Part:BBa_K1477014
Constitutive promoter +RBS+LacZ alpha+Term.
Constitutive promoter (J231202)RBS with the beta-galactosidase alpha fragment peptide coding unit(K1477014).This part is for synthesis and purification of the alpha fragment. It is not designed for blue/white screening.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Characterization and Application of BBa_K1477014
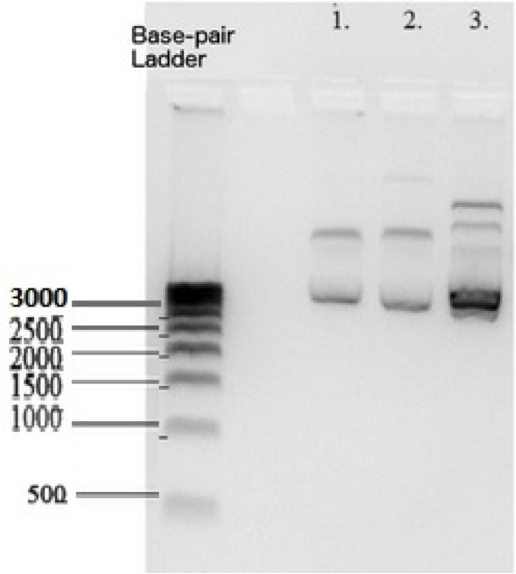
According to the Biorad 500 basepair DNA ladder that we used , K1477014 measures at ~2700 base pairs within pSB1C3 backbone. This matches the sum of the basepairs of individual parts in the device and the backbone. Plasmid DNA isolated from 3 subclones.
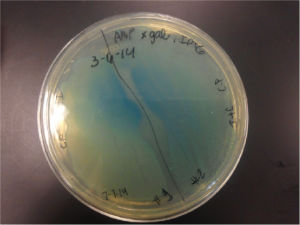
Top10 cells transformed with BBa_K1477014 form blue-green colonies on agar containing X-gal, IPTG, and selective antibiotic confirming the production of a functional β-gal alpha polypeptide and alpha complementation occuring in situ. (2 subclones)
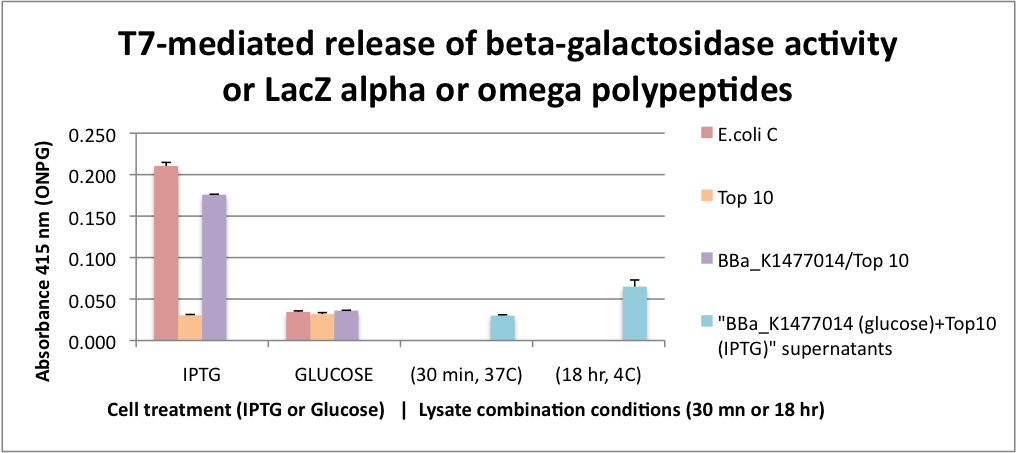
FUNCTIONAL Bgal ALPHA POLYPEPTIDE RELEASED BY CELL LYSIS. This graph represents the results for extracellular alpha complementation of beta galactosidase activity following lytic coliphage liberation of the fragments from transformed cells. To obtain only alpha fragments form BBa_K1477014 we grew the cells on glucose which suppresses the production of the omega polypeptide from the Lac promoter that controls the expression of the delta M15 mutation in the Top 10 host cells. In this assay, we either stimulated beta-gal or omega polypeptide expression with IPTG or suppressed it with glucose. After stimulation or suppression, T7 bacteriophage was added and the cells were incubated for 2 hrs to insure significant cell lysis. Released beta-gal enzyme or beta-gal polypeptide fragments were recovered with SpinX 0.22 micron filter units. Filtrates were combined with ONPG and absorbance was measured at 415 nanometers. Beta-gal activity was recovered from both E. coli C and BBa_K1477014 in ToP10 cells stimulated with IPTG and this activity was surpressed by glucose. In the case of BBa_K1477014 in ToP10 cells, the suppression of beta-gal activity was because of the inhibition of the production of the omega polypeptide. The alpha polypeptide in BBa_K1477014 in ToP10 cells is always produced because it is on a constitutive promoter. To test whether alpha complementation could occur extracellularly, a cell filtrate of IPTG Top 10 cells were combined with a cell filtrate from BBa_K1477014 in ToP10 cells grown on glucose. β-gal activity unfortunately was not detected in this assay. However we found that when the filtrates were combined and incubated at 4C for 18hrs β-gal activity was detectable suggesting that extracellular complementation had occurred at a slow rate.
