Part:BBa_K592025
B0034-amilCP
Purple-blue chromoprotein amilCP (BBa_K592009) with B0034 RBS. One can add any promoter upstream of this gene.
This chromoprotein from the coral Acropora millepora, amilCP, naturally exhibits strong color when expressed. The protein has an absorbance maximum at 588 nm giving it a purple/blue color visible to the naked eye, thereby requiring no instruments to observe. The strong color is readily observed in both LB or agar culture, in less than 24 hours of incubation.
Usage and Biology
This part is useful as a reporter.
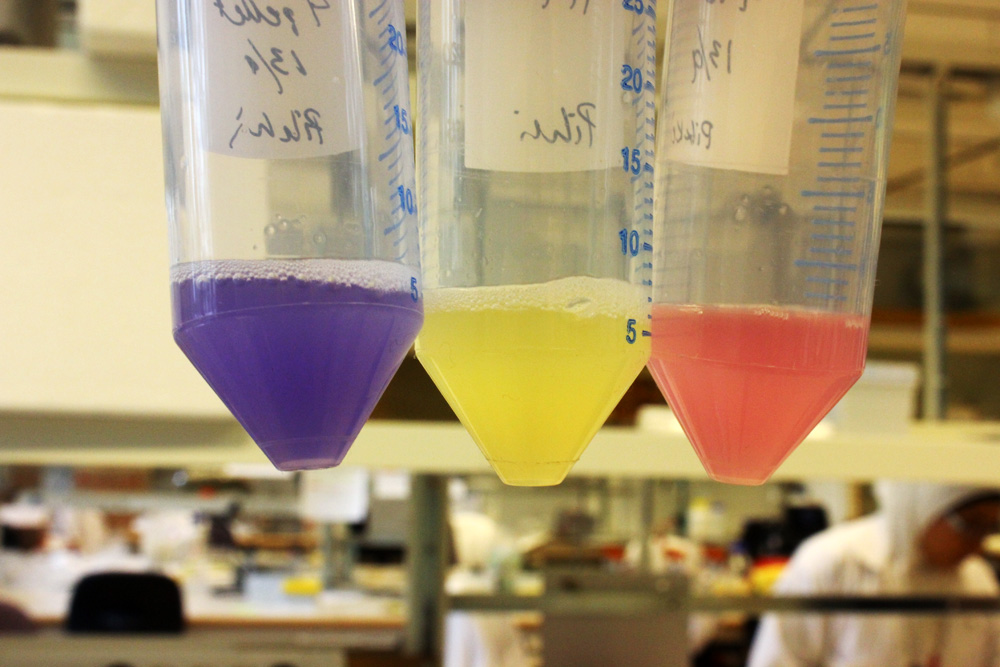
