Part:BBa_K500001
lignin degradation 2
Yeast codon optimization , no terminator codon, from Phanerochaete chrysosporium. Synthetized by Geneart Mn peroxidases (MnP) are extracellular hemeproteins first discovered in the white-rot fungus Phanerochaete chrysosporium, and have been virtually detected in all lignin degrading fungi so far studied. The catalytic cycle of MnP is similar to that of other plant and fungal peroxidases
Contribution
Group: iGEM19_Uppsala_Universitet
Author: Jonas Gockel
Summary: We studied the expression of BBa_K500001 in Pichia pastoris under the control of the AOX1 promoter BBa_K3105675 and an N-terminal fused α-secretion factor BBa_K3105674.
To express BBa_K500001, overhang PCR was performed to add complementary regions towards the commercial pPICZαB vector, in which MnP was cloned by gibson assembly. In this vector, the gene is controlled by the AOX1-promoter and is cloned in-frame to an N-terminal α-secretion factor from S. cerevisiae. MnP was expressed in the X-33 wildtype strain of Pichia pastoris, where secretion was observed (Figure 1).
The full expression circuit can be found here:
BBa_K3105682
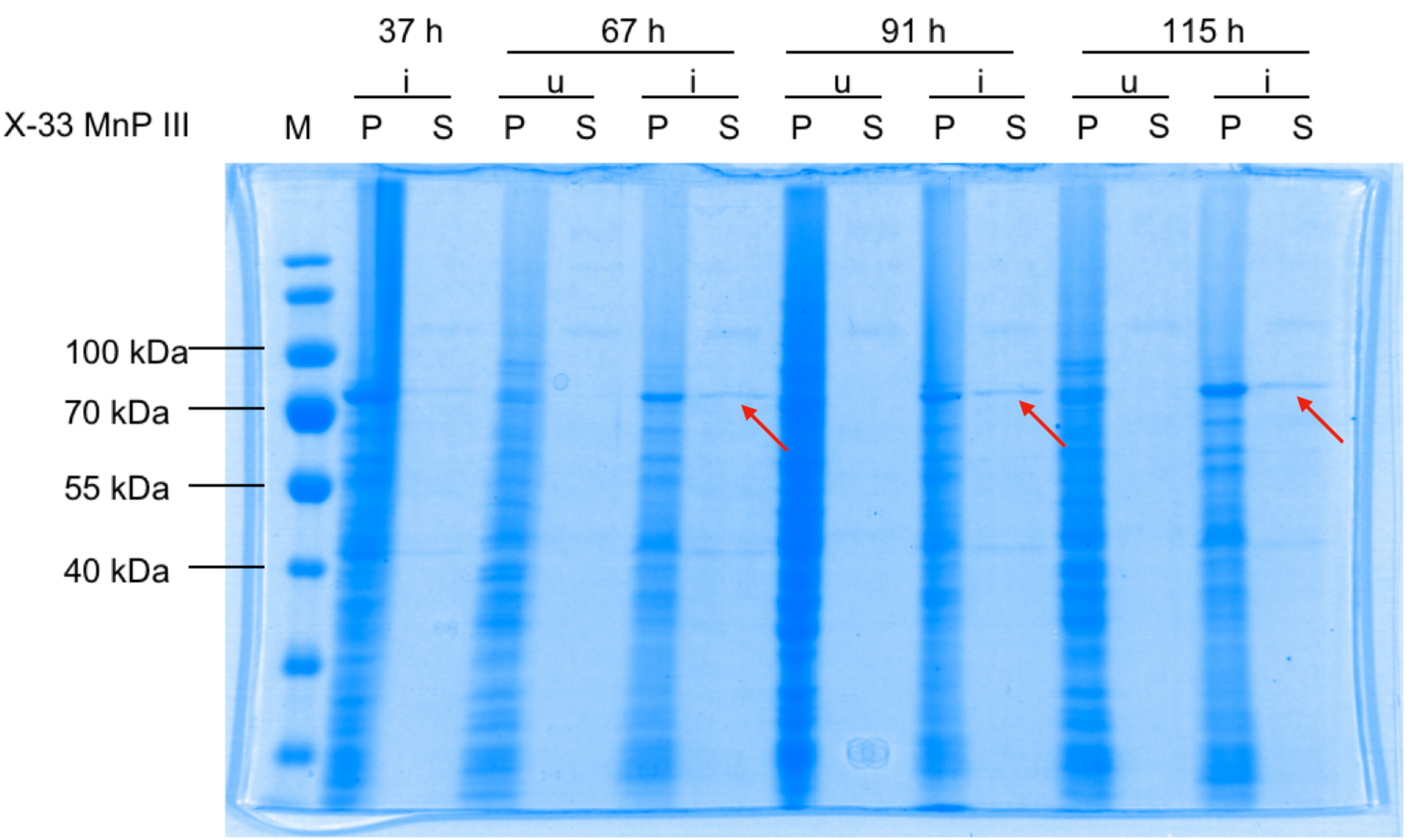
X-33 P. pastoris cells were transformed with pPICZαB_MnP, induced (i), fractionated into pellets (P) and supernatants (S), and analysed by 10 % SDS-PAGE stained with Coomassie Blue. u, uninduced
Induction bands are visible at approximately 70 kDa for both P and S (red arrows), showing secretion of MnP.
Improvement From CPU_CHINA 2021
Group: iGEM Team CPU_CHINA 2021
Author: Peng Luo
Improvement from iGEM CPU_CHINA 2021
Manganese peroxidase (MnP), a glycosylated heme enzyme derived from the white-rot fungus Phanerochaete chrysosporium, can oxidize Mn2+ to Mn3+ under the action of H2O2. Mn3+ can be released outside the enzyme under the action of a chelate such as malonic acid and can oxidise a wide range of phenolic and non-phenolic compounds as a common substrate. The Mn3+-malonic acid chelate can be detected at 469 nm by oxidation of 2,6-dimethyloxyphenol (2,6-DMP), which is also the main enzyme activity detection method for MnP. MnP (E74M) is obtained by mutating the glutamate at position 74 of wild-type MnP (BBa_K3853000) to methionine.
We mutated the glutamate at position 74 of wild-type MnP to methionine through single-point mutation in order to improve the stability of wild-type MnP. We use BBa_K3853057 to construct the expression system to express and purify the protein.
Characterization
1. Identification
After receiving the synthetic plasmid, we electrotransformed it into Pichia pastoris, and selected monoclonal colonies for colony PCR to verify the successful transformation.

