Part:BBa_K584000
pBAD + GFP generator
Characterization
The [http://2011.igem.org/Team:KULeuven K.U.Leuven 2011 iGEM Team] has extensively characterized the existing Biobrick part PBad promoter(BBa_I13453).
The detailed qualitative and quantitative review has been added into the experience page of the BBa_I13453 PBad promoter biobrick.
Usage and Biology
BOKU-Vienna 2019 - Characterization
When designing our project, we planned on incorporating a viability signal that expresses GFP in case of living cells under the control of an arabinose inducible promoter (pBAD). Since the construct that we cloned contained an arabinose inducible promoter (BBa_I13453) that didn’t respond to induction despite correct sequence, we decided to test the respective composite part BBa_K584000 (pBAD + GFP generator) under different arabinose concentrations.
The composite part BBa_K584000 was transformed into competent Escherischia coli (DH10B). Two clones were picked and grown on uninduced LB-media for 1 hour at 37°C on a shaker. 1mL aliquots were transferred into eprouvettes containing 1mL LB-media with different arabinose concentrations to an end concentration of 0, 0.0001, 0.001, 0.01, 0.05, 0.1 mM arabinose. Fluorescence was measured in a plate reader after 15 hours of incubation at 37°C on a shaker together with a fluorescein standard. OD-values were measured with a spectrophotometer right before samples were transferred into the 96-well plate.

Figure 1: Fluorescence at different arabinose concentrations
We calculated the mean fluorescence for each arabinose concentration across both clones, calculated the fluorescence for OD=1 and compared their fluorescence values (see figure 1). Fluorescence did not increase significantly after induction in comparison to the control of 0.04µM fluorescein, which equals the fluorescence of constantly expressed GFP under a medium strength promoter at OD600=0.5.
To verify this data, the experiment was repeated as above, but with 8 hours incubation time and different arabinose concentrations (see figure 2). Again, as in the first experiment, no significant change in fluorescence could be measured after induction.
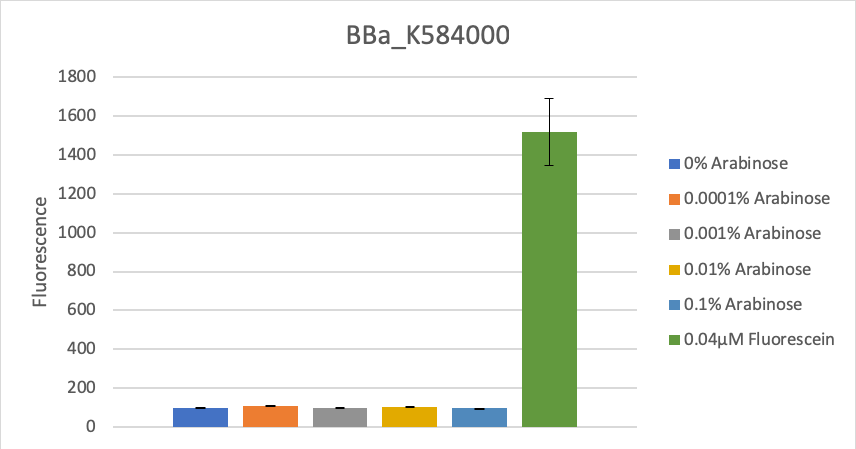
Figure 2: Fluorescence at different arabinose concentrations
Since we could not observe an induction, we checked for correctness of the plasmid by restriction digestion with the enzymes EcoRI and PstI (see figure 3).

Figure 3: Restriction digestion of BBa_K584000 - The band near 1kb matches the size of the insert, the band at 2kb the size of the empty pSB1C3 backbone
The fact that we did not observe induction of gene expression by arabinose could have different reasons. The cells might lack transporters for arabinose (e.g. araE, araFGH) or are capable of effectively degrading arabinose (e.g. by expressing araBAD genes)(Jensen et al., 1993, https://doi.org/10.1111/j.1432-1033.1993.tb19885.x).
Raw data from the plate reader here
Parameters: 485nm excitation, 525nm emission
Materials:
- LB-media with Chloramphenicol
- Tecan Infinite 200 plate reader
- white 96-well plates with white flat bottoms
- Hitachi Photometer U-1900
- L(+)-Arabinose (provider: ROTH; Art.-Nr. 5118.2)
- 2% agarose gel
- Quick-Load Purple 1kb Plus DNA Ladder
Northwestern 2019 - Characterization
The part that we chose to characterize was BBa_K584000 a pBad GFP Generator. We wanted to characterize the optimal concentration of arabinose to induce transcription activity of pBad promoter by measuring the fluorescence per absorbance OD of our cells which have reporter gene (GFP) downstream. To perform this assay, we let E. coli bacteria containing the pBad BBa_K584000 plasmid with sfGFP grow for 9 hours in a plate reader assay. We inoculated the cultures overnight, diluted them to OD = 0.05 in the morning and we let them grow in triplicates up until OD = 0.1. Based on a previous characterization done, we added in arabinose concentration of 0.1% to 0.5% in increments of 0.1% in order to find the concentration at which pBad promoter is most actively induced.
Results
Through performing our characterization assay, we demonstrated the growth of fluorescence per optical density of our pBad+GFP cells over time. From our data, we were able to determine the saturation point at which the pBad promoter was maximally induced. We additionally created a fold activation and dose response curve using the cells which were grown in different concentrations of arabinose. We observed through our experiments that the pBad promoter was strongly induced as the arabinose concentration was increased.
University of Nebraska-Lincoln 2019
Our team has characterized BBa_K584000 to determine the effect of L-arabinose concentration on fluorescence produced by green fluorescent protein. To perform this characterization, BBa_K584000 and BBa_K206000 cells were grown overnight in LB media, with BBa_K206000 functioning as the negative control. In the morning, the cultures were diluted to an OD of 0.05 and allowed to grow to an OD of 0.1. At this point, arabinose was added in the following concentrations: 0%, 0.2%, and 2%. Fluorescence was then recorded at an excitation wavelength of 490 nm and an emission wavelength of 525 nm every hour for eight hours.
Figure 1: Fluorescence vs. time with varying concentration of arabinose.
Results
Our results demonstrate that fluorescence values increased with time. Additionally, an increase in arabinose concentration yielded a greater fluorescence reading, with 0.2% arabinose producing the most GFP. These results are supported by a two-tail t-test of unequal variances with a significance level of 0.05 as shown in figure 2.
Figure 2: p-values for BBa_K584000 when compared to a negative control (BBa_K206000)
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 125
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 65
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 801
Functional Parameters: Austin_UTexas
Burden Imposed by this Part:

Burden is the percent reduction in the growth rate of E. coli cells transformed with a plasmid containing this BioBrick (± values are 95% confidence limits). This BioBrick did not exhibit a burden that was significantly greater than zero (i.e., it appears to have little to no impact on growth). Therefore, users can depend on this part to remain stable for many bacterial cell divisions and in large culture volumes. Refer to any one of the BBa_K3174002 - BBa_K3174007 pages for more information on the methods, an explanation of the sources of burden, and other conclusions from a large-scale measurement project conducted by the 2019 Austin_UTexas team.
This functional parameter was added by the 2020 Austin_UTexas team.
| None |




