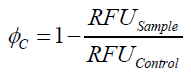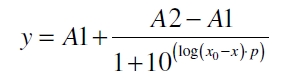Part:BBa_K525305
Fusion Protein of S-Layer SgsE and mCitrine
Fusion protein of S-layer SgsE and mCitrine
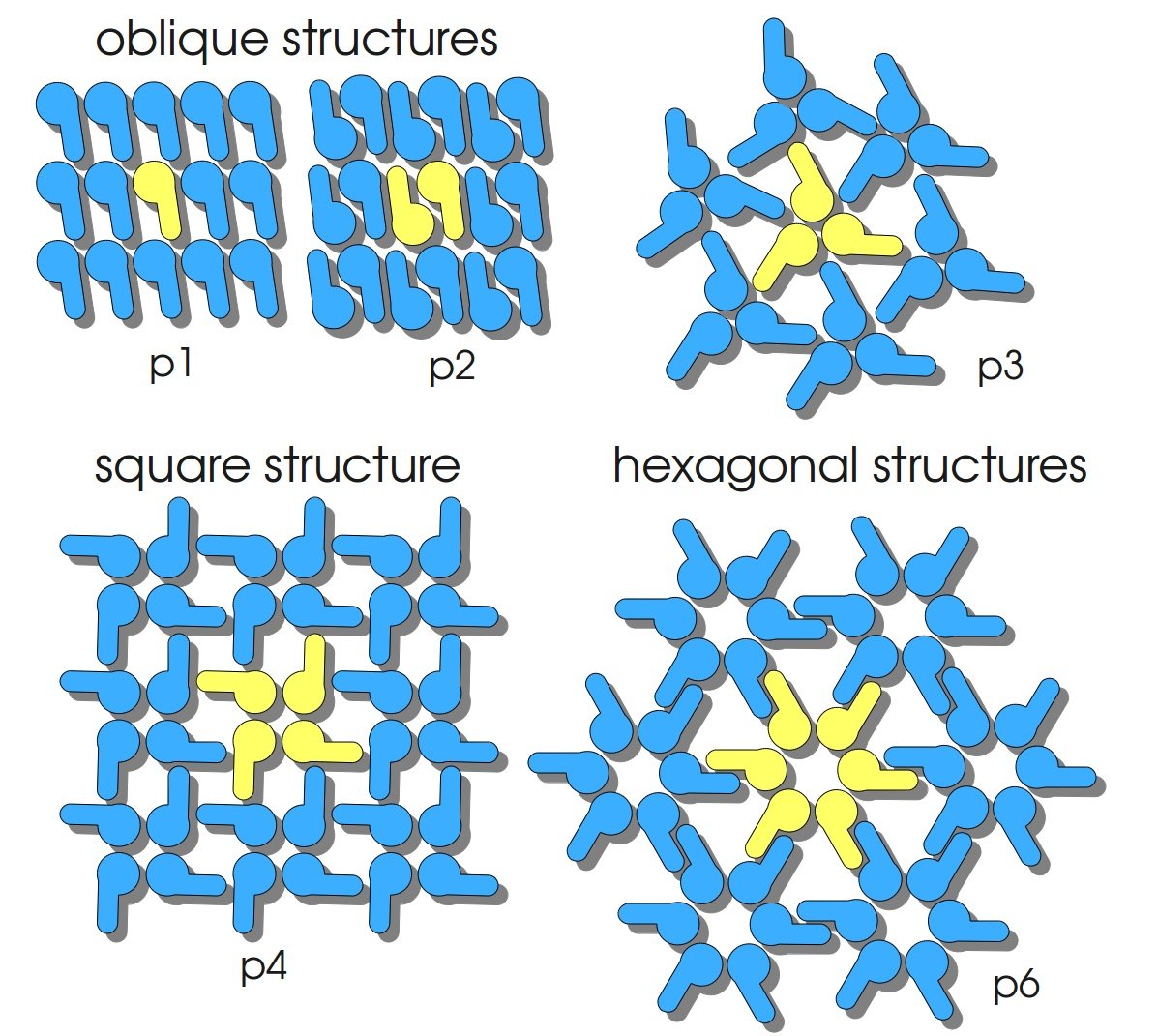
S-layers (crystalline bacterial surface layer) are crystal-like layers consisting of multiple protein monomers and can be found in various (archae-)bacteria. They constitute the outermost part of the cell wall. Especially their ability for self-assembly into distinct geometries is of scientific interest. At phase boundaries, in solutions and on a variety of surfaces they form different lattice structures. The geometry and arrangement is determined by the C-terminal self assembly-domain, which is specific for each S-layer protein. The most common lattice geometries are oblique, square and hexagonal. By modifying the characteristics of the S-layer through combination with functional groups and protein domains as well as their defined position and orientation to eachother (determined by the S-layer geometry) it is possible to realize various practical applications ([http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.2006.00573.x/full Sleytr et al., 2007]).
