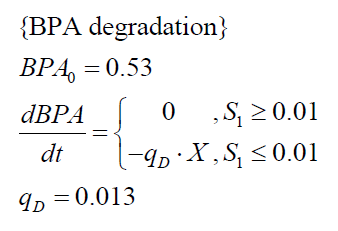Part:BBa_K525515
Fusion protein of BisdA and BisdB
Fusion protein of BisdA and BisdB for improved bisphenol A degradation with Escherichia coli.
Usage and Biology
Expressing this BioBrick in E. coli enables the bacterium to degrade the endocrine disruptor bisphenol A (BPA). The fusion protein works better than a polycistronic expression of the two BioBricks bisdA (BBa_K123000) and bisdB (BBa_K123001).
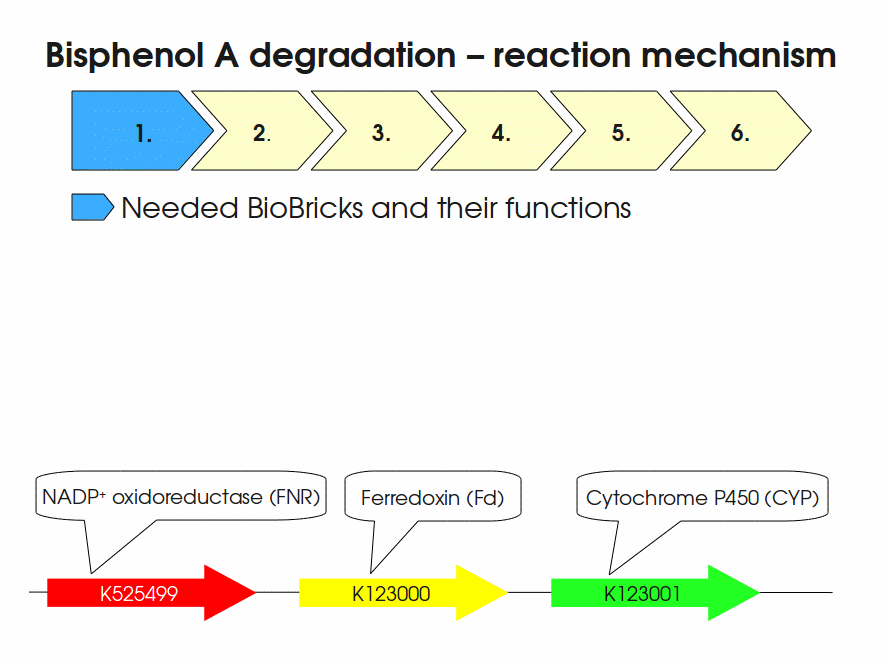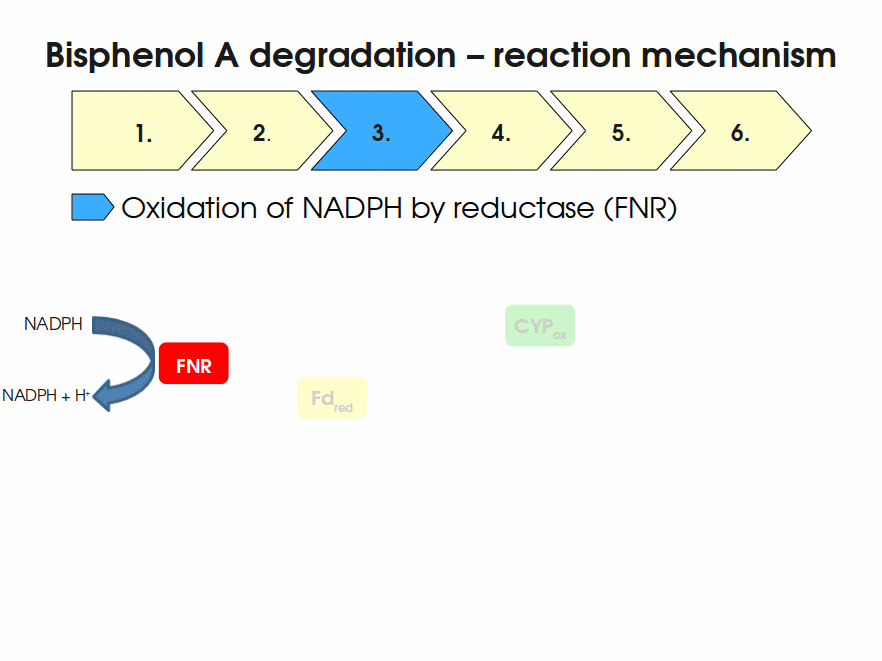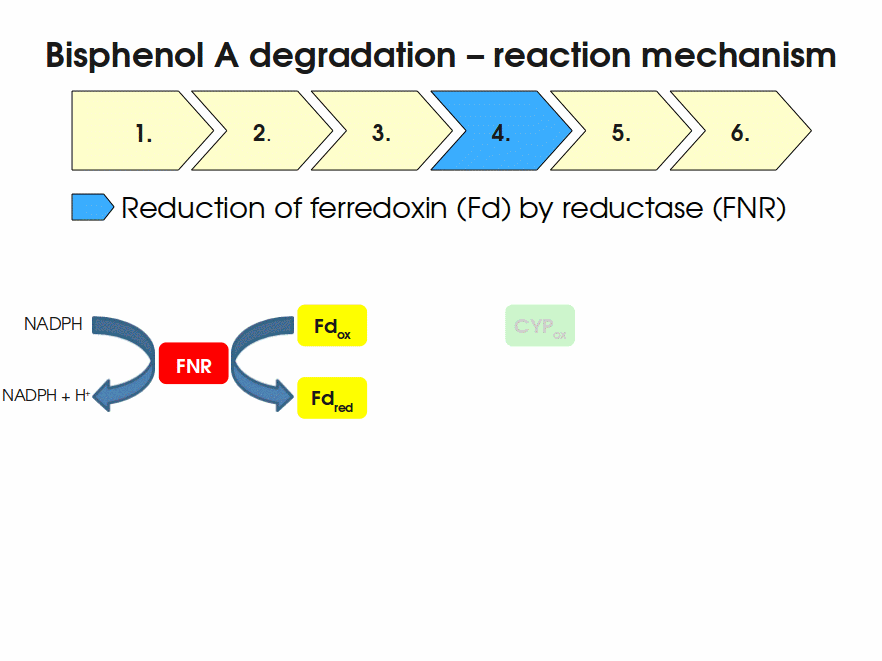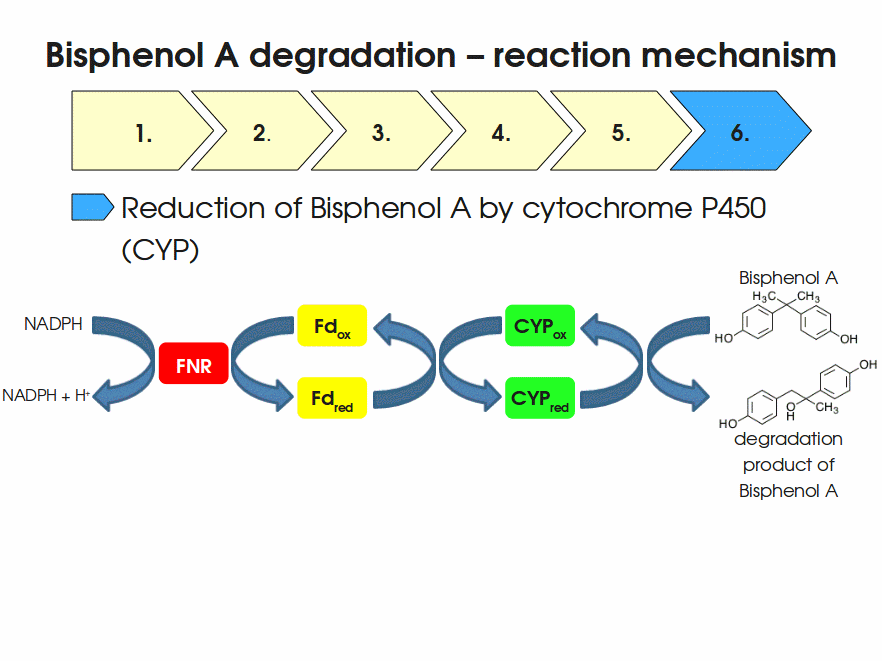
BPA is mainly hydroxylated into the products 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol. In S. bisphenolicum AO1, a total of three genes are responsible for this BPA hydroxylation: a cytochrome P450 (CYP, bisdB), a ferredoxin (Fd, bisdA) and a ferredoxin-NAD+ oxidoreductase (FNR) Sasaki05a. The three gene products act together to reduce BPA while oxidizing NADH + H+. The cytochrome P450 (BisdB) reduces the BPA and is oxidized during this reaction. BisdB in its oxidized status is reduced by the ferredoxin (BisdA) so it can reduce