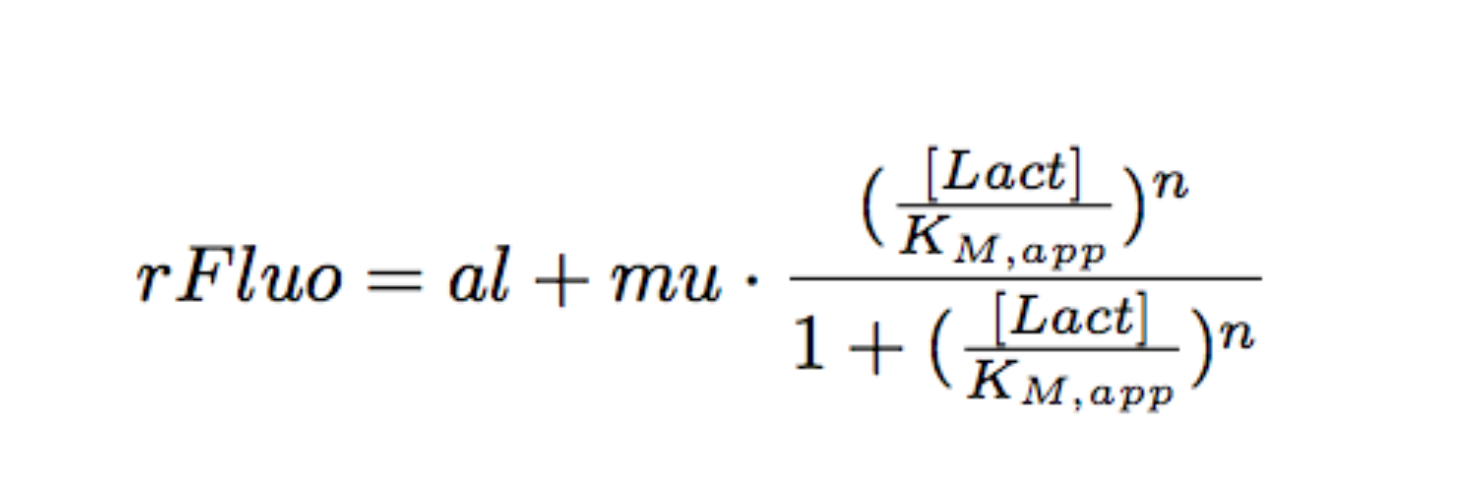Part:BBa_K1847007
lldRO1-J23100-lldRO2
Collection of promoters regulated by LldR.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 78
Illegal NheI site found at 101 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Usage and Biology
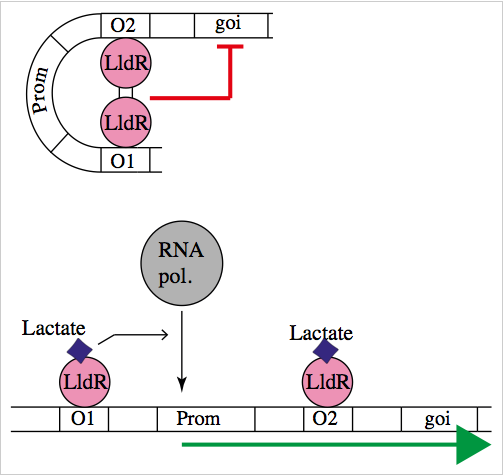
The natural promoter of LldR (Part:BBa_K822000) consists of two operators (O1 and O2) and a promoter which is intercalated between the operators. It regulates the expression of the lldPRD operon, and it is involved in L-lactate metabolism. This promoter is repressed by a dimer of LldR, possibly by forming a DNA loop that does not allow the RNA polymerase to bind to the promoter. LldR can also have a function as an activator [1]. The repression of the promoter can be removed by lactate.
Characterization of the promoter
We wanted to know how the promoter would respond to the lactate presence and how important is the architecture of the promoter in this response. For this regard, we built several promoters. Part:BBa_K1847002, Part:BBa_K1847003, Part:BBa_K1847004, Part:BBa_K1847005, Part:BBa_K1847006, Part:BBa_K1847007, Part:BBa_K1847008, and Part:BBa_K1847009.
Genetic design for characterization with LldR
To test our promoter we designed a plasmid containing the promoter with sfGFP in a medium copy plasmid pSEVA261 and transformed it into Escherichia coli TOP10. Then, we added a second plasmid containing lldR with a medium strong promoter (Part:BBa_J23118) and a strong RBS (Part:BBa_B0034) in pSEVA371 backbone. According to literature [1], LldR will repress the promoter and by the addition of lactate the production of GFP will start, as the repressor will be removed.