Usage and Biology
S-layer proteins can be used as scaffold for nanobiotechnological applications and devices by e.g. fusing the S-layer's self-assembly domain to other functional protein domains. It is possible to coat surfaces and liposomes with S-layers. A big advantage of S-layers: after expressing in E. coli and purification, the nanobiotechnological system is cell-free. This enhances the biological security of a device.
This fluorescent S-layer fusion protein is used to characterize purification methods and the S-layer's ability to self-assemble on surfaces. It is also possible to use the characteristic of mCitrine as a pH indicator ([http://pubs.acs.org/doi/abs/10.1021/bm901071b Kainz et al., 2010]).
Important parameters
| Experiment | Characteristic | Result |
|---|---|---|
| Expression (E. coli) | Localisation | Inclusion body |
| Compatibility | E. coli KRX and BL21(DE3) | |
| Induction of expression | expression of T7 polymerase + IPTG or lactose | |
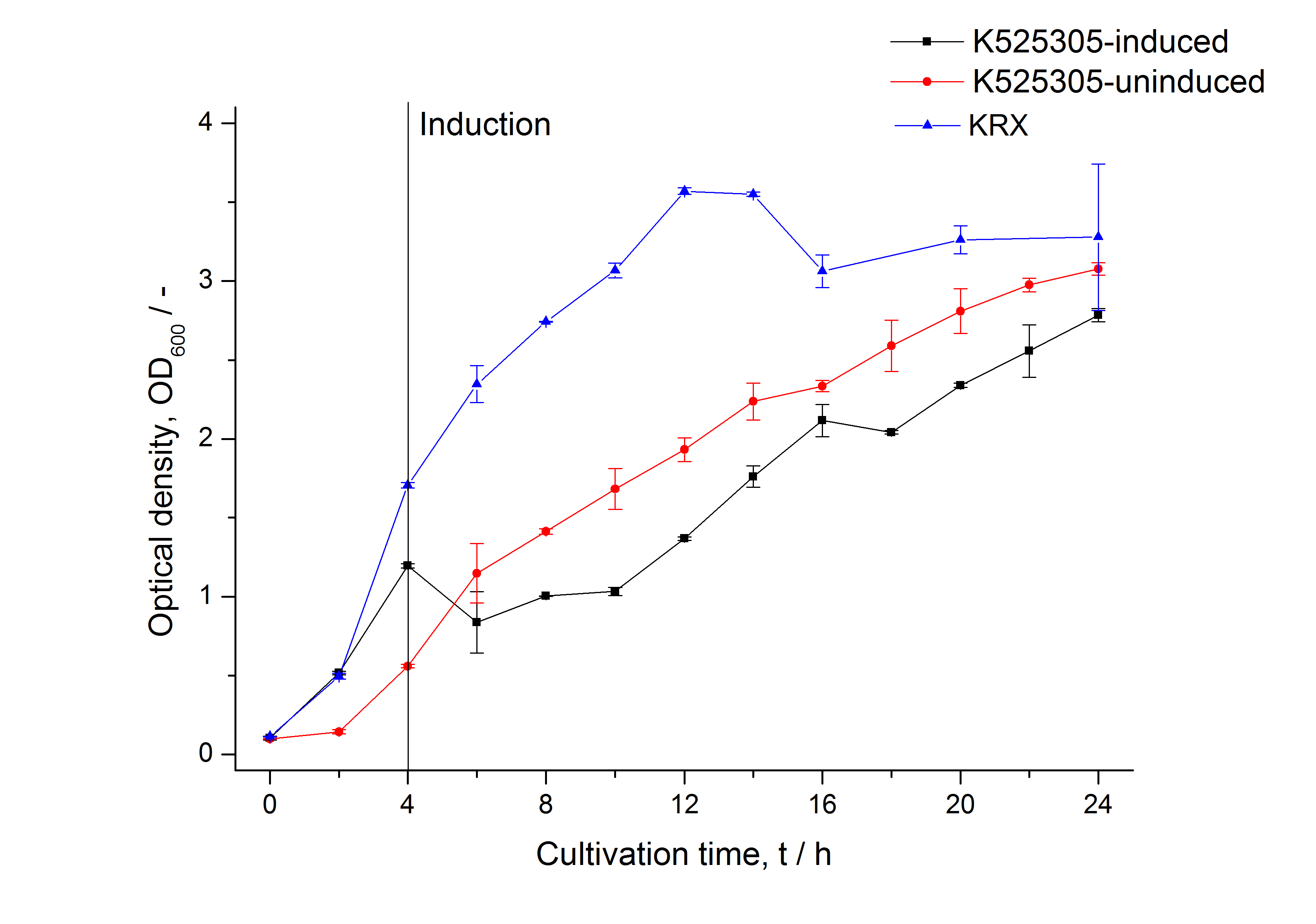
| Specific growth rate (un-/induced) | 0.139 h-1 / 0.071 h-1 | |
| Doubling time (un-/induced) | 4.98 h / 9.78 h | |
| Purification | Molecular weight | 110.2 kDa |
| Theoretical pI | 5.74 | |
| Excitation / emission | 515 / 529 nm | |
| Immobilization behaviour | Saturation protein / bead ratio | 5 - 7 * 10-4 |
| Immobilization time | 4 h |
Sequence and Features
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 167
Illegal BglII site found at 1022 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 76
Illegal AgeI site found at 3121 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 1657
Expression in E. coli
The SgsE (K525301) gen was fused with a mCitrine (BBa_J18931) using [http://2011.igem.org/Team:Bielefeld-Germany/Protocols#Gibson_assembly Gibson assembly] for characterization.