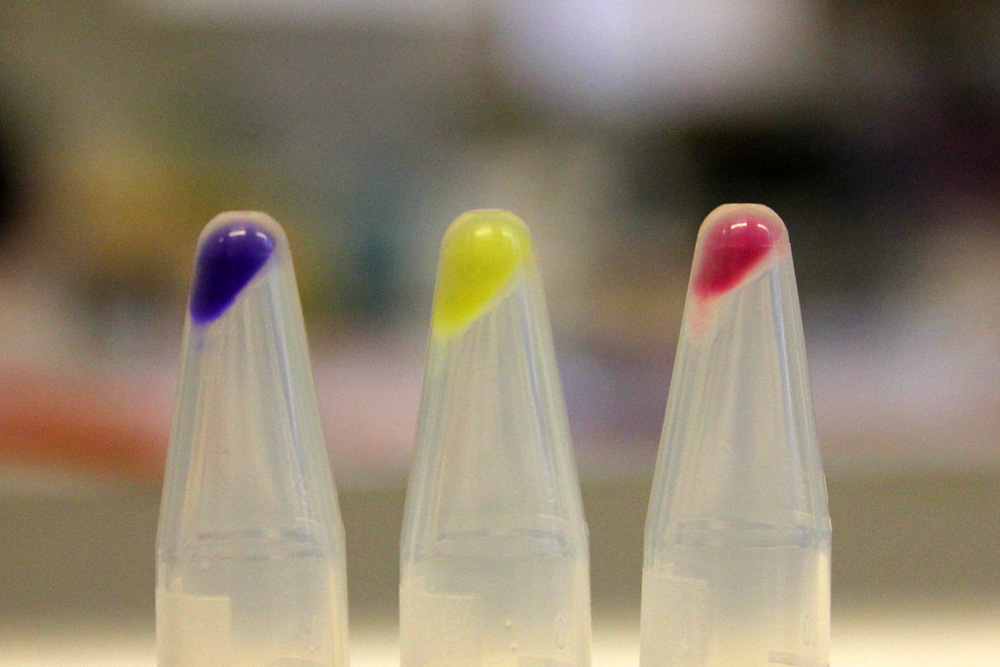
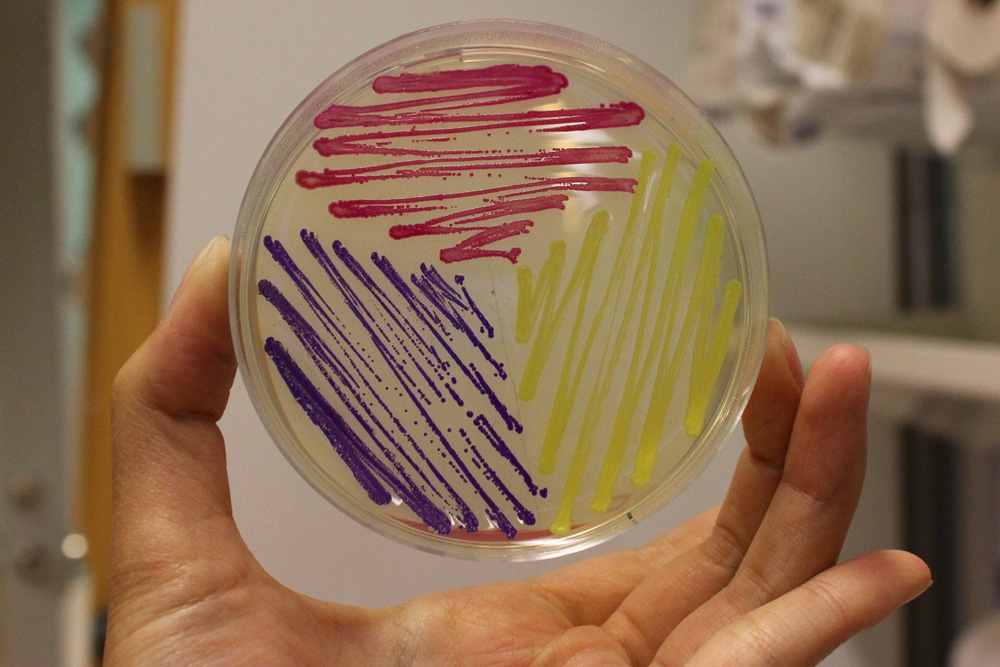
iGEM11_Uppsala-Sweden: Expression of chromoproteins. The images above show E coli constitutively expressing amilCP BBa_K592009 (blue), amilGFP BBa_K592010 (yellow) and RFP BBa_E1010 (red).
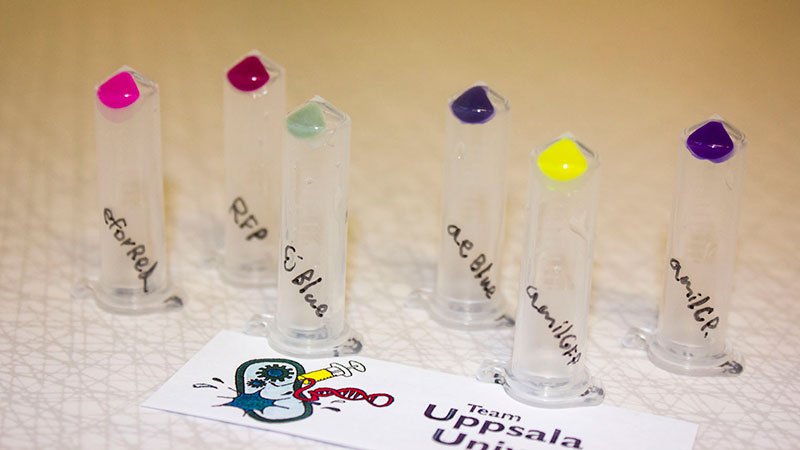
iGEM12_Uppsala_University: The Uppsala chromoprotein collection and RFP. The image shows pellets of E coli expressing chromoproteins eforRed BBa_K592012, RFP BBa_E1010, cjBlue BBa_K592011, aeBlue BBa_K864401, amilGFP BBa_K592010 and amilCP BBa_K592009.