BPA again. The oxidized BisdA is reduced by a ferredoxin-NAD+ oxidoreductase consuming NADH + H+ so the BPA degradation can continue Sasaki05a. This electron transport chain between the three enzymes involved in BPA degradation and the BioBricks needed to enable this reaction in vivo and in vitro are shown in the following figure (please have some patience, it's an animated .gif file):
This BioBrick was characterized using BBa_K525517.
- Allergen characterization of BBa_K525515: Not a potential allergen
The Baltimore Biocrew 2017 team discovered that proteins generated through biobrick parts can be evaluated for allergenicity. This information is important to the people using these parts in the lab, as well as when considering using the protein for mass production, or using in the environment. The allergenicity test permits a comparison between the sequences of the biobrick parts and the identified allergen proteins enlisted in a data base.The higher the similarity between the biobricks and the proteins, the more likely the biobrick is allergenic cross-reactive. In the full-length alignments by FASTA, 30% or more amount of similarity signifies that the biobrick has a Precaution Status meaning there is a potential risk with using the part. A 50% or more amount of identity signifies that the biobrick has a Possible Allergen Status. In the sliding window of 80 amino acid segments, greater than 35% signifies similarity to allergens. The percentage of similarity implies the potential of harm biobricks’ potential negative impact to exposed populations. For more information on how to assess your own biobrick part please see the “Allergenicity Testing Protocol” in the following page http://2017.igem.org/Team:Baltimore_Bio-Crew/Experiments
For the biobrick Part:BBa_K525515, there was a 28.2% of identity match and 52.1% similarity match to the top allergen in the allergen database, glutathione transferase mu class Yv from Sarcoptes scabiei. This means that the biobrick part is not of potential allergen status. In 80 amino acid alignments by FASTA window, no matches found that are greater than 35% for this biobrick. This also means that there is not of potential allergen status.
Important parameters
Tab. 1: Important parameters of BBa_K525515, determined with BBa_K525517.
