Help:Nav/Part Tools
- Registry Help Pages:
- TOC
- At-a-Glance
- FAQ
Contents
Part Pages
- The Main Page acts as the hub for a part. It should contain information necessary to use a part: information, measurements, and characterization so that users know what the part is and how it works. All users can contribute to this page.
- The Design page contains information on how the part was designed (design process) and also includes its sequence and features box by default.
- The Experience page has information from users about the part. This may come in the form of a simple "this works/doesn't work," or an in-depth review. If you've used a part, be sure to share your experience. If you intend to use a part, you should check to see if others have had any experience with it.
- The Information page has editable elements for the part that classify and organize it within the Registry's database; this includes the short description, part type, categories, parameters, etc. Members belonging to the part's author's group are able to edit these elements.
- The Part Tools menu has various software tools that integrate with the part you're viewing.
Related Parts
Your Part is in these parts
The Registry database will return any composite parts that contain your part of interest, and show the DNA status (Availability) and short description for those parts. Only the last (most recently created) composite parts will be returned. In many respects, this tool is similar to the Superpart Search tool, although that will give precedence to available composite parts when returning results.
These parts are in the same categories as Your Part
The Registry database will return parts that belong to same categories as your part of interest. Depending on the categories selected for your part of interest this tool may help you find parts that work in the same chassis, have similar function, etc.
Length in Plasmids
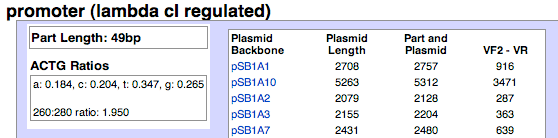
The length in plasmids tool shows the length of your part in relationship to different plasmid backbones. This information is particularly useful when analyzing gel products, either from restriction digests or PCR. Plasmid backbone length, plasmid backbone and part length
- Plasmid Backbone - The plasmid backbone column will list and link to the plasmid backbone in question. Ex. Clicking on pSB1A3 will take you to pSB1C3's main page.
- Plasmid Length - The length of plasmid backbone, from prefix to suffix. Ex. pSB1A3 is 2155bp long.
- Part and Plasmid - The length of the part plus plasmid backbone. Ex. pSB1A3 (2155bo) and the part (49p) are 2204bp total.
- VF2 - VR - Most Registry plasmid backbones have VF2 and VR sites, which are specified for primer use, either for PCR or sequencing. The VF2 and VR primer sites are usually at least 100bp away from the prefix and suffix site, respectively. The band size of your PCR product will be larger than the part length, so this column takes that into consideration. Ex. If you were to PCR the part from the pSB1A3 backbone, then the product would be 363bp.
Edit Sequence and Features
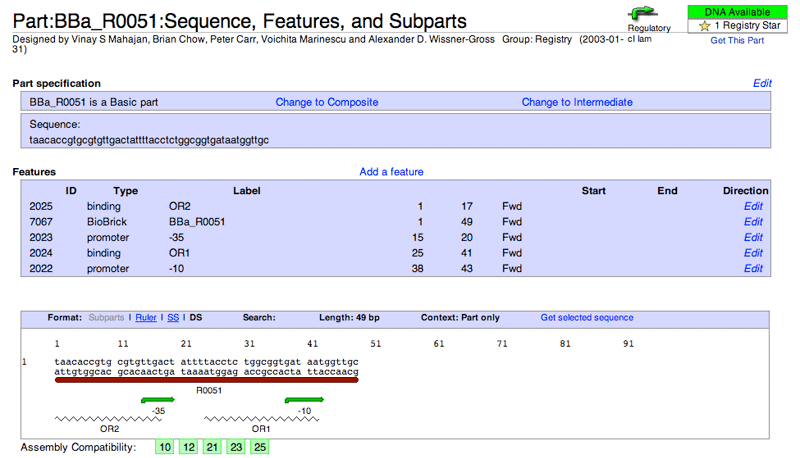
In addition to displaying the Sequence and Features box, the Edit Sequence and Features tool will also list the exact location, type, and name, of the annotations that appear for that part.
If you are the part's creator, or belong to a group that has permission to edit the part, you can use the Edit Sequence and Features tool to add/change the sequence, subparts (and scars), or annotations for the part. See the adding a basic part section or adding a composite part section for more information on this.
Sequence Analysis
The Registry has a variety of Quality Control and Sequence tools which will help you when designing your projects and choosing parts. This Sequence Analysis tool is part specific, at the header it will diagram your basic or composite part out.
You can compare a sequence to that specific part. The sequence comparison tool is useful if you've used an available sample of BBa_R0051 and want to make sure its sequence is correct.
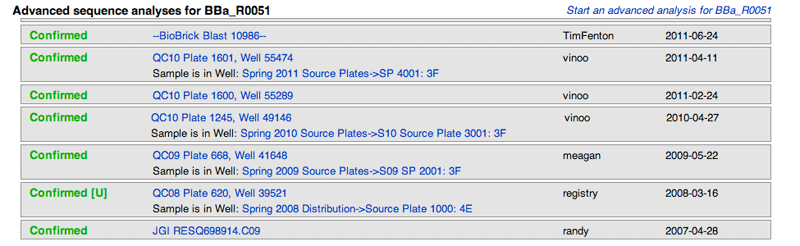
Lastly, all instances of sequence analyses done by the Registry for a part sample will be shown. Each result will show the qualitative value given for that sample, its specific location and ID (which will link to the Sequence Analysis page) and who uploaded the sequence.