Part:BBa_K3781018
3xHA, MocloMania B5
This basic part codes for a 3xHA epitope tag that can be used for western blot detection of your fusion protein. It is based on the HA epitope tag first established in 1988 by J. Field and colleagues which is derived from a small segment of the viral protein hemagglutinin (HA), one of three integral membrane proteins in the Influenza A virus.[1],[2] Fusing your protein of interest with a 3xHA-tag enables its detection using inexpensive commercial primary antibodies against the HA epitope and thus eliminates the need for expensive protein-specific antibodies. As a B5 part, this part is meant to occupy the most downstream position in the MocloMania cloning frame and is thus equipped with an additional stop codon.
size 3.56 kDa
function antibody epitope tag
amino acid sequence YPYDVPDYA GS YPYDVPDYA GS YPYDVPDYA
cloning position B5
plasmid backbone pAGM1301
Data
We were able to successfully clone this basic part into its respective L0 plasmid backbone and to confirm the integrity of the L0 construct via restriction digest and gel electrophoresis, see Figure 1. Furthermore, we were able to include it into the L1 construct L1_sAP_RBD_3xHA, proving its correct adaptation towards MoClo assembly, see Figure 2.
-
 Figure 1 | Test digest of L0_3xHA_B5 using HindIII and BstEII
Figure 1 | Test digest of L0_3xHA_B5 using HindIII and BstEII
1 | L0_3xHA_B5 | 1738 + 609 bp
2 | pAGM1301 | 1834 + 609 + 402 bp
L | Thermofischer GeneRuler Plus Ladder [bp] -
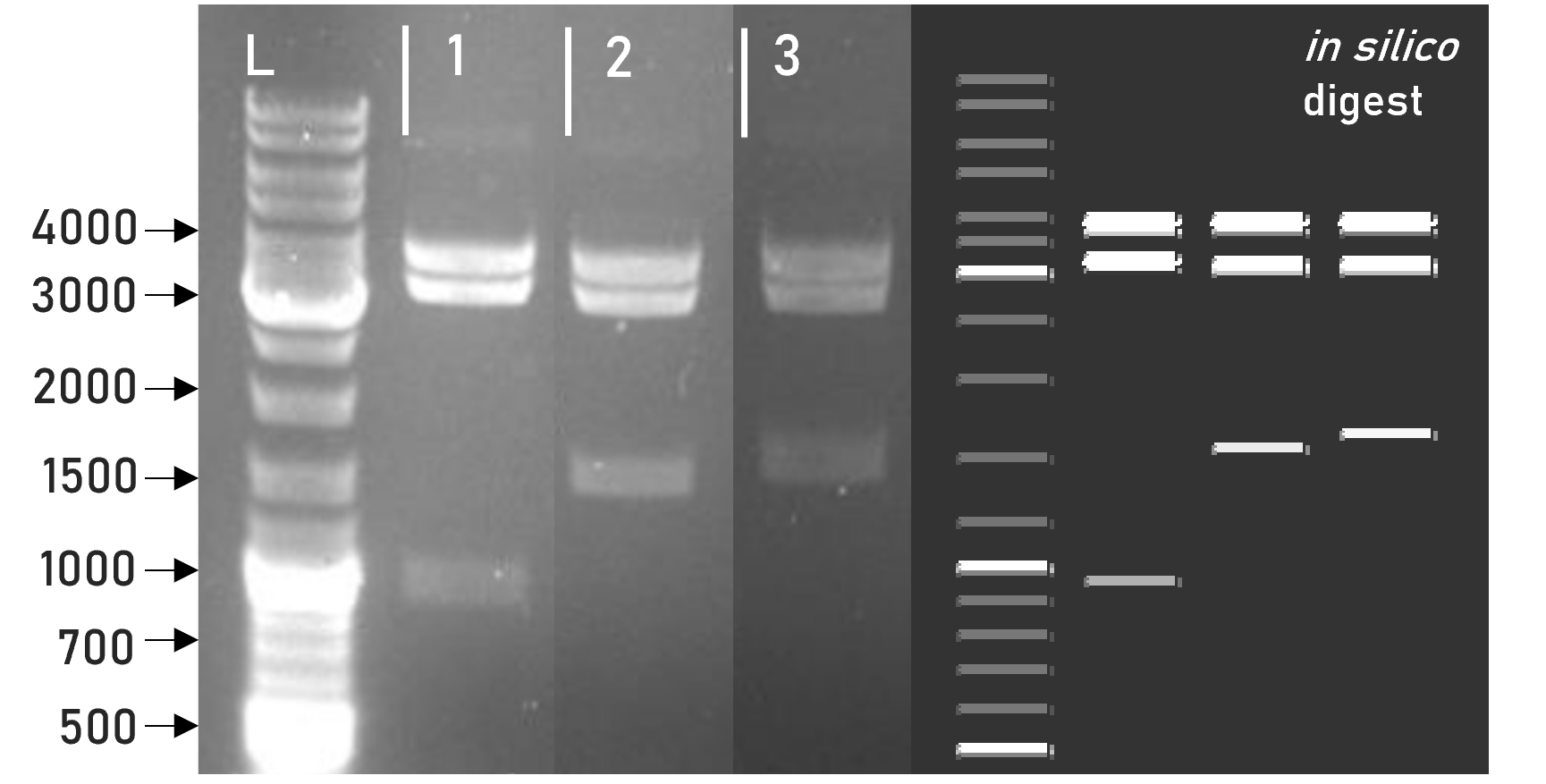 Figure 2 | Test digest of L1 constructs using HindIII
Figure 2 | Test digest of L1 constructs using HindIII
1 | L1_sAP_RBD_3xHA | 3792 + 3198 + 981 bp
2 | L1_sAP_RBD_GST | 3792 + 3198 + 1563 bp
3 | L1_sAP_mCerulean_GST | 3792 + 3198 + 1656 bp
L | Thermofischer GeneRuler Plus Ladder [bp]
L1_sAP_RBD_3xHA has been successfully transfected into Leishmania and resulting recombinant protein expression could be observed via immunostaining on western blot, see Figure 3.