| Experiment | Characteristic | Result |
|---|---|---|
| Expression in E. coli | Compatibility | E. coli KRX, TOP10, MACH1, BL21(DE3) |
| Expression | Constitutive | |
| Optimal temperature | 30 °C | |
| BPA working concentration | 120 mg L-1 (0.53 mM) | |
| Purification | Molecular weight | 59.3 kDa |
| Theoretical pI | 4.99 | |
| High absorbtion | 450 nm (due to CYP) | |
| Degradation of BPA | Completely degradation of 0.53 mM BPA | 21 - 24 h |
| Specific BPA degradation rate | 1.29 10-10 mM cell-1 |
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 603
Illegal BamHI site found at 1341 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 4
Illegal AgeI site found at 1594 - 1000COMPATIBLE WITH RFC[1000]
Bisphenol A degradation with E. coli
The bisphenol A degradation with the BioBricks BBa_K123000 and BBa_K123001 works in E. coli KRX in general. Because [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full Sasaki et al. (2008)] reported problems with protein folding in E. coli which seem to avoid a complete BPA degradation, we did not cultivate at 37 °C and we did not use the strong T7 promoter as [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full Sasaki et al. (2008)] did for expressing these BioBricks but we cultivated at 30 °C and we used a medium strong constitutive promoter (BBa_J23110). 30 °C is in addition the cultivation temperature of S. bisphenolicum AO1. With this promoter upstream of a polycistronic bisdAB gene we were able to completely degrade 120 mg L-1 BPA in about 30 - 33 h. By fusing BBa_K123000 and BBa_K123001 together (using Freiburg BioBrick assembly standard) we could improve the BPA degradation of E. coli even further, so 120 mg L-1 BPA can be degraded in 21 - 24 h. This data is shown in the following figure:
We also carried out these cultivations at different temperatures and BPA concentrations, but the chosen conditions (30 °C and 120 mg L-1 BPA) seem to be the best. Higher BPA concentrations have an effect on the growth of E. coli and higher temperature leeds to a worse BPA degradation (probably due to misfolding of the enzymes). Lower temperature also leeds to less BPA degradation (probably due to slower growth, expression and reaction rate at lower temperatures). These data on the effect of the temperature on the BPA degradation is shown in figure 3.
As shown by [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full Sasaki et al. (2008)], BisdB expressed in E. coli leeds to hardly no BPA degradation. In our experiments we could not detect the BPA degradation products 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol in cultivations with E. coli expressing BBa_K123000 or BBa_K123001 alone (neither via UV- nor MS-detection). The BPA degradation products 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol were identified via MS-MS (m/z: 243 / 225 / 211 / 135) and only occured in cultivations with E. coli expressing BisdA and BisdB together. [http://www.springerlink.com/content/q7864l02734wg32m/ Sasaki et al. (2005)] reported the same MS-MS results for 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol when degrading BPA with S. bisphenolicum AO1 as we observed in our BPA degradation experiments.
We could also identify the BPA degradation products when working with E. coli TOP10 and MACH1 (data not shown) but because we want to fuse BisdA and BisdB to S-layer proteins which we express in E. coli KRX the characterizations of BBa_K123000 and BBa_K123001 were carried out in this strain.
Specificity of bisphenol A degradation with E. coli
In order to access the specificity of the bisphenol A degradation by the bisdA | bisdB fusion protein we tested how well two similar bisphenols, bisphenol F (BPF) and bisphenol S (BPS), were employed. The structure of those bisphenols differs only in the chemical group linking the two phenols from that of bisphenol A (see Figure 4).
BPF and BPS are used in a broad range of applications that involve the use of polycarbonates or epoxy resins and thus can often be found were BPA is also present. Accordingly their presence is a potential disruptive factor that could lead to a false positive signal with our biosensor. This is especially true for BPS that in some cases is used as a substitute for BPA in baby bottles [http://www.nytimes.com/2011/04/18/business/global/18iht-rbog-plastic-18.html]. Studies concerning the environmental pollution with BPF ([http://www.sciencedirect.com/science/article/pii/S0043135401003670 Fromme et al. (2002)]) and the acute toxicity, mutagenicity and estrogenicity of BPF and BPS ([http://onlinelibrary.wiley.com/doi/10.1002/tox.10035/abstract Chen et al. (2001)] and [http://toxsci.oxfordjournals.org/content/84/2/249 Kitamura et al. (2005)]) indicate their potential harmfulness but further research is needed to fully access their impact on human health.
E. coli KRX carrying genes for the bisdA | bisdB fusion protein behind the medium strong constitutive promoter BBa_J23110 with RBS BBa_B0034 was cultivated at 30 °C for 36 h with LB-Medium containing 120 mg L-1 BPA, BPF respectively BPS. The BPF and BPS concentration where determined with a HPLC using the same method as with BPA. Figure 5 shows the degradation of the respective bisphenol after 24 h of cultivation in percent.