Fig. 1 Agarose gel electrophoresis of PCR products of monoclonal colonies of MnP (E74M). lane 2 for MnP(E74M)
2. Proof of the expression
After the expressed protein was re-dissolved by ammonium sulfate precipitation, it was verified by running gel, and the target protein band was observed by SDS-PAGE (Fig. 2).

Fig. 2 SDS-PAGE of MnP(E74M). lane 2 for MnP(E74M)
For assaying the mRNA expression of MnP (E74M), qRT-PCRs were performed. We did data analysis using a variation of the Livak method. To determine the relative expression of MnP (E74M) vs. reference gene ACT1, total RNA was prepared from an equal volum of yeast solution. In Fig. 3, MnP (E74M) reached its peak to a fold difference of 0.11 after 2% methanol inducing for 36 h. The CT values for the MnP (E74M) and the reference gene ACT1 were then used to calculate the fold difference with the following equation:


Fig. 3 qRT-PCR results of MnP (E74M) using the relative quantitative method. Microbial samples were taken every 24 hours and ACT1 was applied as the reference gene. This result indicated that the MnP (E74M) expression reached the maximum after 36 h of fermentation in buffered complex methanol medium (BMMY) medium.
3. Enzyme Activity
MnP activity of MnP (E74M) was measured by monitoring the oxidation of 2,6-dimethyloxyphenol (2,6-DMP) at 469 nm[1]. H2O2 concentration were determined using ε240 = 43.6 M-1 cm-1.The reaction mixtures contained 0.4 mM MnSO4, 50 mM sodium malonate (pH 4.5), and 1 mM 2, 6-DMP. For a 96-well plate, 140 μl of the above reaction mixtures and 20 μl enzyme solution were mixed uniformly in advance and then 40 μl 0.1 mM H2O2 were added to initiate reaction. The concentration of 2, 6-DMP's oxidation products, 2, 2', 6, 6'-tetramethoxydibenzo-1, 1'-diquinone, were determined using ε469 = 49.6 mM-1 cm-1. One unit (U) of MnP activity is defined as the amount of enzyme required to convert 1 μM 2, 6-DMP to 2, 2', 6, 6'-tetramethoxydibenzo-1, 1'-diquinone in 1 minute.
As shown in Fig. 4A, the absorbance of the reaction system with MnP (E74M) continued to rise within 1 min, while the absorbance of the control group (without enzyme) did not change. Through UV-visible spectrum of the reaction system after 1 min, the characteristic absorption at 469 nm was observed (Fig. 4B).

