Part:BBa_K3182107
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 653
Illegal BamHI site found at 580 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Introduction
pT7-CBDcipA-Pln1

This part consists of a carbohydrate binding domain (CBD) from Clostridium thermocellum (C. thermocellum) cellulose scaffolding protein (CipA). This binding domain is a central part Clostridium thermocellum's cellusome and has a strong affinity for cellulose. The CBD was fused to another protein using a flexible GS-linker (-GGGGSGGGGS-), to attach this complex to a polysaccaride material. A thrombin cleavage site (-LVPRGS-) was added to the end of the linker and its breakage will leave a glycine and serine attached to the N-terminal of the fusion protein.
Protease site and use
The thrombin site was added to enable the ability to release the fusion protein down into skin wounds. Because of our integrated human practice we learned that infection span much deeper into wounds that we thought. Simply attaching the CBD-fusion protein to a carbohydrate material wouldn't make the fusion protein reach far into the wound. The thrombin site was also chosen because of thrombins endogenous occurrence in humans.
Assembly compabilities
An internal BamHI recognition sequence (RS) has been added to enable changeable fusion proteins to the CBD. BamHI was chosen because its RS codes for glycine and serine, fitting it to the end of the thrombin site. It is also a cost-effective enzyme and is unaffected by methylated DNA. BamHI is also a part of the RFC21 standard.
CBDcipA crystal structure
Important molecular faces
CBDcipA is composed of a nine-stranded beta sandwich with a jelly roll topology and binds a calcium ion. It further contains conserved residues exposed on the surface which map into two clear surfaces on each side of the molecule. One of faces mainly contains planar strips of aromatic and polar residues which may be the carbohydrate binding part. Further aspect are unknown and unique with this CBD such as the other conserved residues which are contained in a groove.
Carbohydrate binding domain specificity
Since the CBD is from the cellusome of C. thermocellum some researches call it a cellulose binding domain. However, iGEM19 Linköping noticed that this domain could also bind to different sources of polysaccaride materials. This serves as a domain for iGEM19 Linköpings modular bandage, where the polysaccaride material can be changed for anything and not exclusively cellulose.
The choice of carbohydrate binding domain
iGEM Linköping 2019 choose CBDcipA due to many other iGEM teams exploring the possibilities of this domain. Our basic design was influenced by [http://2014.igem.org/Team:Imperial iGEM14 Imperial], [http://2015.igem.org/Team:edinburgh iGEM15 Edinburgh] and [http://2018.igem.org/Team:ecuador iGEM18 Ecuador]. Purification and where to place the fusion protein (N- or C-terminal) was determined by studying the former projects. CBDcipA also originates from a thermophilic bacteria which further increases the domains applications.
Expression system
The part has a very strong expression with a T7-RNA-polymerase promotor (BBa_I719005) as well as a 5'-UTR (BBa_K1758100) region which has been shown to further increase expression in E. coli (BBa_K1758106), ([http://www.ncbi.nlm.nih.gov/pubmed/2676996 Olins et al. 1989]), ([http://www.ncbi.nlm.nih.gov/pubmed/23927491 Takahashi et al. 2013]). Both this part and the part were sfGFP was changed for AsPink (BBa_K3182000) showed great expression.

Antimicrobial Agent - Pln1
Fused to the CBD is a Lactobacillus plantarum antibacterial peptide 1. The peptide is 38 amino acids long and has a secondary structure closely resembling membrane proteins. The function of Pln1 is to inhibit competing bacteria. The AMP has not been studied extensively but is thought to have an α-helical C-terminal and to be amphiphilic. The N-terminus consists probably of a random coil (therefore no stable structure outside the cell membrane) and and has a more hydrophilic nature. With this overlay of Pln1 is has a capability to disrupt membrane bilayers and associate themselves in pore formation inside it, leading to bacterial leakage. Pln1 has better activity to methicillin-resistant S. epidermidis (MRSE), M. luteus and same activity to S. aureus compared to nisin. Therefore, Pln1 in combination with nisin or by replacing it could be a lucrative candidate
The peptide is designed to battle the Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, Enterobacter spp. family of pathogens (ESKAPE). ESKAPE is a family(ies) of bacteria which has multiple understrains that has evolved resistance to the most commonly used antibiotics.
