RBS
Part:BBa_B0030
Designed by: Vinay S Mahajan, Voichita D. Marinescu, Brian Chow, Alexander D Wissner-Gross and Peter Carr IAP, 2003. Group: Antiquity (2003-01-31)
RBS.1 (strong) -- modified from R. Weiss
Strong RBS based on Ron Weiss thesis. Strength is considered relative to BBa_B0031, BBa_B0032, BBa_B0033 and BBa_B0034.
Usage and Biology
IIT Madras 2016's Characterization
Modelling
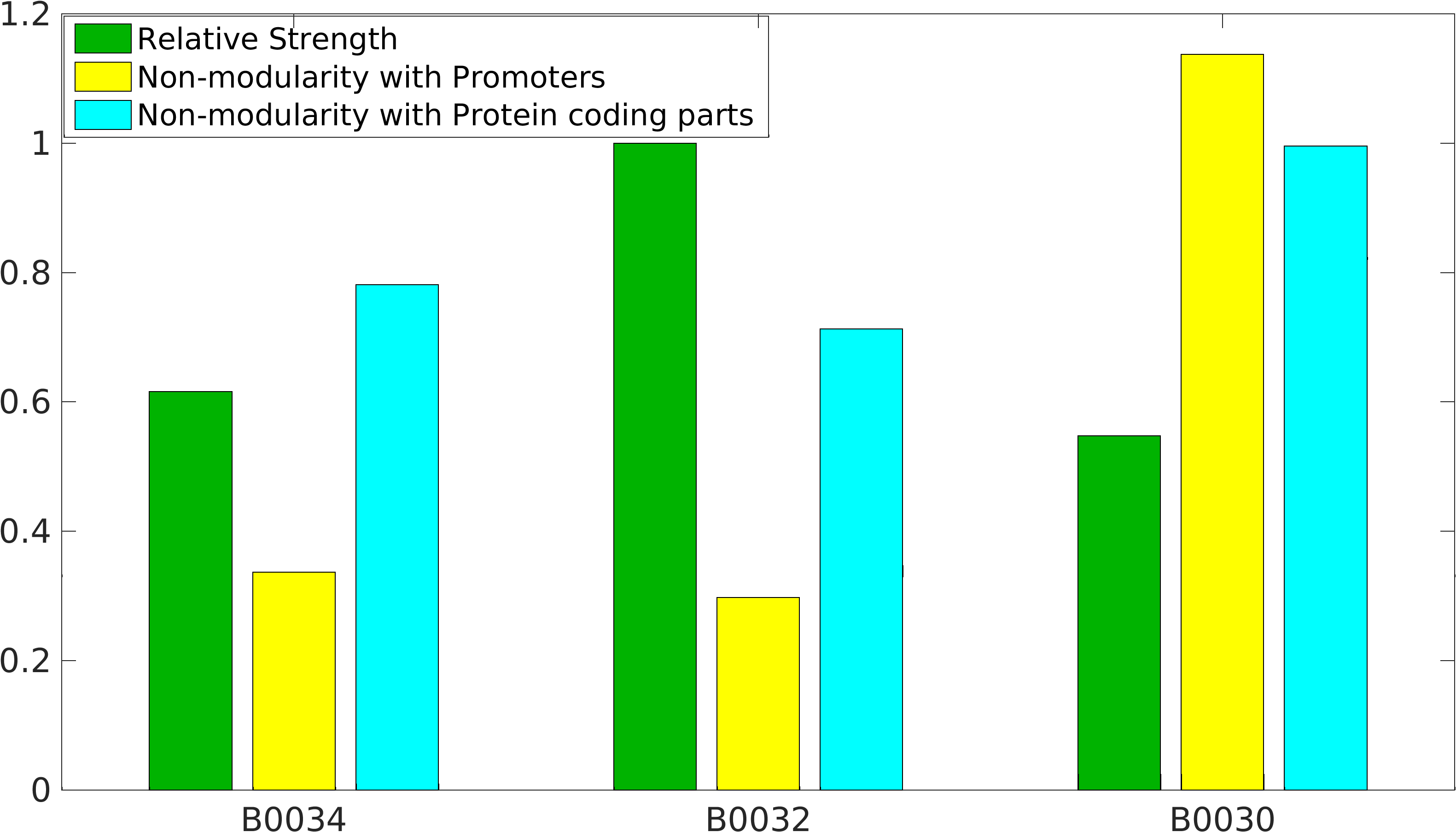
Global non-modularity towards promoters & protein coding parts and relative strength was estimated for RBSs B0030, B0032, B0034 in our [http://2016.igem.org/Team:IIT-Madras/Model#Modularity_of_RBS_parts modelling]
Experimentation
Biobrick RBSs B0030, B0031, B0032, B0034 were used in our 'Noise in Device' experiment to understand the role of RBS parts in giving rise to noise.
Note: not compatible with R0053 promoter due to likely transcript secondary structure This combination yielded very low gfp expression (see BBa_I7108). [jcb 8/3/05].
Sequence and Features
Assembly Compatibility:
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Team Warsaw 2010's measurement
RBS strength (relative to B0034): 91,84%Functional Parameters
| biology | -NA- |
| efficiency | 0.6 |
[edit]
Categories
Parameters
//rbs/prokaryote/constitutive/community
//ribosome/prokaryote/ecoli
//chassis/prokaryote/ecoli
//direction/forward
//regulation/constitutive
//ribosome/prokaryote/ecoli
//chassis/prokaryote/ecoli
//direction/forward
//regulation/constitutive
| biology | |
| efficiency | 0.6 |

 1 Registry Star
1 Registry Star