|
In order to optimize the function of the lux promoter, We creat a hybrid promoter which combine plux with OmpR, and in this case, plux can be lead to expression by HSL and obtain by phosphorylation of OmpR protein..
According to the research of it before, we camp up with a program . Keep the sequencing of Lux Box, -35box and -10box., Then put the sequencing of C3 point of OmpR promoter into -35box and -10box.

Figure 1 structural representation of Plux promoter

Figure 2 : the analysis of conserved sequence of pLux by Weblogo
Site-directed mutagenesis at position 3,5 and 3/5 of BBa_R0062
On the basis of the nucleotide sequence of the lux pR promoter, the 20 base pair inverted repeat ACCTGTAGGA TCGTACAGGT might consititude a protein binding site. And we also learned that mutagenesis at position 3 and position 5 might cause dramatic change on the expression of downstream gene. Therefore, we generated 3 mutants (BBa_K658006 BBa_K658007 BBa_K658008) of the promoter lux pR by site-directed mutagenesis at position 3, 5 and 3/5.
By testing their strength in IR-GFP devices, we prove that mutagenesis at position 3 and position 5 can change the expression of downstream gene. Mutated promoters lux pR-3 (BBa_K658006) and lux pR-5 (BBa_K658007) dramatically enhanced the expression of the downstream gene compared with wild type promoter lux pR (R0062), while mutated promoter lux pR-3/5 (BBa_K658008) gave an even weaker expression of the downstream gene than promoter lux pR (R0062).
Based on the information about the mutated promoters, we constructed a series of population-control devices which can maintain the cell density of bacteria population at several certain values.
Lux pR strength testing device