Usage and Biology
Expression, purification and protease treatment
The antimicrobial agents which iGEM Linköping used were all expressed the same way. Early expression tests showed great promise but with low concentrations. The early tests were done with standard E. coli expression, cultures were grown to an OD600 of 0.4-1.0 and induced with 0.5 mM IPTG, and let to express the agents for 16 hours in 18 degrees Celsius. Harvest of the cells was done at 3500 rpm for 30 minutes.
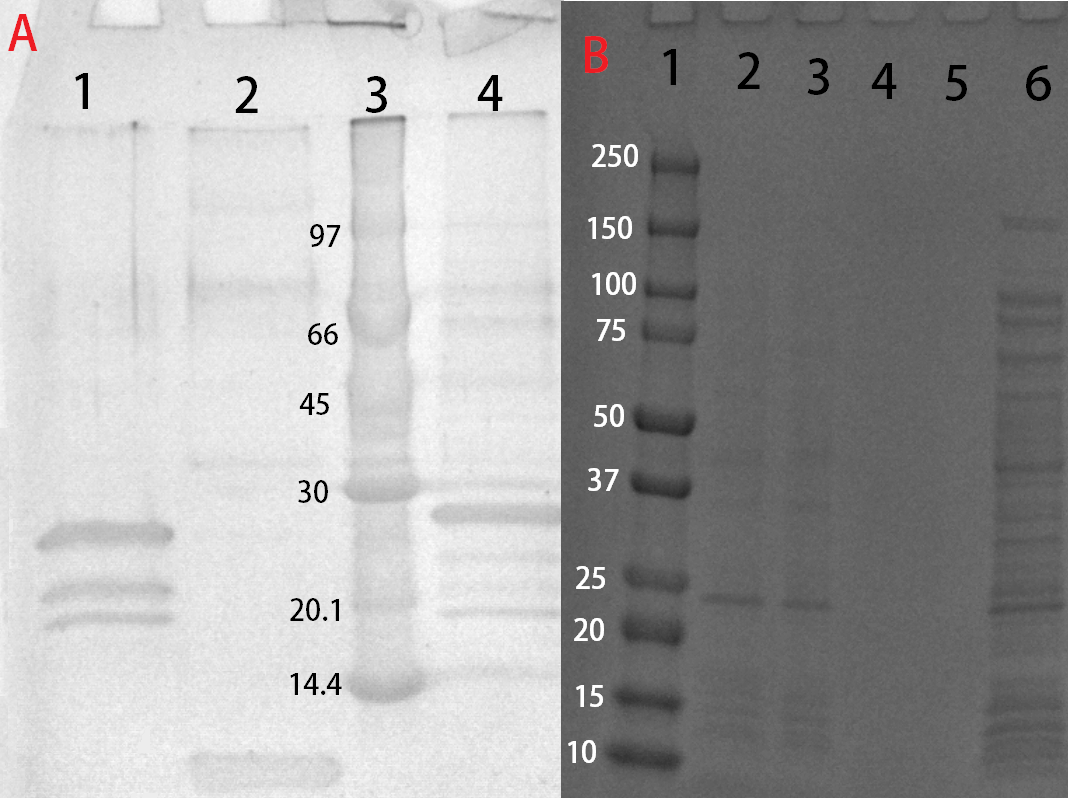
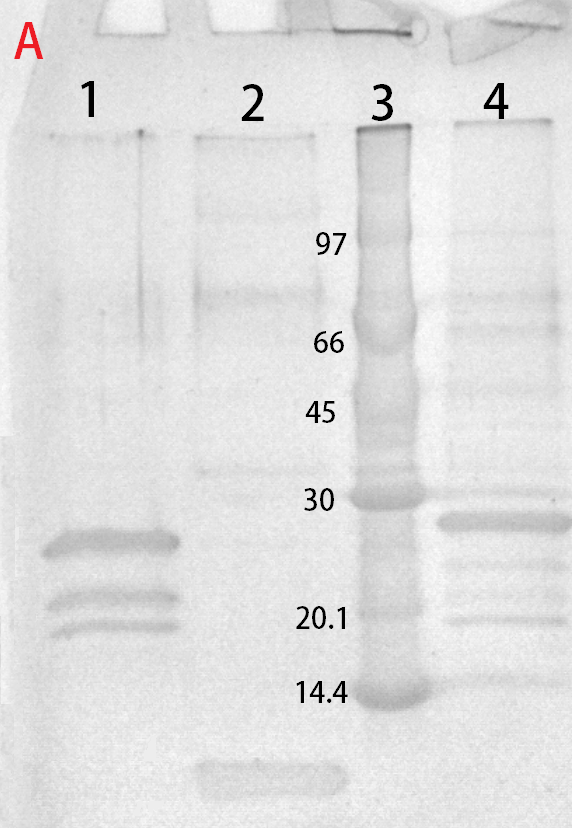
After harvest the cells was re-suspended in phosphate buffered saline (PBS) buffer and sonicated for 6 minutes, 30 % amplitude and 30 seconds on, 30 seconds off. The soluble fraction was then purified by attaching the CBD-fusion to cellulose. This showed very feint bands on SDS-PAGE analysis. Because of the low yield from the soluble fraction, a detergent (Triton X100, 1%) was used to re-suspend the lipid-soluble proteins. In this fraction a much higher protein concentration could be found.