Genetic design for characterization with LldP-LldP
The second step for our characterization was to add an L-lactate permease (LldP), which despite its name is an active transporter of L-lactate, D-lactate and glycolate. To understand the effect of LldP (L-lactatep permease) in our system, we tried the following conditions:
- Plasmid with the promoter and sfGFP in medium copy number plasmid+ J23114-B0032-lldP-lldR (LldP and LldR are under the control of the same low promoter) in low copy number plasmid
- Plasmid with the promoter and sfGFP in medium copy number plasmid + J23118-B0034-lldP-lldR (LldP and LldR are under the control of the same medium promoter) in low copy number plasmid
Plate reader
E. coli TOP10 strains were grown overnight in Lysogeny Broth (LB) containing kanamycin (50 µg/mL) and chloramphenicol (12.5 µg/mL) at 37°C and 200 rpm. Cultures were diluted 1:50 in fresh LB with the corresponding antibiotic and transferred to a 96-well plate (200 µL/well). Cultures were grown for 90 min to arrive to exponential phase and then different concentrations of lactate were added. Samples were always made in triplicates and a blank of LB with the corresponding lactate concentration was done. During 7 h the absorbance at OD600 and fluorescence (excitation 488 nm and emission 530 nm) were measured with intervals of 7 min. The plate was always kept at 37°C in a Tecan Infinite M200 Pro Plate Reader. We calculated dose-response curves from the exponential phase of the bacteria using normalized fluorescence corrected by optical density.
Modeling
Each experimental data set was fitted to a Hill function using the Least Absolute Residual method (Fitting Toolbox in MatLab).
Where:
- al: leakiness in µM/min
- mu: constant in µM/min
- Km: half-maximum effective concentration (a representation of sensitivity) in µM
- n: Hill coefficient (a representation of cooperativity), unitless
| J23118-B0034-lldR | J23118-B0034-lldP-lldR | J23114-B0032-lldP-lldR | |
|---|---|---|---|
| Coefficient values | KM=697.7 (75.16, 1320) | KM=1977 (474.5, 3479) | KM=1337 (470, 2203) |
| al = 1.338e+04 (8443, 1.832e+04) | al = 1102 (-353.6, 2558) | al = 2e+04 (1.767e+04, 2.233e+04) | |
| mu = 1.798e+04 (1.1e+04, 2.496e+04) | mu = 4252 (2395, 6109) | mu = 2.599e+04 (2.028e+04, 3.17e+04) | |
| n1 = 0.8499 (0.1814, 1.518) | n1 = 1.481 (0.1392, 2.824) | n1 = 0.8996 (0.4539, 1.345) | |
| Goodness to fit | sse: 1.1364e+10 | sse: 3.7467e+08 | sse: 2.0117e+10 |
| rsquare: 0.9586 | rsquare: 0.9131 | rsquare: 0.9826 | |
| fe: 7 | dfe: 7 | dfe: 7 | |
| adjrsquare: 0.9408 | adjrsquare: 0.8758 | adjrsquare: 0.9752 | |
| rmse: 4.0292e+04 | rmse: 7.3160e+03 | rmse: 5.3609e+04 |
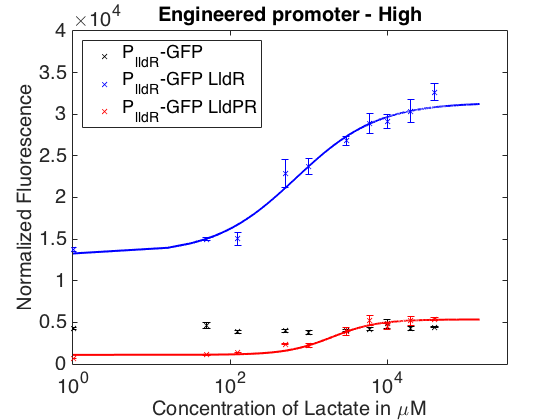
As can be seen in the picture above, when PlldR GFP is alone, the fluorescence levels do not vary with lactate concentration. However, if PlldR GFP is expressed in a system with constitutive expression of LldR, at low concentrations of lactate there is repression of the promoter, while at high concentrations there is activation. Activation is higher when there is constitutive expression of LldP-LldR. This promoter, however, is very leaky when LldP-LldR is added.
This promoter is one from our designed and tested synthetic promoters based on the wild-type PlldR. In the following table, one can see the ON/OFF ratios and the KM of the wild-type promoter and the three more prominent members of the library.
Table 1: ON/OFF ratio
| LldR | Additional LldP | ||
|---|---|---|---|
| J23118-B0034-lldR | J23118-B0034-lldP-lldR | J23114-B0032-lldP-lldR | |
| Part:BBa_K822000 (wild-type) | 10.35 | 8.04 | 1.16 |
| Part:BBa_K1847008 | 15.26 | 23.96 | 1.42 |
| Part:BBa_K1847009 | 2.5 | 24.34 | 0.96 |
| Part:BBa_K1847007 | 2.15 | 3.85 | 1.29 |
Table 2: KM (µM)
| J23118-B0034-lldR | J23118-B0034-lldP-lldR | J23114-B0032-lldP-lldR | |
|---|---|---|---|
| Part:BBa_K822000 (wild-type) | 955 | 1930 | 720.2 |
| Part:BBa_K1847008 | 1075 | 1751 | 337.5 |
| Part:BBa_K1847009 | 977.5 | 2361 | 459.8 |
| Part:BBa_K1847007 | 697.7 | 1977 | 1337 |
References
- J Bacteriol. 2008 Apr; 190(8): 2997–3005.
| None |