To test the strength of promoters lux pR(BBa_R0062) and its 3 mutants lux pR-3 (BBa_K658006), lux pR-5(BBa_K658007), lux pR-3/5(BBa_K658008) , we constructed four devices( BBa_K658016 BBa_K658017 BBa_K658018 BBa_K658019).
If promoter lacl+pL(BBa_R0011) is induced by isopropyl-b-D-thiogalactopyranoside (IPTG), this device will be switched on. At sufficiently high cell density, this device produces greenish tint visible by naked eye. By measuring florescent intensities at steady state of the cell growth for these four IR-GFP devices, the strength of a promoter lux pR could be defined.
Four lux pR strength testing devices (BBa_K658016 BBa_K658017 BBa_K658018 BBa_K658019) were first cloned into plasmid pSB1A2 respectively, followed by transformation into E.coli strain BL21. Fluorescence was measured when cell growth reached a steady state (around 20h).
The results are shown in following figures:
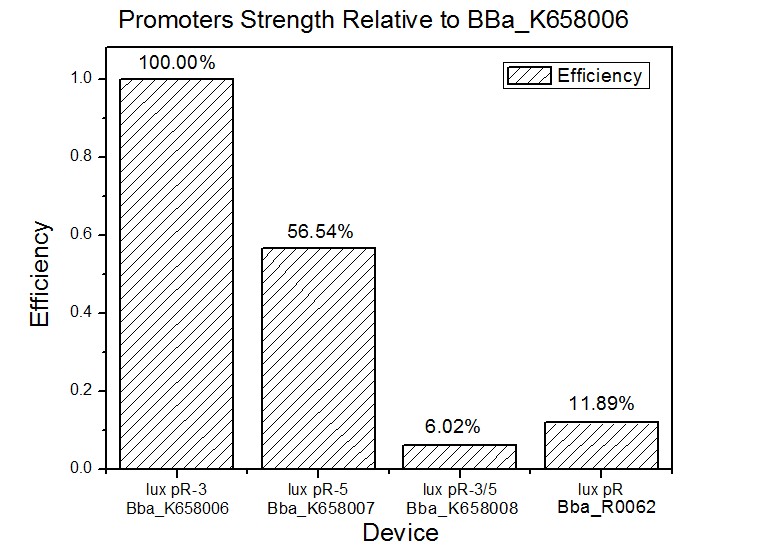
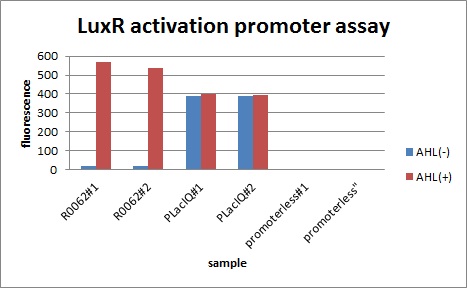
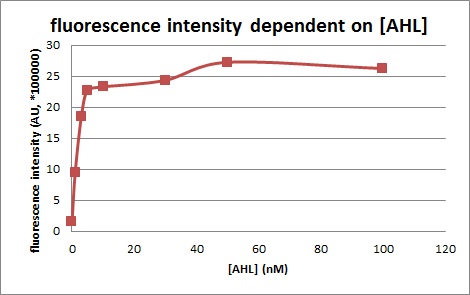
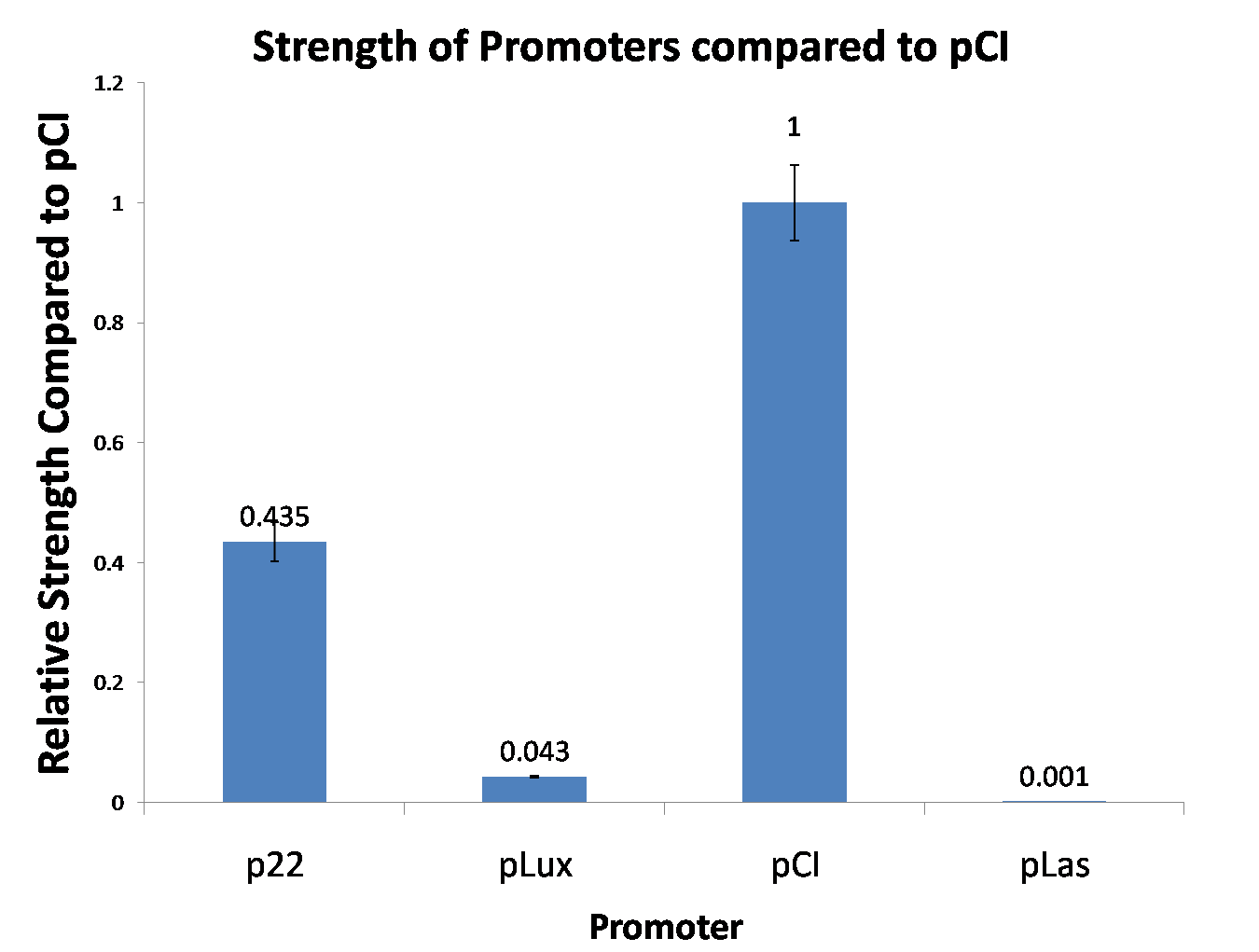
Figure 1, figure 2 and figure 3 illustrate that mutated promoters lux pR-3 (BBa_K658006) and lux pR-5 (BBa_K658007) dramatically increased the fluorescence intensity at steady state compared with wild type promoter lux pR (R0062), while mutated promoter lux pR-3/5 (BBa_K658008) gave an even weaker expression of GFP than promoter lux pR (R0062). It might be explained that the mutagenesis at position 3 and position 5 of the sequence of lux pR (R0062) changed the binding strength between promoter lux pR and protein luxR.
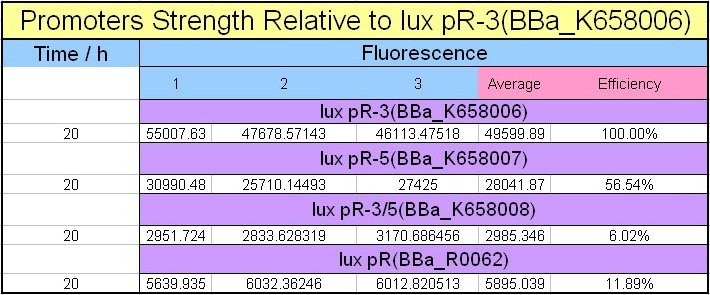 Figure 1 Promoters strength relative to lux pR-3 (BBa_K658006).