The results of the experiment indicate that the bisdA | bisdB fusion protein has a high specificity for the degradation of BPA. In addition it is possible that the decrease in BPF and BPS concentration is due to internalization of those bisphenols or a endogenous enzyme of E. coli KRX and not the bisdA | bisdB fusion protein was responsible. It can be assumed that false positive signals because of BPF or BPS present in a sample are unlikely.
Modelling of intracellular bisphenol A degradation
The modelling was done with the software [http://www.berkeleymadonna.com/ Berkeley Madonna] using the [http://en.wikipedia.org/wiki/Runge–Kutta_methods#Common_fourth-order_Runge.E2.80.93Kutta_method common fourth-order Runge-Kutta] method to solve the equations. The model was fitted to the measured data shown above by the function "curve fit" in Berkeley Madonna to calculate the parameters, constants etc.
To model the BPA degradation by E. coli carrying BioBricks for BPA degradation (BBa_K123000 and BBa_K123001) the cell growth has to be described first to calculate a specific BPA degradation rate per cell. The observed growth of E. coli on (our) LB medium was [http://en.wikipedia.org/wiki/Diauxie diauxic] with two different growth phases. Cell growth is a [http://en.wikipedia.org/wiki/First_order_kinetics#First-order_reactions first-order reaction] and is mathematically described as
with the specific growth rate µ and the cell count X. The specific growth rate is dependent on the concentration of the growth limiting substrate (e.g. glucose) and can be described as
with the substrate concentration S, the Monod constant KS and the maximal specific growth rate µmax ([http://www.annualreviews.org/doi/abs/10.1146/annurev.mi.03.100149.002103 Monod, 1949]). Because LB medium is a complex medium we cannot measure the substrate concentration so we have to assume an imaginary substrate concentration. Due to the diauxic growth two different substrates with different Monod constants and consumption rates are necessary to model the cell growth. The amount of a substrate S can be modelled as follows
with the specific substrate consumption rate per cell qS. The whole model for the diauxic growth of E. coli on LB medium with two not measurable (imaginary) substrates looks like:
The specific BPA degradation rate per cell qD is modelled with an equation like eq. (3). In the beginning of the cultivations, when E. coli growths on the "good" imaginary substrate S1, no BPA degradation is observed. When this substrate is consumed, the BPA degradation starts. The model for this diauxic behavior is as follows:
Fig. 6 shows a comparison between modelled and measured data for cultivations with BPA degrading E. coli. In Tab. 2 the parameters for the model are given, obtained by curve fitting the model to the data.

Tab. 2: Parameters of the model.
| Parameter | BBa_K525512 | BBa_K525517 |
|---|---|---|
| X0 | 0.112 108 mL-1 | 0.138 108 mL-1 |
| µmax | 1.253 h-1 | 1.357 h-1 |
| KS,1 | 2.646 AU-1 | 1.92 AU-1 |
| KS,2 | 265.1 AU-1 | 103.1 AU-1 |
| S1,0 | 1.688 AU | 1.166 AU |
| qS,1 | 0.478 AU 10-8 cell-1 | 0.319 AU 10-8 cell-1 |
| S2,0 | 16.091 AU | 6.574 AU |
| qS,2 | 0.295 AU 10-8 cell-1 | 0.191 AU 10-8 cell-1 |
| BPA0 | 0.53 mM | 0.53 mM |
| qD | 8.76 10-11 mM cell-1 | 1.29 10-10 mM cell-1 |
The specific BPA degradation rate per cell qD is about 50 % higher when using the fusion protein compared to the polycistronic bisdAB gene. This results in an average 9 hours faster, complete BPA degradation by E. coli carrying BBa_K525517 compared to BBa_K525512 as observed during our cultivations. The fusion protein between BisdA and BisdB improves the BPA degradation by E. coli.
Interpretation of the results
Misfolding seems to be a problem when expressing BisdA and BisdB in E. coli. To reduce this, the cultivation conditions were improved for the BPA degradation with the polycistronic bisdAB gene in E. coli first, compared to the literature ([http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full Sasaki et al., 2008]). Problems with misfolding of BisdA and BisdB could be reduced by lowering the temperature and growth rate and by using a weaker promoter for expression.
