Part:BBa_K316003
XylE - catechol 2,3-dioxygenase from P.putida with terminator
Catechol or catechol 2,3-dioxygenases (C2,3O) + O(2) is converted by a ring cleavage into 2-hydroxymuconate semialdehyde which is the toxic and bright yellow-coloured substrate1. This is a key enzyme in many (soil) bacterial species used for the degradation of aromatic compounds. Catechol 2,3-dioxygenase2 was originally isolated from Pseudomonas putida and is a homotetramer of C230 monomers. The tetramerization interactions position a ferrous ion critical for enzymatic activity. It has been deduced that intersubunit interaction is essential to produce a functioning enzyme after performing N and C terminal modifications on the monomer. Coming together the subunits generate an active site. The reaction itself takes place within seconds after the addition by Pasteur pipette or spraying of catechol at a 100mM stock solution diluted with DDH20 (used by our lab.) The toxic byproduct is thought to interfere with cell wall integrity and cellular machinery such that exposed cells gradually die.
Safety
Catechol is classed as irritant in the EU but as toxic in the USA, as well as being a possible carcinogen. It should therefore be handled with care and proper safety equipment. More information is available on the Material Safety Data Sheet[http://www.sciencelab.com/msds.php?msdsId=9927131].
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 337
Illegal NgoMIV site found at 509
Illegal AgeI site found at 860 - 1000COMPATIBLE WITH RFC[1000]
Part Characterisation
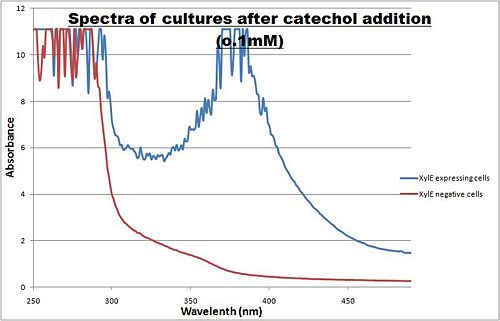
The enzymatic reaction catalysed by XylE can also serve as a powerful reporter. The substrate - catechol - is colourless. However within seconds of its addition, the colonies or liquid cultures of XylE-expressing cells become clearly yellow3 to the naked eye. This reaction allows direct measurement of XylE activity by measuring product concentrations, which absorbs light in the visible spectrum, at 380nm. In contrast to other common reporters, like GFP,S a XylE assay, as stemming from Pseudomonas putida assay would also be functional in thermophile environments. The spectrophotometric assay compared the spectra of two cultures of E.coli (one XylE gene transformed and the other not) were compared on addition of 0.1mM Catechol substrate.
Figure I. XylE assay : Peak absorbance of catechol breakdown product (2-hydroxymuconic semialdehyde).
A spectrophotometric assay of two cultures of E.coli (Blue: contains BBa_K316004, Red: not expressing XylE ) were compared on addition of 0.1mM Catechol substrate. The spectra show that in XylE transformed cells, a broad peak appears at about 380nm. The absorbance at this particular wavelength is due to the yellow product of the reaction (2-hydroxymuconic semialdehyde (HMS)).
Characterisation data was obtained using GFP-XylE constructs BBa_K316007 and XylE under two different promoters: B. subtilis derived Pveg BBa_K316005 and J23101 BBa_K316004 from E. coli. These are described on our wiki[http://2010.igem.org/Team:Imperial_College_London/Results] and the aforementioned parts pages.
References
<biblio>
- 1 pmid=10368270
- 2 pmid=12519074
- 3 pmid=6405380
</biblio>
| n/a | XylE - catechol 2,3-dioxygenase from P.putida with terminator |