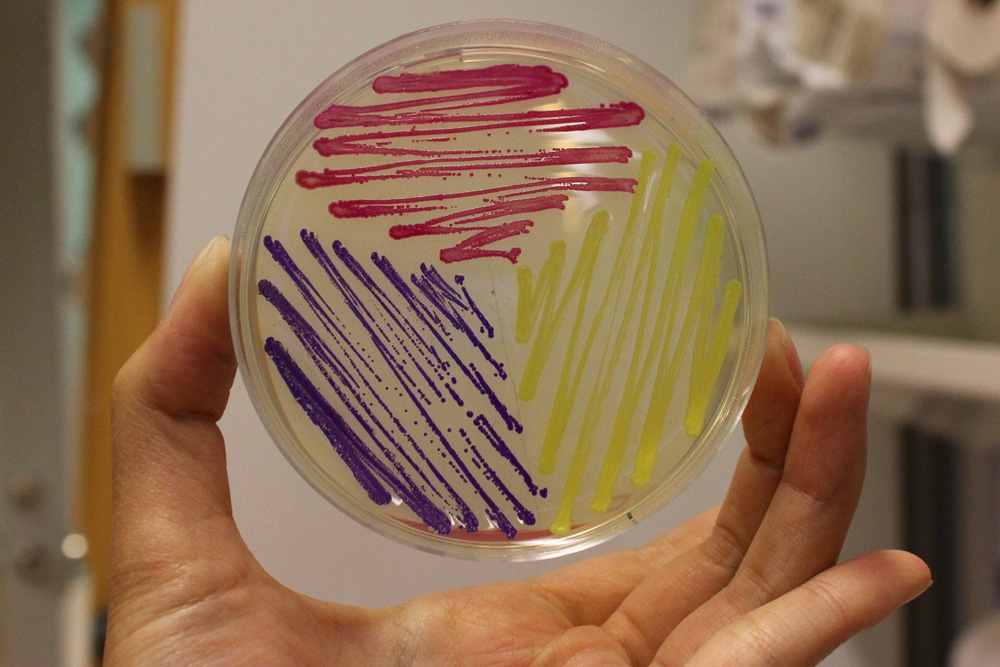
This part is useful as a reporter. In the pictures below it's built into BBa_K592015.
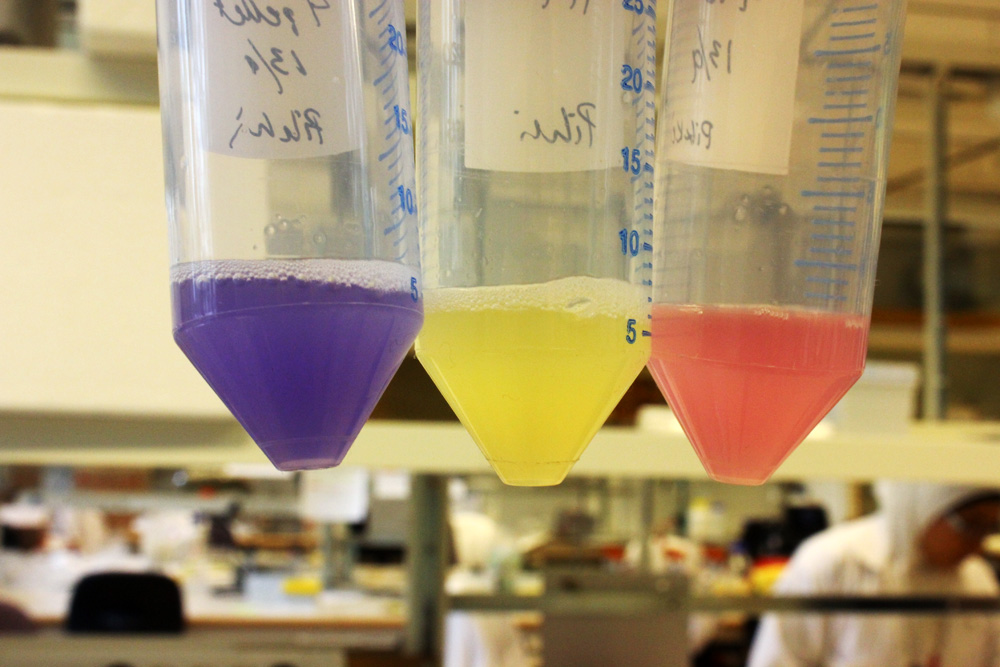
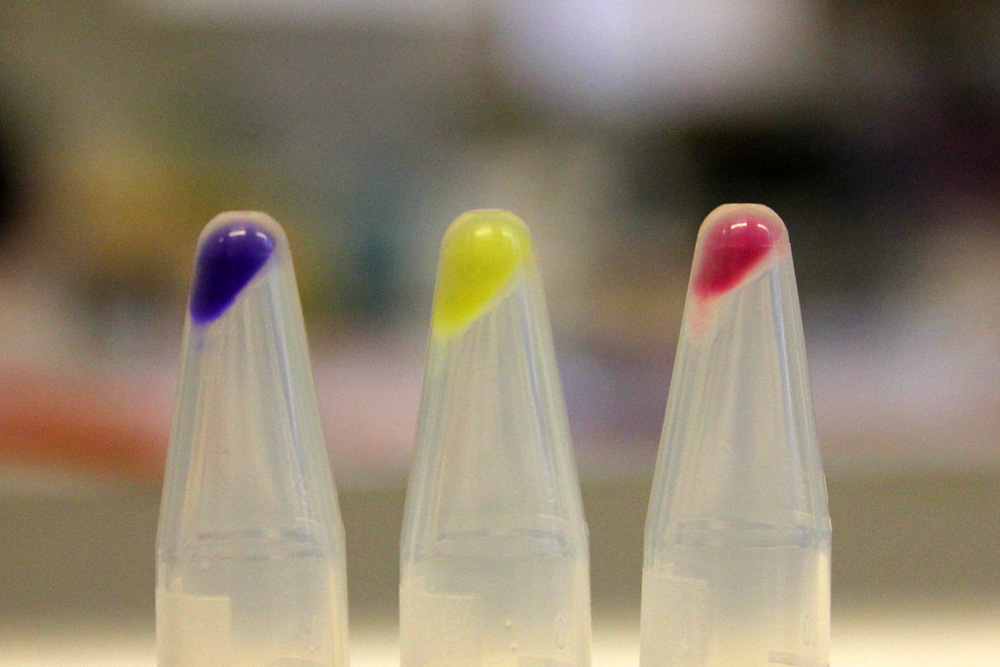
iGEM11_Uppsala-Sweden: Expression of chromoproteins. The images above show E coli constitutively expressing amilCP BBa_K592009 (blue), amilGFP BBa_K592010 (yellow) and RFP BBa_E1010 (red). Also see the green chromoprotein cjBlue BBa_K592011.
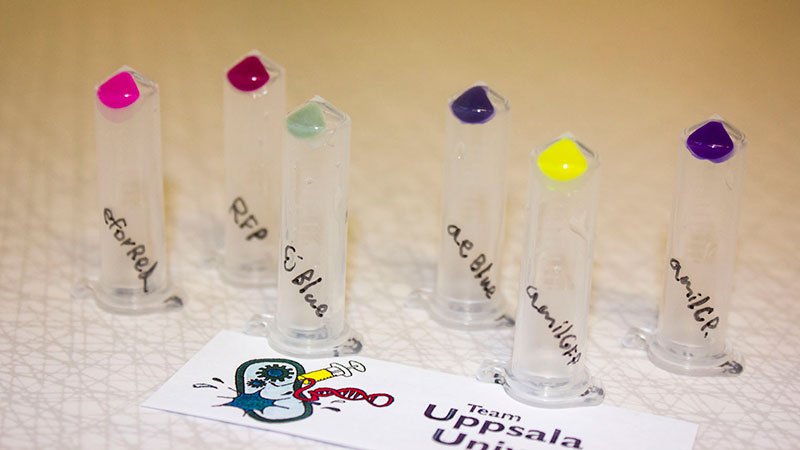
iGEM12_Uppsala_University: The Uppsala chromoprotein collection and RFP. The image shows pellets of E coli expressing chromoproteins eforRed BBa_K592012, RFP BBa_E1010, cjBlue BBa_K592011, aeBlue BBa_K864401, amilGFP BBa_K592010 and amilCP BBa_K592009.
References
[http://www.ncbi.nlm.nih.gov/pubmed/18648549] Alieva, N. O., et al. 2008. Diversity and evolution of coral fluorescent proteins. PLoS One 3:e2680.
[1] Levy, Appelbaum, Leggat, Gothlif, Hayward, Miller, & Hoegh-Guldberg. (2007). Light-responsive cryptochromes from a simple multicellular animal, the coral Acropora millepora. Science (New York, N.Y.), 318(5849), 467-70. GenBank: AY646075.1
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
CONTRIBUTION: Uppsala 2018
Authors iGEM Uppsala 2018, Matilda Brink, Elin Ramstrom.
The original native AmilCP sequence (Part:BBa_K592009) was codon optimized for E.coli K12 with a codon optimization tool provided by Integrated DNA Technologies (IDT). The codon optimized basic AmilCP part is attached to a constitutive promotor, RBS and a double terminator; Part:BBa_K2669003.
Purpose
The Uppsala 2018 team have performed a codon optimization on the original native AmilCP basic part in order to get a blue protein expression that can be maintained for a longer period of time through several generations of growth, hence making the expression of the blue chromoprotein more stable.
Method
In order to conduct a stability assay through liquid experiment, the codon optimized basic part and the original native AmilCP part was attached to a constitutive promotor, RBS and a double terminator separately. The two Biobrick parts were ordered as customized genes from IDT inserted in pUCIDT (Amp).
Both vectors were transformed into DH5-aplha E.coli cells through single tube transformation and yielded phenotypically blue colonies for cells containing plasmid with original part and plasmid containing codon optimized part. Stability assay through liquid experiemt was performed according to protocol [2].