When degrading BPA with E. coli using the BisdA | BisdB fusion protein, both domains (BisdA and BisdB) are active and correctly folded because otherwise there would be no BPA degradation measured and no BPA degradation products 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol could be detected, which could definetely be identified via MS-MS. The about 50 % higher specific BPA degradation rate in E. coli expressing BisdA | BisdB fusion protein could be explained either by improved folding properties of the fusion protein or by the closer distance of BisdA and BisdB in the fusion protein leeding to a faster electron transfer and therefore a more efficient reaction. In this case, which has to be further analysed by cell-free enzyme assays, a natural cytochrome P450 class I electron transport system was converted into a more effective class V electron transport system, demonstrating the new possibilities of synthetic biology. In addition, this new class V system would work without alanine-rich linker, potentially changing the view on cytochrome P450 depending electron transport chains.
Methods
Cultivations

- Chassis: Promega's [http://www.promega.com/products/cloning-and-dna-markers/cloning-tools-and-competent-cells/bacterial-strains-and-competent-cells/single-step-_krx_-competent-cells/ E. coli KRX]
- Medium: LB medium supplemented with 100 mg L-1 Ampicillin and 120 mg L-1 bisphenol A (Sigma, 97 %)
- BPA is thermally stable -> you can autoclave it together with the medium
- 100 mL culture in 300 mL shaking flask without baffles (Schott) with silicon plugs
- Cultivation temperature: 24 °C, 30 °C or 37 °C, tempered with Infors AG AQUATRON at 120 rpm
- for characterizations: automatic sampling every three hours with Gilson fraction controller F2XX cooled (< 4 °C) with Julabo F10 water bath
- the characterization experiment setup is shown on the picture on the right
Extraction with ethylacetate
- mix 100 µL culture supernatant with 100 µL internal standard bisphenol F (Alfa Aesar, 98 %) , 100 µg L-1)
- add 200 µL ethylacetate (VWR, HPLC grade) for extraction
- vortex (30 s)
- centrifuge for phase separation (5 min, 5000 g)
- take a bit from upper phase and put it in a clean eppi
- SpeedVac at 40 °C to remove ethlyacetate
- solve remaining BPA in water (HPLC grade), vortex (30 s)
- solubility of BPA in water only 300 mg L-1
- for LC-MS analysis of BPA, 300 mg BPA L-1 is rather too much
- if you want to detect or expect higher concentrations of BPA, solve it in an acetonitrile-water-mix
HPLC method
- C18 reverse phase column
- Isocratic method: 45 % Acetonitrile
- Flow = 0.6 mL min-1
- UV-detection at 227 nm
- Internal standard: 100 mg L-1 bisphenol F (BPF)
- Column:
- Eurospher II 100-5 C18p by [http://www.knauer.net/ Knauer]
- Dimensions: 150 x 4.6 mm with precolumn
- Particle size: 5 µm
- Pore size: 100 Å
- Material: silica gel
- Software:
- Clarity (Version 3.0.5.505) by [http://www.dataapex.com/ Data Apex]
- Autosampler:
- Midas by [http://www.spark.nl/ Spark Holland]
- Tray cooling: 10 °C
- Pump:
- L-6200A Intelligent Pump by [http://www.hitachi.com/ Hitachi]
- UV-Detector:
- Series 1050 by [http://www.hp.com/ Hewlett Packard]
LC-ESI-qTOF-MS(-MS)
HPLC method
- Column: C18 reverse phase column (Knauer [http://beta.knauer.net/products/column-detail-view/productdetail/vertex_plus_column_50_x_2_mm_blueorchid_175_18_c18-1.html Blue Orchid])
- dimension: 50 x 2 mm
- Pore size: 175 Å
- Particle size: 1.8 µm
- Flow: 0.4 mL min-1
- Column temperature: 30 °C
- Gradient:
- 0 - 1.05 min: 45 % acetonitrile
- 2.55 min: 95 % acetonitrile
- 6.00 min: 95 % acetonitrile
- 6.15 min: 45 % acetonitrile
- 12.00 min: 45 % acetonitrile
- VWR Hitachi LaChrom ULTRA HPLC equipment
- Software: HyStar 3.2, HyStarPP, mircrOTOF Control
Ionization method
- Using Bruker Daltonics micrOTOFQ
- ESI in negative mode
- Mass range: 50 - 1500 m/z
- End plate offset: - 500 V, 107 nA
- Capillary: 2500 V, 4 nA
- Nebulizer: 3 bar
- Dry gas: 8 L min-1
- Quadrupole
- Ion energy: 5 eV
- Low mass: 100 m/z
- Collision energy: 10 eV
- Collision RF: 150 Vpp
- Transfer time: 70 µs
- Pre puls storage: 7 µs
MS-MS
- Isolated mass: 243.1 +/- 0.1
- Collision energy: 30 eV
Contribution
Group: NAU-CHINA iGEM 2020
We find that there’s no detailed information about “LC-ESI-qTOF-MS(-MS)”, which is a method to detect the pollutant of BPA[1].