Thessaloniki 2019's Characterization
Goal
Aiming at the characterization of Part:BBa_K1477014 that contains the LacZα peptide of β-Galactosidase (Part:BBa_I732006), Thessaloniki 2019 examined the expression of the LacZα fragment in Part:BBa_K1477014 in relation to vectors containing a universal constitutive Anderson Family Promoter (Part:BBa_J23100) and various RBS parts from the Community RBS Collection of the Registry Part:BBa_B0031, [Part:BBa_B0030]], Part:BBa_B0032, Part:BBa_B0033 and Part:BBa_B0034, by conducting a colorimetric β-Galactosidase assay.
Methods
β-Galactosidase is an enzyme that is commonly used as a reporter marker to monitor gene expression. It is encoded by the LacZ gene and its function in the cell is to cleave lactose to glucose and galactose. β-Galactosidase assay builds on the α-complementation phenomenon, according to which the LacZ enzyme splits into two peptides, LacZα and LacZω, neither of which is active by itself. Activation of the enzyme occurs when these two peptides reassemble and form a single unit of the Galactosidase enzyme.
In E. coli strains such as DH5α and XL1-Blue the mutated LacZω fragment is naturally found in the bacterial genome, so when a vector containing the LacZα fragment is inserted through bacterial transformation, an active form of the β-Galactosidase unit that can cleave its respected substrates can be formed. The strength of a certain RBS, being related to gene expression, can be measured via the expression of this universally used reporter.
LacZ’s activity can be quantified using an artificial substrate such o-nitrophenyl-beta-d-galactopyranoside (ONPG). This synthetic compound is also cleaved to yield galactose and o-nitrophenol which has a yellow color. When ONPG is in excess over the enzyme in a reaction, the production of o-nitrophenol per unit time is proportional to the concentration of beta-Galactosidase. Thus, the production of yellow color can be used to determine enzyme concentration and, therefore, strength of the examined RBS, since its function is immediately related to gene expression.
Miller Units are the units of measurement used β-Galactosidase assays, named after Jeffrey Miller who introduced the protocol concerning the determination of β-Galactosidase activity.
To achieve measurable response of the enzyme’s activity, we inserted the coding sequence for the LacZα fragment (Part:BBa_I732006) into a universal promoter (Part:BBa_J23100) as well as a second promoter (Part:BBa_J23102), followed by a universal RBS (Part:BBa_B0034). Constructs containing the universal promoter were followed by the rest of the RBS parts of the Community RBS Collection available in the iGEM Distribution kit for 2019 (Part:BBa_B0030, (Part:BBa_B0031), Part:BBa_B0032 & Part:BBa_B0033) were also assembled to obtain comparable results. A bi-directional terminator was added (Part:BBa_B0015) and the constructs were inserted into a high copy number pSB1C3 vector. 3A assembly was followed for the creation of all constructs and the produced vector was then transformed and expressed into E. coli DH5α cells.
A detailed version of the protocol we used regarding the β-Glactosidase assay can be found on our wiki here.
Reaction time for sample with vector containing promoter BBa_J23102 was 2 hours, while for the rest of the samples reaction time was 4 hours. Results were obtained using a plate reader measuring in 420nm to detect the yellow colour of o-nitrophenol.
Results
 Figure 1. β-Galactosidase assay. "B00NN" indicates the RBS part (as in Part:BBa_B00NN) used with promoter BBa_J23100, while "J23102" indicates the use of a vector containing promoter BBa_J23102 with universal RBS BBa_B0034. Increased strength of expression is indicated by yellow colour.
Figure 1. β-Galactosidase assay. "B00NN" indicates the RBS part (as in Part:BBa_B00NN) used with promoter BBa_J23100, while "J23102" indicates the use of a vector containing promoter BBa_J23102 with universal RBS BBa_B0034. Increased strength of expression is indicated by yellow colour.


Figure 2. Miller Units of β-Galactosidase assay. Results demonstrate the expression strength of coding sequence LacZα as a result of RBS strength. Highlighted is the output of the [[Part:BBa_K1477014]]. P100 indicates the use of Promoter BBa_J23100 while P102 indicates the use of Promoter BBa_J23102. RBS NN indicates the use of a RBS as in Part:BBa_B00NN.
Figure 3. Miller Units β-Galactosidase assay normalized to BBa_B0034. Results demonstrate the expression strength of coding sequence LacZα as a result of RBS strength in relation to strength of universal RBS BBa_B0034. Highlighted is the output of [[Part:BBa_K1477014]], as well as the output of the sample containing a vector with Promoter BBa_J23100 and RBS BBa_B0034. P100 indicates the use of Promoter BBa_J23100 while P102 indicates the use of Promoter BBa_J23102. RBS NN indicates the use of a RBS as in Part:BBa_B00NN.
| None |