Results

Figure 3: Results after 10 generations of growth. The upper row represents 10 different culture pellets of the cells with the plasmid containing the codon optimized sequence of AmilCP. The lower row represents 10 different culture pellets with the original native AmilCP (part:BBa_K592009) sequence incorporated in the plasmid.
Already after 10 generations of growth it was clearly visible that the color intensity was better distributed in the cells containing the plasmid with the codon optimized AmilCP sequence. The result confirms that the plasmid containing codon optimized part generates better maintenance of protein expression than the plasmid containing the original native part. The improved stability of expression makes this Biobrick part a good candidate as a reliable reporter or biosensor.
IMPROVEMENT REFERENCE: IONIS_PARIS 2017
The part used to improve BBa_K592009 (Group: iGEM11_Uppsala-Sweden) is : BBa_K2282006 (Group: iGEM17_IONIS-PARIS). https://parts.igem.org/Part:BBa_K2282006
IMPROVEMENT REFERENCE: Austin_UTexas 2017
The original blue chromoprotein part (BBa_K592009) was codon-optimized in order to increase its overall stability as a color reporter. The modified blue chromoprotein part is: BBa_K2253002 (Group: iGEM17_Austin_UTexas). Designed by Surta Dave.
IMPROVEMENT REFERENCE: DTU-Denmark 2017
A 6x his-tag was added to this part, to the C-terminal end of the expressed amilCP. The new part was submitted as BBa_K2355002 (Group: iGEM17_DTU-Denmark), Designed by:Philip Sørensen.
IMPROVEMENT REFERENCE: BNU-China 2017
The part was added to the end of a silk protein from honeybee. The new part is : BBa_K2220028(Group: iGEM17_BNU-China), Designed by: Jiawei Xing.
IMPROVEMENT REFERENCE: IISER Pune India 2017
The part was assembled with B0034 RBS and B0015 double terminator. The new part is : BBa_K2258003(Group: iGEM17_IISER Pune India), Designed by: Jyothish S.
IMPROVEMENT REFERENCE: Peshawar 2017
One of the most used chromoprotein used in iGEM is AmilCP blue, BBa_K592009. Because our modelling team was doing homology models for other proteins and AmilCP doesn't have a homology model on it's page, we decided to do one for AmilCP blue.
Click <a href="http://2017.igem.org/Team:Peshawar/Model" target="_blank"> here</a>to check out our modeling page!
Contribution: Valencia UPV iGEM 2018
Authors: Carolina Ropero, Adrián Requena Gutiérrez Part BBa_K2656018 is the amil Chromoprotein coding sequence BBa_K592009 compatible with both BioBricks and [http://2018.igem.org/Team:Valencia_UPV/GB3 Golden Braid 3.0] assembly methods.
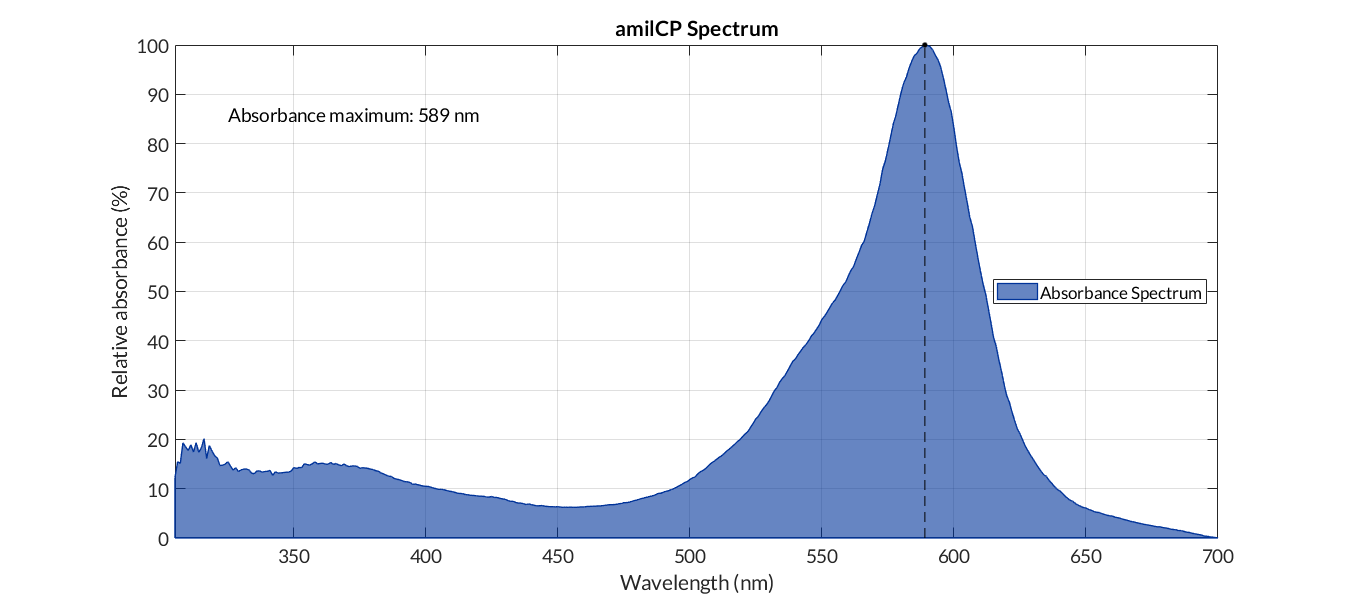
In order to carry out a correct characterization of the protein and to be able to use it to make measurements of the different transcriptional units that we have assembled with it, we obtained the absorbance spectrum in the conditions of our equipment. To do so, we assembled composite part BBa_K2656113. By using this [http://2018.igem.org/Team:Valencia_UPV/Modeling#spectra protocol] with the parameters of Table 1, Figure 1 was obtained.