-
 Figure 3 | Immunoblot of L1 transfected Leishmania cell cultures | stained against HA
Figure 3 | Immunoblot of L1 transfected Leishmania cell cultures | stained against HA
1 | L1_sAP_RBD_3xHA | 28.5 kDa
n.c. 1 | negative control 1 | untransfected Leishmania culture
n.c. 2 | negative control 2 | Leishmania culture
transfected with empty L1 expression vector
p.c. | mCherry-Ha8His | 28 kDa
L | Thermofischer PageRuler Protein Ladder [kDa]
1. AB | ms anti-HA | 1:3,000
2. AB | rb anti-ms HRP | 1:10,000
When stained against the HA-epitode, cultures transfected with 1 | L1_sAP_RBD_3xHA show a clear protein band at around 28 kDa which closely corresponds to the fusion protein's in silico predicted size. Despite the construct containing a sAP secretion tag, recombinant protein can be detected in both the cell lysate, P+, as well as in the cell culture supernatant, S+. The reason for this dysfunctional secretion mediation is unclear, but since this expression pattern has only been seen on few blots, biological variation inbetween cell cultures might factor into the effective processing of sAP secretion signal. Further western blot analysis will hopefully provide a more in-depth view on the qualitative nature of secretion vs. cytosolic expression of L1_sAP_RBD_3xHA.
As for L0_3xHA_B5, the protein bands of adequate size detected with anti-HA immunostaining verify the functionality of this L0 basic part as an effective antibody epitope tag.
The MocloMania collection
This basic part is part of the MocloMania collection, the very first collection of genetic parts specifically designed and optimized for Modular Cloning assembly and recombinant protein expression in the protozoan parasite Leishmania tarentolae.
Are you trying to express complexly glycosylated proteins? Large antibody side chains? Human proteins that require accurate post-translational modification? Then Leishmania might be just the right organism for you! Leishmania tarentolae’s glycosylation patterns resemble those of human cells more closely than any other microbial expression host, while still delivering all the benefits of microbial production systems like easy transfection and cultivation.[3] So instead of relying on mammalian cell lines, try considering Leishmania as your new expression host of choice!
Our MocloMania collection will allow you to easily modify your protein of choice and make it suitable for downstream detection and purification procedures - all thanks to the help of Modular Cloning. This cloning system was first established by Weber et al. in 2011 and relies on the ability of type IIS restriction enzymes to cut DNA outside of their recognition sequence, hereby generating four nucleotide overhangs.[4] Every basic part in our collection is equipped with a specified set of overhangs that assign it to its designated position within the reading frame. These so-called cloning positions are labelled B2-B5 from upstream to downstream. By filling all positions with the basic parts of your choice, you can easily generate variable genetic constructs that code for the fusion protein of your desire.
We furthermore provide a specifically domesticated Leishmania expression vector, named weird_plex, which will package your fusion construct into a functional transcriptional unit that is optimized for high expression in Leishmania.
The best part? Because of the type IIS restriction properties and the specifity of the generated overhangs, restriction and ligation of your construct can all happen simultaneously in a simple one-step, one-pot reaction. This will safe you a lot of time and frustration in your cloning endeavours!
Do we have your attention? In the table below you can find some basic information on how our cloning system, along with most other MoClo systems, is set up. Please feel free to check out our wiki to find more information on Leishmania and Modular Cloning as well as to understand how this basic part integrates into our part collection. See you there!
| Level | What does this level contain? | antibiotic resistance | Enzyme used for ligation |
| L0 | The foundation to every MoClo construct which are basic genetic units, such as coding sequences, promoters, terminators | spectinomycin | BbsI |
| L1 | Several L0 parts assembled into a functional transcriptional unit, e.g. consisting of promoter, coding region and terminator | ampicillin | BsaI |
| L2 | Multiple transcriptional units added into one multi-gene construct, e.g. a protein of interest fused to a selection marker | kanamycin | BbsI |
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Reference Literature
- ↑ J. Field, J. Nikawa, D. Broek, B. MacDonald, L. Rodgers, I. A. Wilson, R. A. Lerner, M. Wigler: Purification of a RAS-responsive adenylyl cyclase complex from Saccharomyces cerevisiae by use of an epitope addition method. In: Molecular and cellular biology. Band 8, Nummer 5, Mai 1988, S. 2159–2165, PMID 2455217, PMC 363397
- ↑ White JM, Hoffman LR, Arevalo JH, et al. (1997). "Attachment and entry of influenza virus into host cells. Pivotal roles of hemagglutinin". In Chiu W, Burnett RM, Garcea RL (eds.). Structural Biology of Viruses. Oxford University Press. pp. 80–104
- ↑ Langer T, Corvey C, Kroll K, Boscheinen O, Wendrich T, Dittrich W. Expression and purification of the extracellular domains of human glycoprotein VI (GPVI) and the receptor for advanced glycation end products (RAGE) from Rattus norvegicus in Leishmania tarentolae. Prep Biochem Biotechnol. 2017 Nov 26;47(10):1008-1015. doi: 10.1080/10826068.2017.1365252. Epub 2017 Aug 31. PMID: 28857681.
- ↑ Weber E, Engler C, Gruetzner R, Werner S, Marillonnet S (2011) A Modular Cloning System for Standardized Assembly of Multigene Constructs. PLoS ONE 6(2): e16765. https://doi.org/10.1371/journal.pone.0016765
| None |