iGEM16_CLSB-UK:The consensus promoter Part:BBa_J23119 was assembled with this reporter protein. The construct Part:BBa_K2078002 was carried on the pSB1C3 plasmid for expression and amplification in E-Coli.
Expression of the part in Synechocystis 6803 was attempted but no colour could be seen after successful transformation. We haven't done enough tests to confirm if this was due to lack of expression (unlikely given the promoter was a strong, constitutive promoter) or because natural pigments mask its colour.
iGEM16_Glasgow: This BioBrick part was ligated to P25, a native Streptococcus thermophilus promoter and transformed into S. thermophilus using the shuttle vector pMG36ET. The expected blue colour was visible in E. coli when using the same BioBrick construct, though slightly less vibrant due to the low copy number of the plasmid. However, in S. thermophilus, no blue colour was observed with transformant colonies. This suggests that amilCP is not suitable as a reporter for this organism.
References
[http://www.ncbi.nlm.nih.gov/pubmed/18648549] Alieva, N. O., et al. 2008. Diversity and evolution of coral fluorescent proteins. PLoS One 3:e2680.
CSMU_Taiwan 2020 Improve
Group: CSMU iGEM 2020
Author: Huan-Jui Chang, Cheng-Ruei Yang, Hung-Yu Chen
Introduction
This year, CSMU wants to solve the problems revolving around diagnosing oral cancer, and we decided to use toehold switches as our detection device. Toehold switches are composed of 3 main parts: trigger binding site, ribosome binding site, and a reporter gene. We wanted to find the most suitable reporter gene which can be measured with ease. Therefore, we had tried different kinds of reporter genes. Including mRFP which has been mentioned in our contribution, invertase which is our new part. We also had tried amilCP, which is a blue chromoprotein that is visible to the naked eye. With our improvement, amilCP's expression can be controlled, just like adding an on and off switch. AmilCP would not be expressed without a specific trigger under the regulation of toehold switches. By doing so, it could turn out to be a useful diagnostic tool.
Method
We followed the protocol we created. We expressed three of them, the original part ( BBa_K1087003), the toehold switch with the trigger, and the toehold switch without the trigger in vitro environment, using the cell-free PURExpress protein synthesis kits and incubating them for six hours at 37°C.