| Parameter | Value | ||
| Number of samples | 3 | ||
| Wavelength measurement range (nm) | [300-700] | ||
| Table 1. Parameters used to obtain the spectum | |||
IMPROVEMENT REFERENCE: NWU-China 2019
The N-terminus of this protein was added with a secreted peptide (Kp-Sp,BBa_K2981006). The new part is : BBa_K2981007(Group: iGEM19_NWU-China), Designed by: Junjie Zhu.
Contribution: Duesseldorf 2019
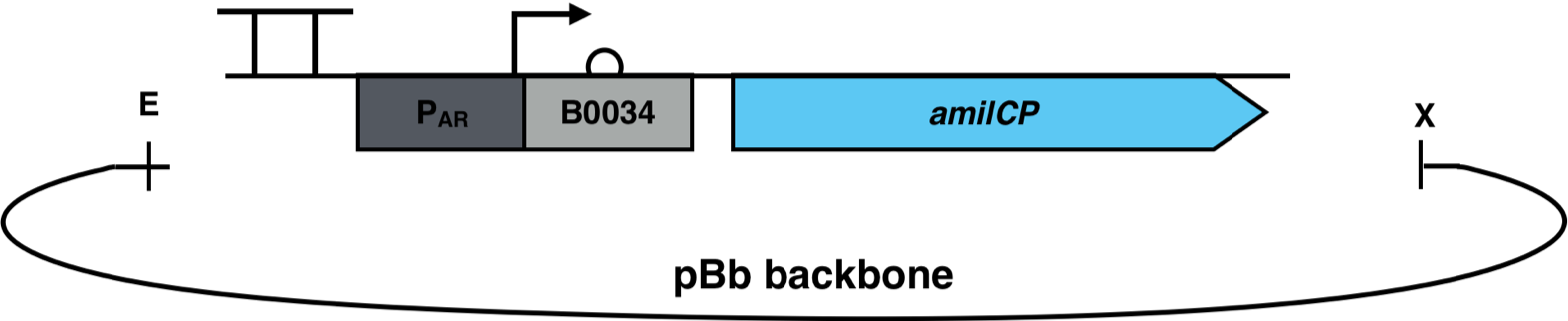
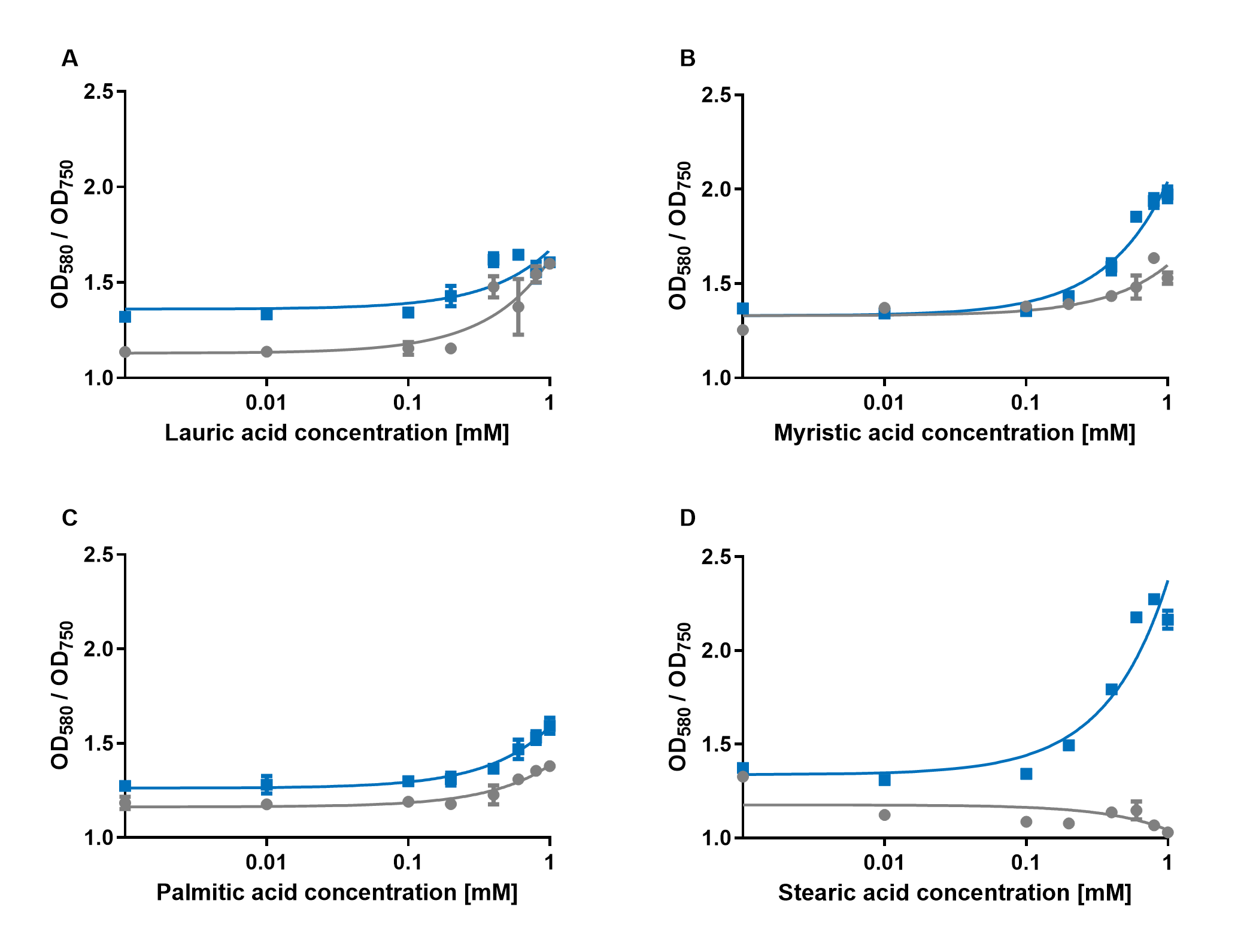
iGEM Duesseldorf characterized this protein in its capacity to be a biosensor molecule for fatty acids. It was assembled into the composite part BBa_K2924020 and tested with several fatty acids, at several concentrations.