But we have found the similar method “LC-MS/MS”. This method uses "methyl alcohol - water" as the fluid phase, separated by the Waters ACQUITY UPLC HSS T3 chromatographic column separation and determined by the UPLC/MS/MS quantitative assay. The method could meet the requirement for the determination of BPA reference value limit specified in the Sanitary Standard for Drinking Water (GB 5749-2006), with high reproducibility and precision.
The samples were eluted under the following mass spectrometry conditions: Electrospray anion source (ESI-) mode; Multi-response monitoring mode (MRM) scanning; Colliding gas (CUR) : 0.17mpa; Atomized gas (GS1) : 0.24 MPa; Add hot gas (GS2) : 0.22 MPa; Ion source spray voltage (IS) : -4500 kV; Temperature (TEM) : 500℃; Scanning time: 50.0ms.
Here’re the results about the research.
Table1.Linear equation, correlation coefficient, LOD and LOQ of BPS, BPF and BPA in pure water(Zhou Xiaoxin, et al., Journal of Environmental Hygiene. 2020)
| Compound | Linear equation | Correlation coefficient | LOD(μg/L) | LOQ(μg/L) |
|---|---|---|---|---|
| BPS | y=3.11×106x-4.4×105 | 0.9999 | 0.002 | 0.007 |
| BPF | y=5.22×104x-5.2×104 | 0.9974 | 0.05 | 0.17 |
| BPA | y=1.87×104x+1.64×104 | 0.9996 | 0.15 | 0.50 |
Table2. Precision and recovery of BPS, BPF and BPA in different water samples (N = 6)(Zhou Xiaoxin, et al., Journal of Environmental Hygiene. 2020)
| Sample name | Compound | Background/(μg/L) | Added Scalar/(μg/L) | Amount recycled/(μg/L) | Average recovery rate/% | RSD/% |
|---|---|---|---|---|---|---|
| 2.00 | 1.91~2.05 | 100 | 3.19 | |||
| Pure water | BPS | Not detected | 7.50 | 7.26~7.77 | 101 | 2.95 |
| 25.0 | 22.9~26.8 | 103 | 5.90 | |||
| 2.00 | 1.91~2.06 | 98.4 | 2.85 | |||
| BPF | Not detected | 7.50 | 7.67~7.79 | 103 | 0.65 | |
| 25.0 | 24.1~25.5 | 98.7 | 2.44 | |||
| 2.00 | 1.93~2.09 | 101 | 3.09 | |||
| BPA | Not detected | 7.50 | 7.33~7.51 | 99.1 | 0.85 | |
| 25.0 | 23.7~25.1 | 98.1 | 2.06 | |||
| 2.00 | 2.07~2.20 | 108 | 2.31 | |||
| Spring water | BPS | Not detected | 7.50 | 7.13~7.30 | 96.4 | 1.03 |
| 25.0 | 21.6~22.5 | 88.3 | 1.71 | |||
| 2.00 | 1.93~2.00 | 98.3 | 1.47 | |||
| BPF | Not detected | 7.50 | 7.23~7.50 | 98.7 | 1.37 | |
| 25.0 | 21.8~24.1 | 92.4 | 3.57 | |||
| 2.00 | 2.03~2.06 | 102 | 0.57 | |||
| BPA | Not detected | 7.50 | 7.33~7.51 | 99.2 | 0.85 | |
| 25.0 | 24.1~26.9 | 102 | 4.03 | |||
| 2.00 | 1.96~2.08 | 102 | 2.46 | |||
| Tap water | BPS | Not detected | 7.50 | 6.57~6.95 | 90.1 | 2.35 |
| 25.0 | 23.3~24.4 | 95.8 | 1.92 | |||
| 2.00 | 1.90~1.99 | 97.3 | 1.65 | |||