Antimicrobial activity in solution
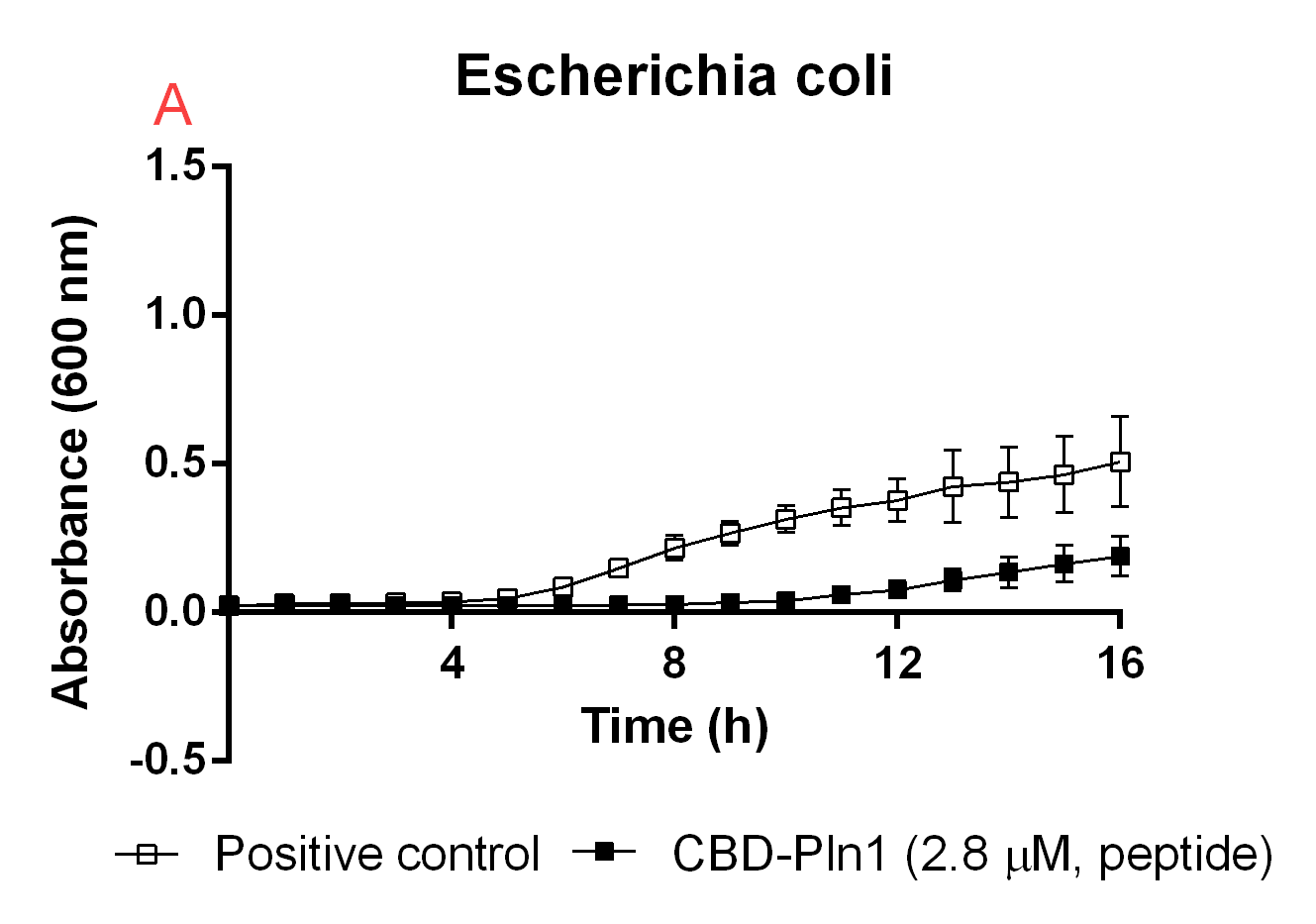
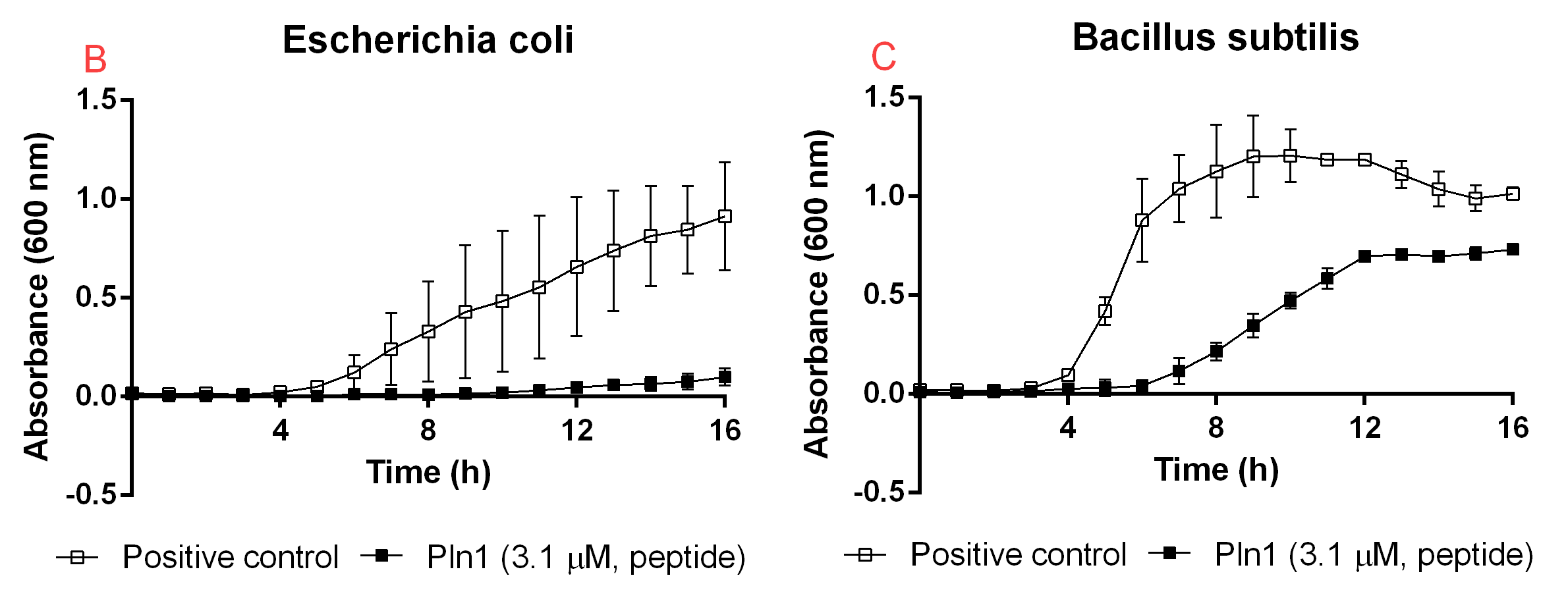
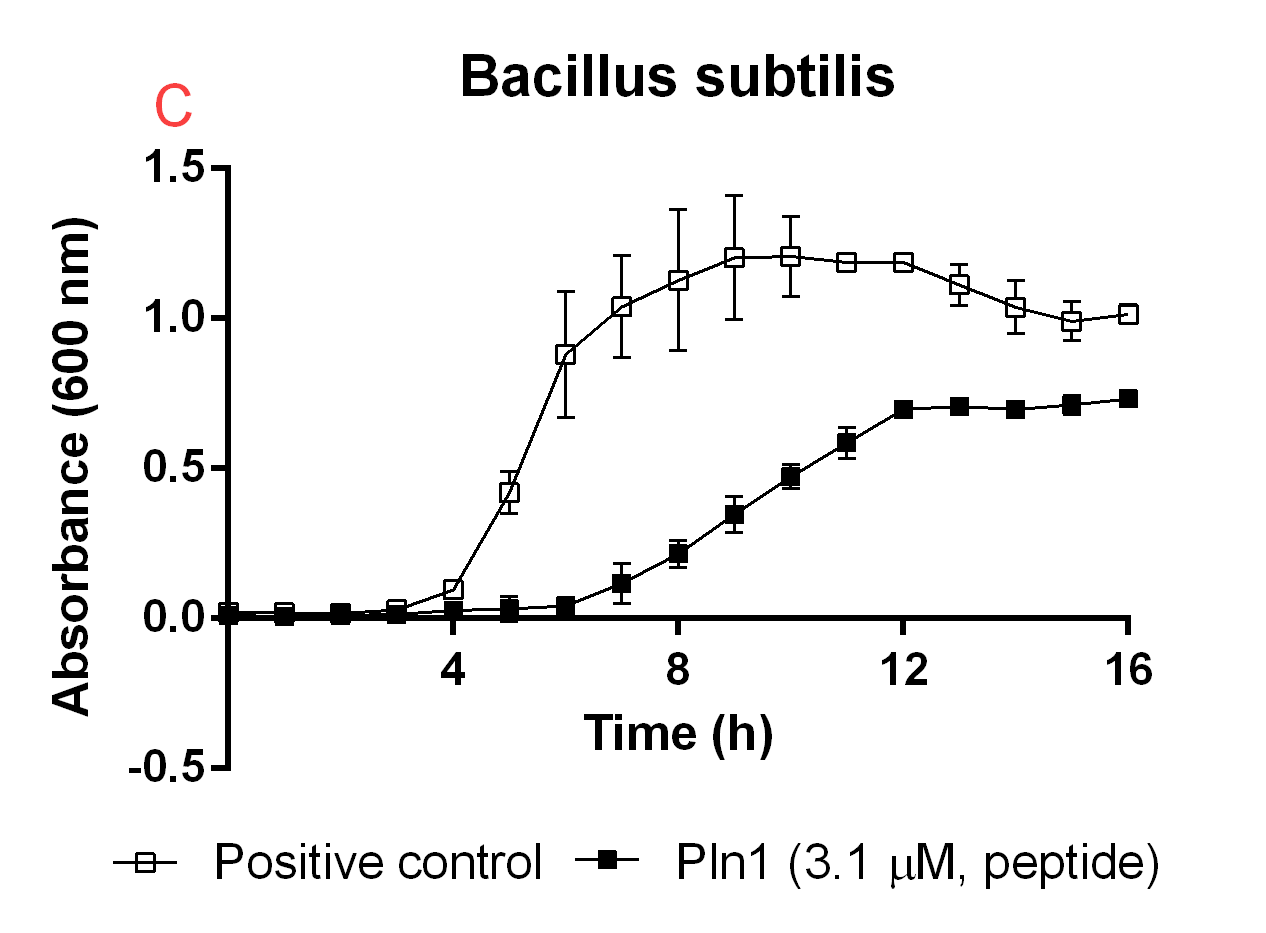 Figure B. A: CBD-Pln1 2.8 uM was added to an E. coli BL21 (DE3) 0 OD culture, the positive control is water at the same volume as the peptide added. B: Cleaved Pln1 was added to an E. coli BL21 (DE3) 0 OD culture. The positive control contain thrombin and thrombin cleavage buffer with the same volume as Pln1 (40 uL). C: Pln1 was added to a B. subtilis 0 OD culture. The positive control contain thrombin and thrombin cleavage buffer with the same volume as Pln1 (40 uL).
Figure B. A: CBD-Pln1 2.8 uM was added to an E. coli BL21 (DE3) 0 OD culture, the positive control is water at the same volume as the peptide added. B: Cleaved Pln1 was added to an E. coli BL21 (DE3) 0 OD culture. The positive control contain thrombin and thrombin cleavage buffer with the same volume as Pln1 (40 uL). C: Pln1 was added to a B. subtilis 0 OD culture. The positive control contain thrombin and thrombin cleavage buffer with the same volume as Pln1 (40 uL).