Fig. 4 The detection of 2, 2', 6, 6'-tetramethoxydibenzo-1, 1'-diquinone. Mutant 2# refers to MnP (E74M). Control group refers to the reaction system without enzyme. A: The absorbance change at 469 nm in the reaction system within 1 min. B: UV-visible spectrum of the reaction system after 1 min.
4. Thermostability
To evaluate thermal stability, the purified MnP (E74M) were incubated in 20 mM sodium malonate buffer (pH 5.5) with 100 mM NaCl at different temperatures for 6 h (Fig. 5) and the residual enzyme activity were measured and calculated every 2 h. The relative enzyme activity under different temperatures were calculated with the following equation:

As shown in Fig. 5, relative enzyme activity of MnP (E74M) under different incubation temperatures displayed distinct characteristics. It exhibited good stability when the temperature wasn't high (below 60°C), and the relative enzyme activities after 6 h incubation at room temperature, 37℃ and 50℃ were 106.1%, 97.26% and 83.19%, respectively. When the temperature exceeded 60℃, a sharp decline of enzyme activity within 2 h could be observed, and it gradually decreased in the following 4 hours.

Fig. 5 Thermal stability of MnP (E74M). The initial MnP activity before incubation was set as 100%.
Compared to the MnP without any mutation, MnP (E74M)'s thermostability showed the same trend and it had significantly improved as it exhibited higher relative enzyme activity at almost all temperature gradient we set after 6 h incubation (Fig. 6).

Fig. 6 Effect of temperature on the stability of MnP (E74M) and MnP after 6 h incubation. The initial MnP activity before incubation was set as 100%. Mutant 2# refers to MnP (E74M). *P < 0.05, **P < 0.01.
5. pH stability
To evaluate pH stability, the purified MnP (E74M) were incubated in 20 mM sodium malonate buffer with 100 mM NaCl under pH 3-7 for 12 h at room temperature. The relative enzyme activity at different pH conditions were calculated with the following equation:

Compared to the MnP without any mutation, MnP (E74M) exhibited higher relative enzyme activity at pH 4 and there was no significant difference under other pH conditions (Fig. 7). This may be due to the fact that after mutating the glutamate at position 74 to methionine, the latter retained hydrogen bond with S66 and A136 (Fig. 8). These result may indicated that the hydrogen bond with S66 and A136 is important for MnP's pH stability as our other mutant exhibited poorer pH stability.

Fig. 7 Effect of pH on the stability of MnP (E74M) and MnP after 12 h incubation. The initial MnP activity before incubation was set as 100%. Mutant 2# refers to MnP (E74M). **P < 0.01.

Fig. 8 Position of Glu74 in wild-type MnP. The residue in wheat color refers to 74E. The dashed line indicates the hydrogen bond formed by glutamate at position 74 and the surrounding amino acid residues. For glutamate, the red dashed line indicates the hydrogen bond acceptor, while the blue dashed line indicates the hydrogen bond donor. Green ball indicates Ca2+ and purple ball indicates Mn2+.
References
[1] Wariishi, H., Valli, K. & Gold, M. H. Manganese(II) oxidation by manganese peroxidase from the basidiomycete Phanerochaete chrysosporium. Kinetic mechanism and role of chelators. The Journal of biological chemistry 267, 23688-23695 (1992).
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 220
Illegal BglII site found at 512 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
| n/a | lignin degradation 2 |