The SgsE|mCitrine fusion protein was overexpressed in E. coli KRX after induction of T7 polymerase by supplementation of 0,1 % L-rhamnose and 1 mM IPTG using the [http://2011.igem.org/Team:Bielefeld-Germany/Protocols/Downstream-processing#Expression_of_S-layer_genes_in_E._coli autinduction protocol] from promega.

Purification of SgsE fusion protein
As seen in the analysis of the cultivations with expression of SgsE | mCitrine fusion proteins, these proteins form inclusion bodies in E. coli. Inclusion bodies have the advantage that they are relatively easy to clean-up and are resistant to proteases. So the first purification step is to solve and set-free the inclusion bodies. This step is followed by two filtrations (300 kDa UF and 100 kDa DF/UF) to further concentrate and purify the S-layer proteins. After the filtrations, the remaining protein solution is dialized against ddH2 for 18 h at 4 °C in the dark. The dialysis leads to a precipitation of the water-insoluble proteins. After centrifugation of the dialysate the water-soluble S-layer monomers remain in the supernatant and can be used for recrystallization experiments.
The fluorescence of the collected fractions of this purification strategy is shown in the following figure A:

A lot of protein is lost during the purification especially after centrifugation steps. The fluorescence in the urea containing fractions is lowered due to denaturation of the fluorescent protein. Some fluorescence could be regenerated by the recrystallization in HBSS. This purification strategy is very simple and can be carried out by nearly everyone in any lab being one first step to enable real do it yourself nanobiotechnology.
Immobilization behaviour
After purification, solutions of monomeric SgsE S-layer proteins can be recrystallized and immobilized on silicon dioxide beads in HBSS (Hank's buffered saline solution). After the recrystallization procedure the beads are washed with and stored in ddH2O at 4 °C in the dark. The fluorescence of the collected fractions of a recrystallization experiment with BBa_K525305 are shown in fig. X. 100 mg beads were coated with 100 µg of protein. The figure shows, that not all of the protein is immobilized on the beads (supernatant fraction) but the immobilization is pretty stable (very low fluorescence in the wash). After the immobilization, the beads show a high fluorescence indicating the binding of the SgsE | mCitrine fusion protein.

Optimal bead to protein ratio for immobilization
To determine the optimal ratio of silica beads to protein for immobilization, the degree of clearance ϕC in the supernatant is calculated and plotted against the concentration of silica beads used in the accordant immobilization experiment (compare fig. A):
The data was collected in three indipendent experiments. The fluorescence of the samples was measured in the supernatant of the immobilization experiment after centrifuging the silica beads. The fluorescence of the control was measured in a sample which was treated exactly like the others but no silica beads were added. 100 µg protein was used for one immobilization experiment. The data was fitted with a sigmoidal dose-response function of the form
with the Hill coefficient p, the bottom asymptote A1, the top asymptote A2 and the switch point log(x0) (R² = 0.874).
The fit indicates that a good silica concentration for 100 µg of protein is 150 - 200 mg mL-1. This set-up leads to saturated beads with low waste of protein. So a good protein / bead ratio to work with is 5 - 7 * 10-4.

Methods
References
Kainz B, Steiner K, Möller M, Pum D, Schäffer C, Sleytr UB, Toca-Herrera JL (2010) Absorption, Steady-State Fluorescence, Fluorescence Lifetime, and 2D Self-Assembly Properties of Engineered Fluorescent S-Layer Fusion Proteins of Geobacillus stearothermophilus NRS 2004/3a, [http://pubs.acs.org/doi/abs/10.1021/bm901071b Biomacromolecules 11(1):207-214].
Sleytr UB, Huber C, Ilk N, Pum D, Schuster B, Egelseer EM (2007) S-layers as a tool kit for nanobiotechnological applications, [http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.2006.00573.x/full FEMS Microbiol Lett 267(2):131-144].
//function/reporter/fluorescence
//proteindomain/internal
| biology | S-Layer |
| chassis | E. coli |
| emission | 529 nm |
| excitation | 515 nm |
| origin | Geobacillus stearothermophilus |