Antimicrobial activity immobilized and released
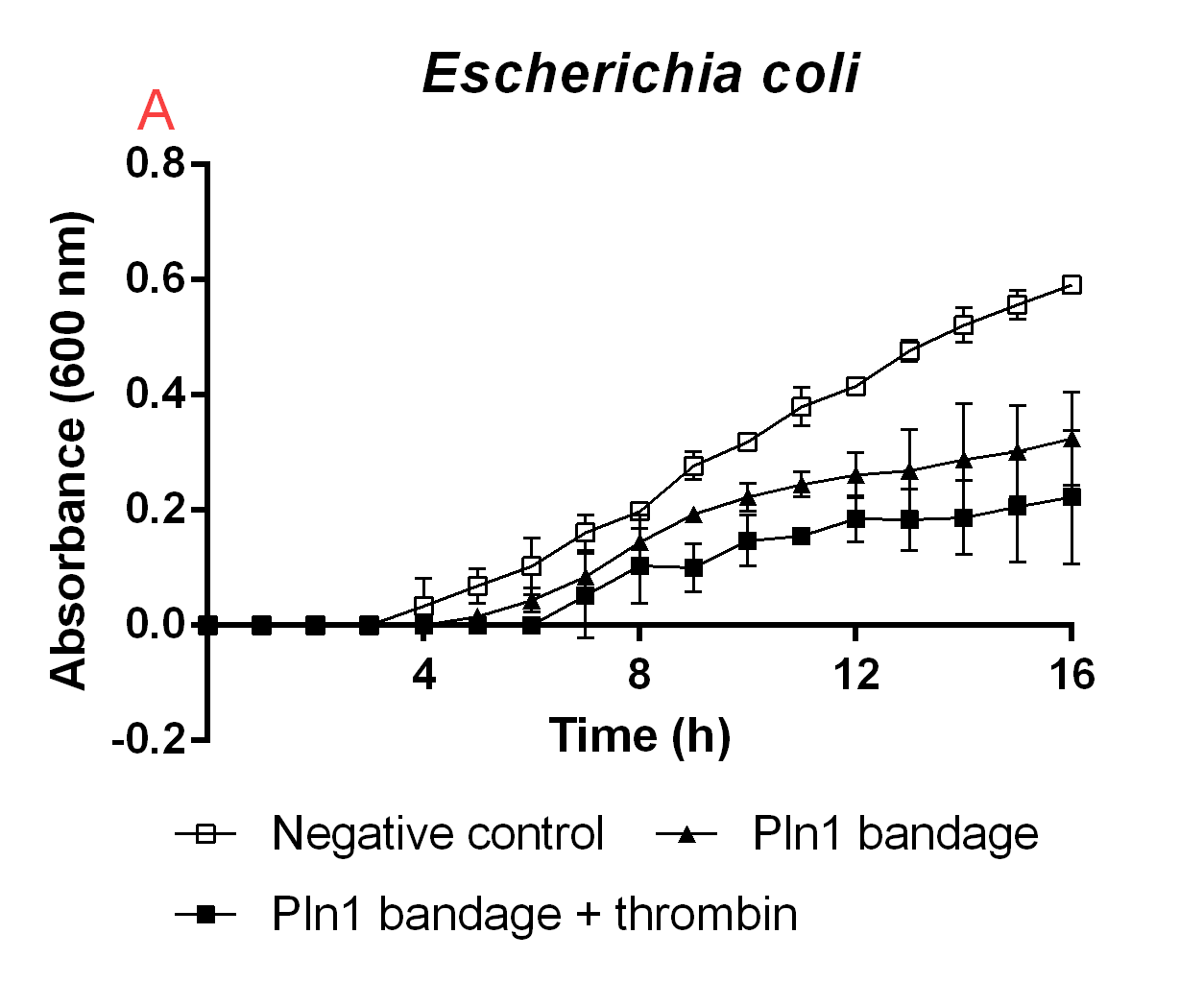
Figure B. A cellulose bandage (Epiprotect) incubated with E. coli BL21 (DE3) 0 OD cultures.
| None |