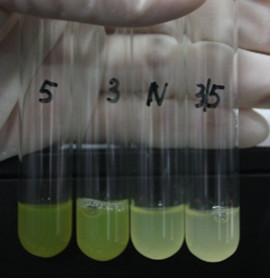 Figure 3: Fluorescence of four IR-GFP devices at 20h. 5, 3, 3/5, N represent for IR-3-GFP (BBa_K658017), IR-5-GFP (BBa_K658018), IR-3/5-GFP (BBa_K658019) and IR-GFP (BBa_K658016) respectively.

Application----a series of population-control devices
The study of the mutated promoters lux pR-3 (BBa_K658006), lux pR-5(BBa_K658007) and lux pR-3/5(BBa_K658008) can be applied to construct a series of population-control devices based on iccdB0.6 (BBa_K658003). These devices—iccdB0.6(BBa_K658003), iccdB-3(BBa_K658009), iccdB-5(BBa_K658010) and iccdB-3/5(BBa_K658011) program the steady-state cell density maintaining at different levels.
With time limited, we only tested the performance of iccdB-3(BBa_K658009) compared to that of iccdB0.6(BBa_K658003) by measuring its steady-state cell density.
The results are shown in following figures:
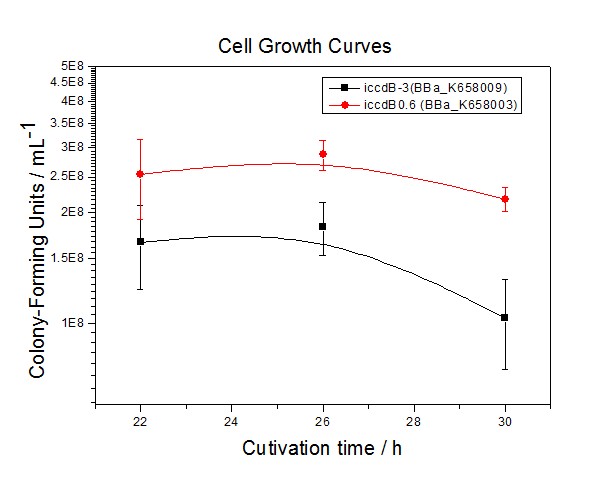 Figure 4 Experimentally measured steady-state cell density of iccdB0.6(BBa_K658003) and iccdB-3 (BBa_K658009).
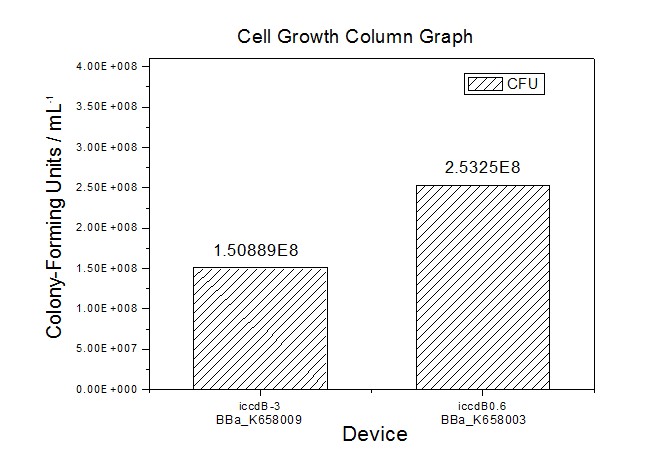 Figure 5 Average of experimentally measured cell densities of BL21’s cells with iccdB0.6 (BBa_K658003) and its mutant iccdB-3 (BBa_K658009).

Figure 4 and Figure 5 illustrate that the population-control device iccdB-3 programs a relatively lower steady-state cell density compared with iccdB0.6.
This matched the result of the test on four lux pR promoters’ strength in our IR-GFP device (BBa_K658016) mentioned above. As is shown in figure 2, promoter lux pR-3 has the highest strength of the four. It is probable that mutation at position 3 lowers the threshold for the binding reaction between LuxR/AHL protein complex and promoter lux pR, which starts the Quorum Sensing system at a relatively earlier period with a lower cell density compared with circuits regulated by wild type promoter lux pR (BBa_R0062).
Once the QS system is started, downstream killer protein expresses. The viable cell density reaches a steady state when cell growth rate equals to its death rate. Generally, steady-state cell density seems to fluctuate at the cell density when QS is started. Thus, the higher strength a promoter has, the earlier the population-control device is started, leading to a lower steady-state cell density.
| 
 1 Registry Star
1 Registry Star