| BPF | Not detected | 7.50 | 7.36~7.53 | 99.2 | 0.98 | |
| 25.0 | 22.9~25.5 | 99.1 | 4.11 | |||
| 2.00 | 1.96~2.05 | 99.3 | 1.71 | |||
| BPA | Not detected | 7.50 | 7.34~7.56 | 98.8 | 1.05 | |
| 25.0 | 22.7~25.2 | 96.0 | 3.92 |
This study analyzed 5 parts of pure water, 5 parts of mineral water, 1 part of distilled water and 10 parts of terminal water respectively. The results showed that BPS, BPF and BPA were not detected in 11 commercially available samples and 6 terminal water samples but BPA was detected in 4 parts of terminal water with concentrations of (0.59 ~ 5.23) g/L.
Thus, “LC-MS/MS” has the advantages of strong qualitative ability, strong stability and high sensitivity, and is often used in the determination of bisphenol compounds.
Reference
Zhou Xiaoxin, Hu Xiaojian. Determination of BPS, BPF and BPA in drinking water by SUPER high performance liquid chromatography tandem mass spectrometry[J]. Journal of Environmental Hygiene, 2020,10(02):196-200.
References
Monod J (1949) The growth of bacterial cultures, Annu Rev Microbiol [http://www.annualreviews.org/doi/abs/10.1146/annurev.mi.03.100149.002103 3:371-394].
Sasaki M, Tsuchido T, Matsumura Y (2008) Molecular cloning and characterization of cytochrome P450 and ferredoxin genes involved in bisphenol A degradation in Sphingomonas bisphenolicum strain AO1, Journal of Applied Microbiology [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2672.2008.03843.x/full 105(4):1158-1169]
Sasaki M, Maki J, Oshiman K, Matsumura Y, Tsuchido T (2005) Biodegradation of bisphenol A by cells and cell lysate from Sphingomonas sp. strain AO1, Biodegradation [http://www.springerlink.com/content/q7864l02734wg32m/ 16(5):449-459]
Fromme H, Küchler T, Otto T, Pilz K, Müller J, Wenzel A (2002) Occurrence of phthalates and bisphenol A and F in the environment, Water Research [http://www.sciencedirect.com/science/article/pii/S0043135401003670 36(6):1429-1438]
Chen M, Ike M, Fujita M (2002)Acute toxicity, mutagenicity, and estrogenicity of bisphenol-A and other bisphenols, Enviromental Toxicology [http://onlinelibrary.wiley.com/doi/10.1002/tox.10035/abstract 17(1):80-86]
Kitamura S, Suzuki T, Sanoh S, Kohta R, Jinno N, Sugihara K, Yoshihara S, Fujimoto N, Watanabe H, Ohta S (2005) Comparative Study of the Endocrine-Disrupting Activity of Bisphenol A and 19 Related Compounds, Toxicol. Sci. [http://toxsci.oxfordjournals.org/content/84/2/249 84(2):249-259]
//function/degradation/bisphenol
//proteindomain/internal
| chassis | E. coli |
| origin | Sphingomonas bisphenolicum AO1 |