The experiments with amilCP showed that the production of the chromoprotein is increased by a higher concentration of fatty acids in the medium. The best result was achieved with the fatty acid stearic acid. The biosensor also worked for the chain lengths from C14:0 and C16:0. The EVC is in these experiments lower than the samples with amilCP. By adding lauric acid (C12:0) to the culture medium the production of amilCP increased, but the EVC also increased a lot and is close to amilCP, so it is not certain if this result can be used for an nearly distinct conclusion.
In conclusion, amilCP can be used as a biosensor molecule for measurements by absorption, if no fluorescence plate reader is available in the lab. However, for the use as a biosensor, the results have to be confirmed by using controls with empty vector strains and control for absorption of the molecule that is to be detected in the sample.
iGEM19_SCU-China:
The yamilCP(Part: BBa_K3221205) was improved from amilCP, which is a blue chromoprotein gene from Acropora millepora submitted by Team Uppsala-Sweden 2011.
We expected amilCP to be expressed efficiently in S. cerevisiae, so we chose amilCP to be improved. We processed codon optimization on original amilCP sequence and added a consensus sequence (AAAAAA) for S. cerevisiae expression. And the new sequence was named yamilCP.
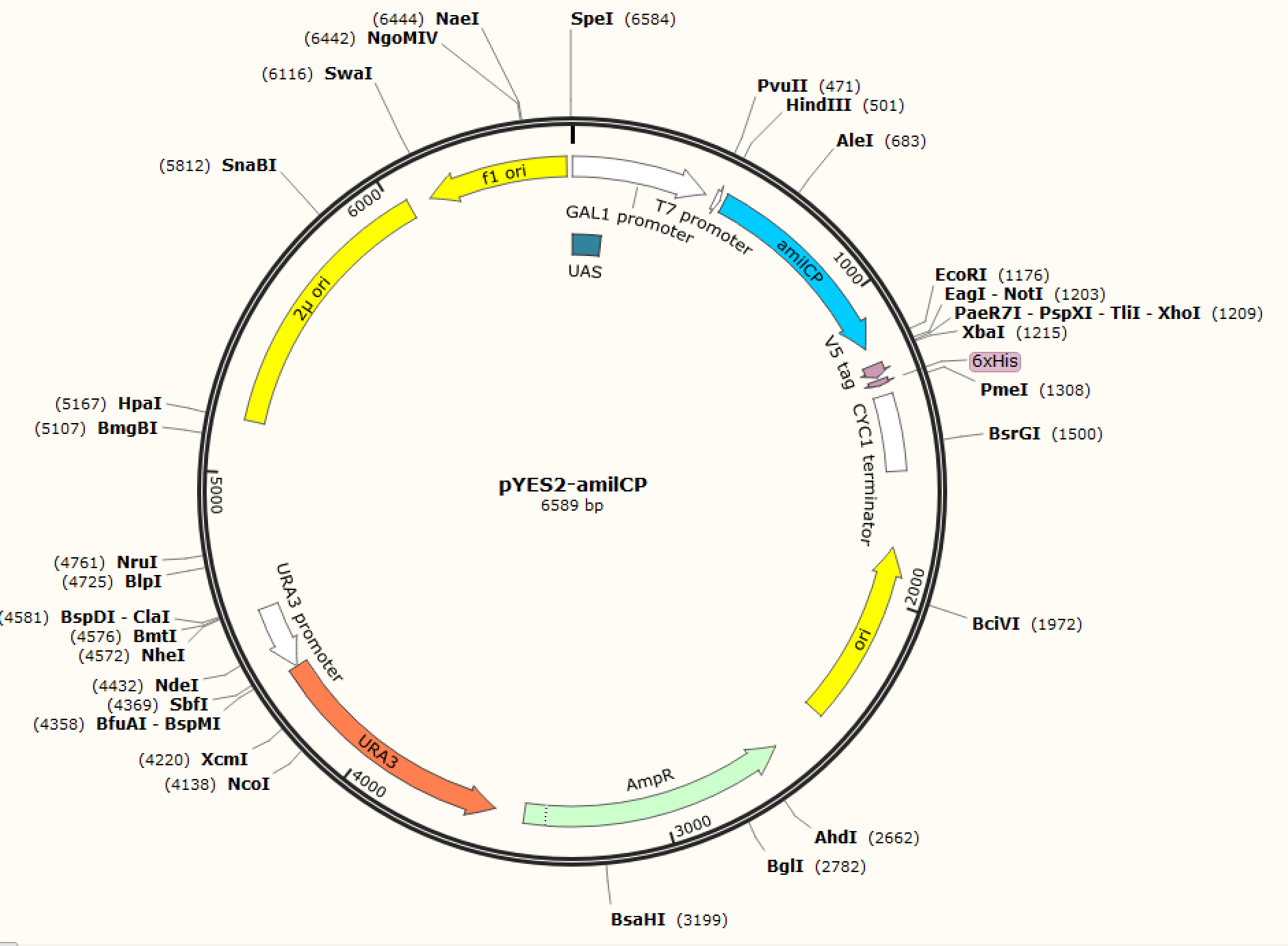
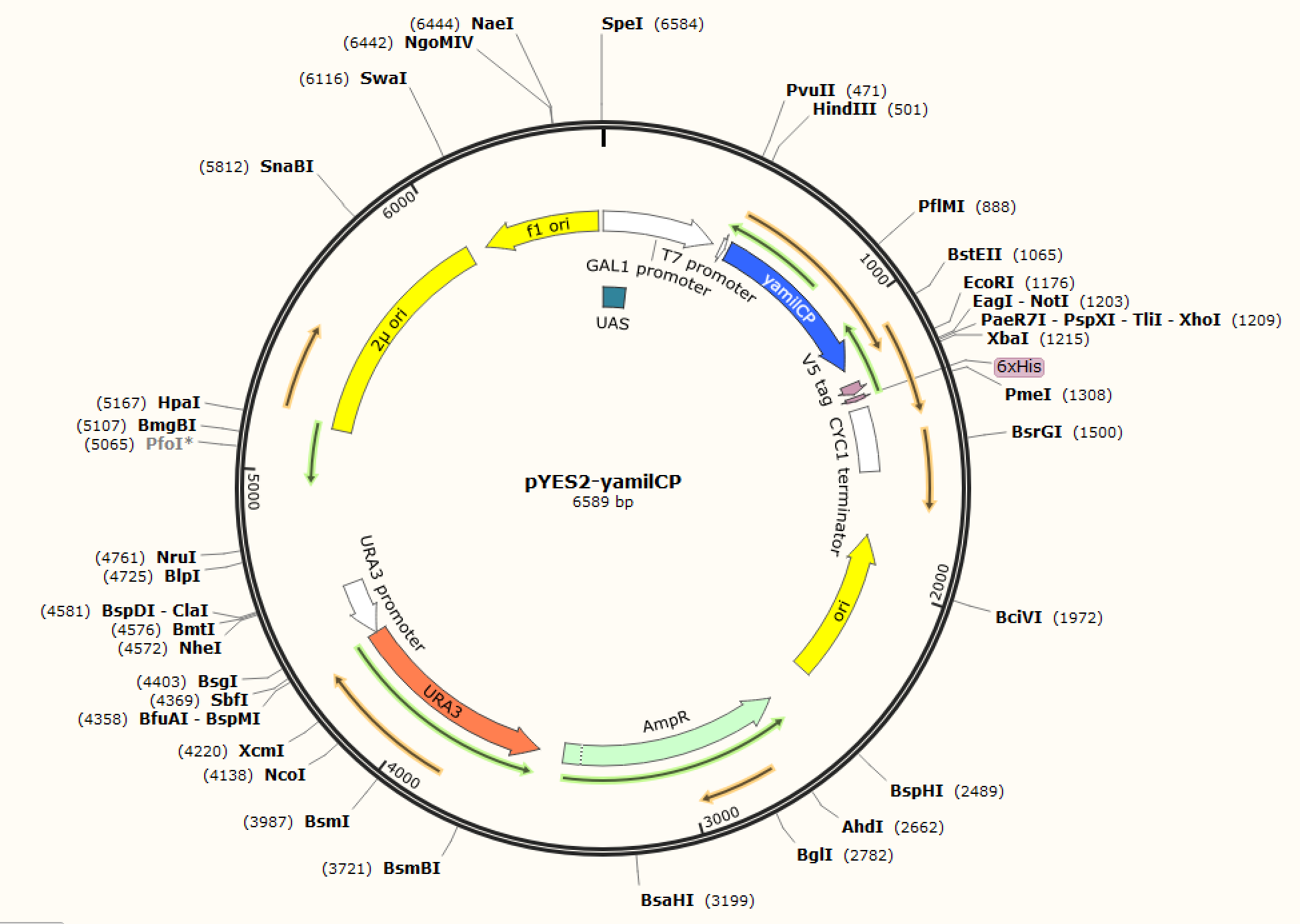
Figure. 1 The map of plasmid pYES2-amilCP Figure. 2 The map of plasmid pYES2-yamilCP
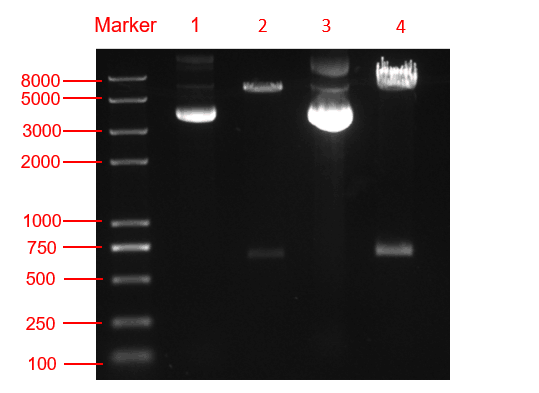
We respectively cloned the amilCP and yamilCP on the vector of pYES2-NTA.There are the maps of pYES2-amilCP and pYES2-yamilCP (Figure.1 and Figure.2). And we verified the construction of them by double restriction enzyme digestion(Figure.3) and sequencing.
