Part:BBa_K2273014
| Part Information | |
|---|---|
| RFC standard | RFC 25 |
| Partner Tag | BBa_K2273015: mini. SpyCatcher |
| Original Biobrick Part | BBa_K1159201: SpyTag |
| His-tagged SpyTag | BBa_K2273016: SpyTag-His |
| Submitted by | [http://2017.igem.org/Team:TU_Dresden TU Dresden] |
SpyTag codon adapted for Bacillus subtilis
This is the codon optimized version of Biobrick BBa_K1159201 improved for ideal production in Bacillus subtilis. It codes for an oligopeptide forming an covalent isopeptide bond with a protein called SpyCatcher BBa_K1159200. There is also an shortened and codon optimized version of this protein called the Minimal SpyCatcher BBa_K2273015. The Spytag and the Minimal SpyCatcher can be fused as a tag to proteins of interest and will then mediate covalent bonding of these proteins.
Improvement
Codon adaption:
The original part was used in E. coli but the SpyTag was thought to be implemented throughout TU_Dresdens project in B. subtilis. Therefore the sequence was codon adapted for this purposes. The sequence was proposed by researchersthat investigated the secretion of SpyTag/SpyCatcher system in B. subtilis.
His-Tag:
The SpyTag was also submitted to the registry with an added C-terminal His-Tag to purify the protein constructs when SpyTag/SpyCatcher link together.
Application
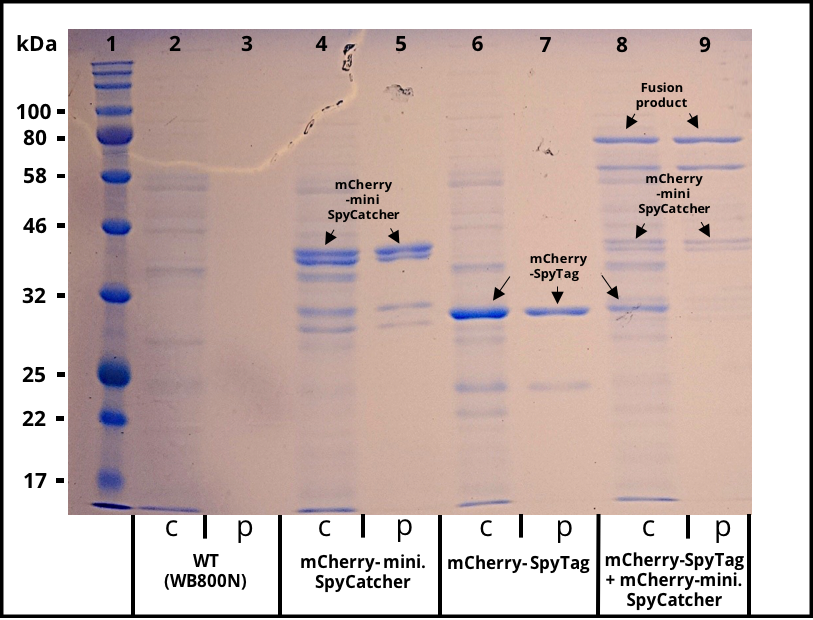
The 2017 team of [http://2017.igem.org/Team:TU_Dresden/Project/Secretion TU_Dresden] used the SpyTag together with its partner mini. SpyCatcher to investigate the potential of using B. subtilis secretion system for the production of self conjugation multi protein complexes. The his-tagged SpyTag was C-terminally fused to mCherry which functions as a reporter protein. The secretion was mediated through the N-terminal signal peptide of AmyE. The secretion of the functional mCherry fusion protein could be proven (Figure 1 and Figure 2). The functionality of the SpyTagr was proven through SDS-Page (Figure 3). After incubating supernatants containing mCherry-mini. SpyCatcher and mCherry-SpyTag the fusion product was detected, proving the covalent bonding of the Tag partners.

Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
| None |