Results
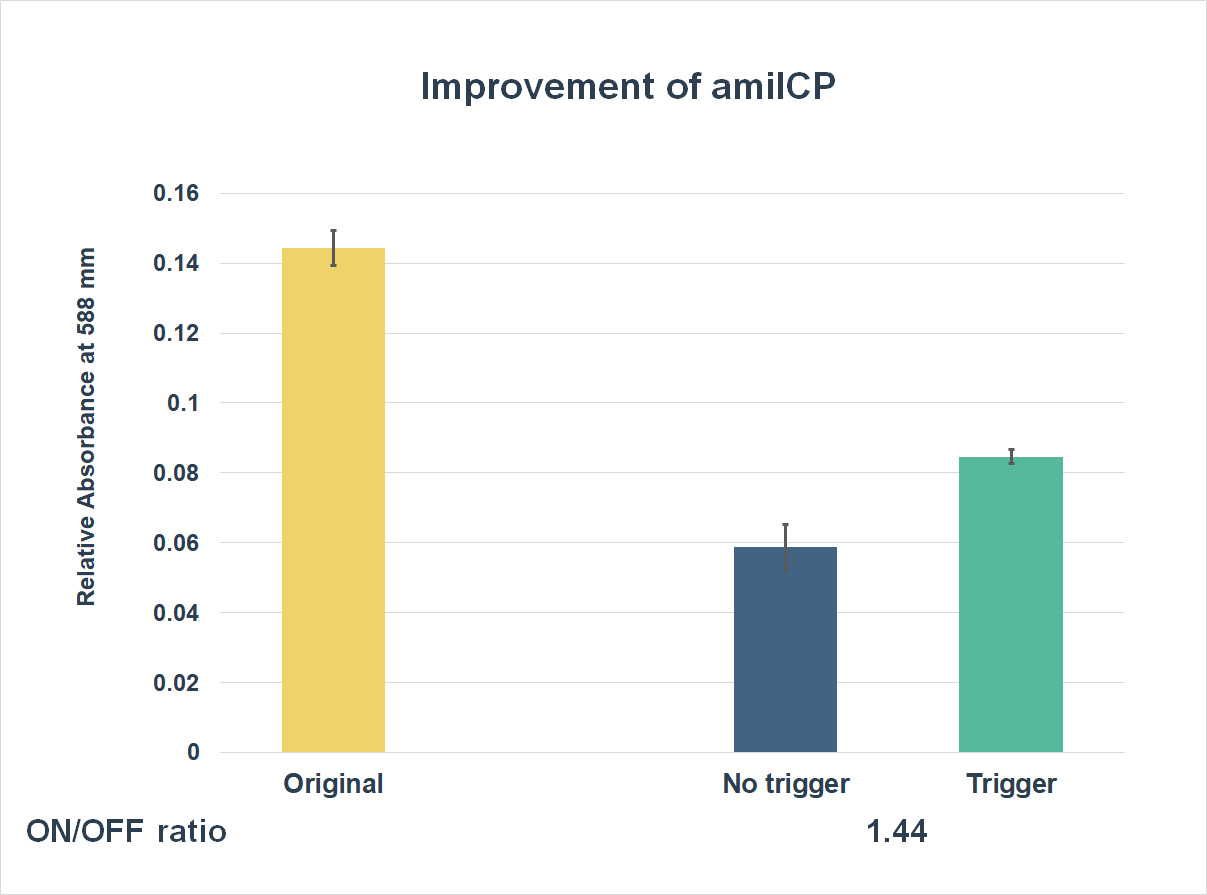
This is the graph that focuses at the sixth hour. It shows the results of our experiments with the existing part and the improved part. The vertical axis shows the relative absorbing wavelength at 588 mm. And the horizontal axis represents the name of the three. The yellow bar represents the original part, and the two bars on the right represent the improved part. The blue one refers to the condition without trigger, while the green one refers to the condition with the trigger.
The expression of our improved part is lower than the original ones. However, comparing the green one and the blue one, the green one expressed better. The ON/OFF ratio is 1.44, which was much higher than 1. Representing that there were significant differences between the ON state and the OFF state. The result proved that amilCP with toehold switches could only be triggered by a specific sequence, which also means the amilCP expression is lower in the absence of the trigger.
Conclusion
We successfully added the new function on amilCP. With our improvement, AmilCP's expression can be controlled, just like adding an on and off switch. AmilCP would not be expressed without a specific trigger. By doing so, it could turn out to be a useful diagnostic tool.
Reference
1. Alieva, N. O., Konzen, K. A., Field, S. F., Meleshkevitch, E. A., Hunt, M. E., Beltran-Ramirez, V., Miller, D. J., Wiedenmann, J., Salih, A., & Matz, M. V. (2008). Diversity and evolution of coral fluorescent proteins. PloS one, 3(7), e2680. https://doi.org/10.1371/journal.pone.0002680
2. Part: BBa_K1087003: https://parts.igem.org/Part:BBa_K1087003
3. Part:BBa_K592025: https://parts.igem.org/Part:BBa_K592025
4. Pardee, K., Green, A. A., Takahashi, M. K., Braff, D., Lambert, G., Lee, J. W., Ferrante, T., Ma, D., Donghia, N., Fan, M., Daringer, N. M., Bosch, I., Dudley, D. M., O'Connor, D. H., Gehrke, L., & Collins, J. J. (2016). Rapid, Low-Cost Detection of Zika Virus Using Programmable Biomolecular Components. Cell, 165(5), 1255–1266. https://doi.org/10.1016/j.cell.2016.04.059</p>
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
//function/reporter/color
| abs | 588 nm |