Figure. 3 The double restriction enzyme digestion verification result. lane 1: pYES2-amilCP plasmid, lane 2: pYES2-amilCP plasmid digestion, lane 3: pYES2-yamilCP plasmid, lane4: pYES2-amilCP plasmid digestion, Marker:Trans 2K plus Ⅱ Marker.
Then we transformed pYES2-amilCP and pYES2-yamilCP into S. cerevisiae respectively. Later on, We observed the expression of blue chromoprotein with the induction of galactose (Figure.4).
Figure. 4 The colony transformed pYES2-amilCP and pYES2-yamilCP expressed blue chromoprotein.
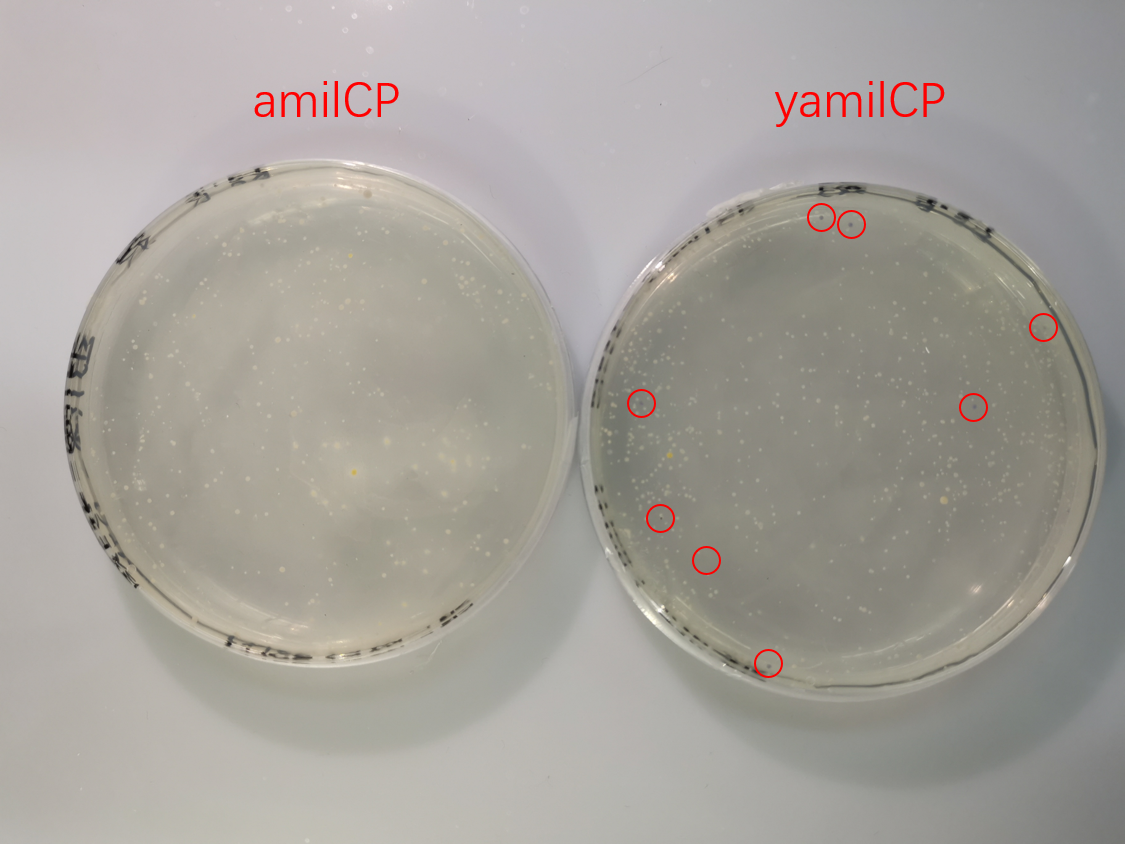
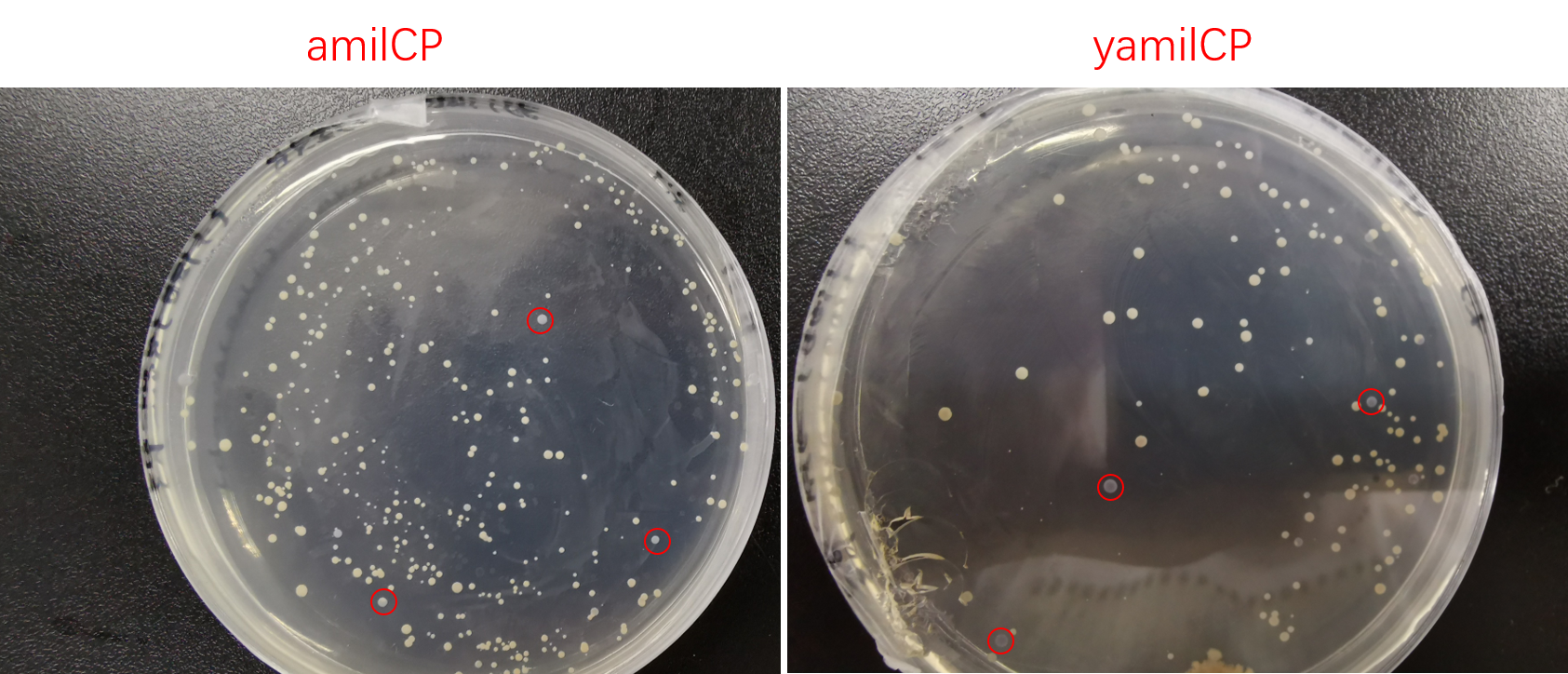
To test the expression effciency of pYES2-amilCP and pYES2-yamilCP in S. cerevisiae, we transformed them into BY4741 and used galactose to induce blue chromoprotein expression. After 10 days, we observed blue colonies on the pYES2-yamilCP plate at first and there was no blue colony on the pYES2-amilCP plate, which confirmed that yamilCP expressed more quickly than amilCP (Figure.5). As Figure. 5, there is no blue clony (0/356) on the amilCP plate while 8 colonies (8/575) on the yamilCP plate. It shows that our improvement is successful.
As the result, the improved yamilCP was successfully expressed, and the expression of yamilCP is more quickly than that of amilCP in S. cerevisiae.
Figure. 5 The transformation results of pYES2-amilCP and pYES2-yamilCP into S. cerevisiae BY4741 with galactose induction after 10 days.
//collections/chromoprotein/uppsala
//function/reporter/pigment
| abs | 588 nm |
| color | Blue |