Part:BBa_K864400
Ptac, trp & lac regulated promoter
The Ptac promoter is a functional hybrid promoter, derived from the trp and lac promoters, that are regulated by trp and lac [1]. This part also exist together with lacI, part BBa_K180000
[1] Proc. Natl. Acad. Sci. USA, Vol. 80, pp. 21-25
Improvement by Evry_Paris-Saclay 2017
This part has been improved to be the pPsiTac1, a strong psicose inducible promoter. For more information on the improved part, please go to the page of Part:BBa_K2448016.
Characterization by SHSBNU_China 2019
Description
In our design, the induction signal will be detected and stored in the plasmid DNA sequence of our genetically modified E. coli. The bacteria will be gathered from the capsule after it left human body and sent to lab for further quantitative analysis in order to represent the inflammation level in colon. Therefore, in our project, the fluorescence intensity will be the index that represents levels of gut inflammation.
Since then, we first needed to develop a standard quantitative measurement protocol, so we studied the following articles: Zong, Y et al (2017)[1] and Zhang, H. M. et al(2015).

Following reference, we determined the experimental procedure:
Flat-bottom 96-well plates and sealing film were used throughout the study. Bacteria harboring parts/circuits of interest were inoculated from plates to LB medium and grown overnight (8−12 h, 1000 rpm, 37 °C, mB100-40 Thermo Shaker). Ten microliters of each overnight culture was sequentially diluted into 130 μL of fresh medium twice; the total dilution fold was 196. After growing the diluted cultures for ∼3 h, we diluted the exponentially growing cultures 700-fold using fresh medium; the dilution process was as follows: 10 μL of cell culture is added to 130 μL of M9 medium, which is followed by diluting 3 μL of this into 147 μL. Then, cultivation continued (1000 rpm, 37 °C, mB100-40); atspecifictimepoints,a2−50μL aliquot of each culture was transferred to a new plate containing 200 μL of PBS with 2 mg/mL kanamycin preadded to terminate protein expression. For the time course of cell growth after 700-fold dilution, OD 600 was recorded using Varioskan Flash (Thermal Scientific); the time interval was 5 min.

The graph below shows our result.

Ptac, trp & lac regulated promoter</br>
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
1. pTAC_RFP characterization
Description
pTAC promoter was taken from part BBa_K864400. This promoter is a hybrid of two operons: the trp and lac operons. This promoter is inducible by IPTG and commonly used in Escherichia coli for overproduction of proteins. E. coli NM522 strain that we used in our lab constitutively produces the LacIq protein, a strong pTAC promoter repressor. However, in absence of IPTG, we observed a strong leakage when plating our BBa_K1934000 transformants. Therefore, we decided to put a RFP reporter ORF under control of the pTAC promoter to characterize the promoter-driven transcriptional noise.
Expression of the pTAC-RFP fusion in presence of increasing amount of the inductor IPTG
Experimental design
The RFP coding sequence (BBa_E1010) was placed in silico under the control of the pTAC promoter (BBa_K864400), a strong RBS (BBa_B0030) and a bidirectional terminator (BBa_B0011). IDT performed the DNA synthesis and delivered the part as gBlock. The construct was cloned by conventional ligation into pSB1C3 plasmid and then transformed into E. coli NM522 strain. In order to study the efficiency of the pTAC promoter for the overproduction of proteins, recombinant clones were grown overnight in LB at 37°C in duplicate in three different induction conditions (IPTG concentrations): 0 mmol.L-1, 1 mmol.L-1 and 5 mmol.L-1. OD600 of each culture was measured every hour over six hours. E. coli NM522 strain was grown overnight in LB at 37°C in the same three induction conditions as control.Results
- Noise of the pTAC: fluorescence in absence of IPTG
We studied the noise of the promoter by comparing the normalized fluorescence between the construction and the NM522 strain without any induction of IPTG.

An ANOVA was made to see if there was a time effect between the two populations. We obtained a p-value of 0.61, suggesting that time had no effect on pTAC-RFP expression in absence of IPTG (α<0.05). Given this result, we gathered data to analyze if there was a significant difference between the two strains. A Student test was performed with the variance not equal. The p-value of 5.44*10-4 indicated that the strain carrying pTAC-RFP transcriptional fusion displayed a higher fluorescence than the control strain.
- pTAC induction by increasing concentration of IPTG
Then we studied the induction of the promoter by comparing the normalized fluorescence of the construction under the induction of [IPTG] = 0 and 1 mmol.L-1.

An ANOVA test was made to see if there was a time effect between the two populations. A p-value of 0.21 indicated that time had no effect on the fluorescence induction (α<0.05). Data was therefore gathered in order to compare the strain fluorescence with 1 mmol.L-1 IPTG and no IPTG. We realized a Student test with the variance not equal and obtained a p-value of 8.57*10-3, showing a difference of fluorescence due to the presence of IPTG in the medium.
Finally, we compared the expression of the pTAC-RFP transcriptional fusion in 2 concentrations of IPTG: 1 and 5 mmol.L-1.

An ANOVA was made to see if there was a time effect between the two populations. A p-value of 0.06 indicated that time had no effect (α<0.05). From there, we could gather the data to analyze if there was a significant difference between the two concentrations of IPTG.
We realized a Student test with the variance not equal and obtained a p-value of 0.61, indicating that no significant difference of fluorescence was observed with the rise of IPTG concentration.
Conclusion
In the case of RFP, the pTAC promoter seemed to not enable a gene tune because statistics showed that there was a significant noise, even in the absence of IPTG.
2. pTAC_CFP characterization
Description
pTAC promoter was taken from part part BBa_K864400 . This promoter is a hybrid of two operons: the trp and lac operons.This promoter is inducible by IPTG and commonly used in Escherichia coli for overproduction of proteins. Escherichia coli NM522 strain that we used in our lab constitutively express the LacIq protein, a strong pTAC promoter repressor. However, in absence of IPTG, we observed a strong leakage when plating our BBa_K1934000 transformants. Therefore, we decided to put a CFP reporter ORF under control of the pTAC promoter to characterize the promoter-driven transcriptional noise.
Expression of the pTAC-CFP fusion in presence of increasing amount of the inductor IPTG
Experimental design
The CFP coding sequence (BBa_E2020) was placed in silico under the control of the pTAC promoter (BBa_K864400), a strong RBS (BBa_B0030) and a bidirectional terminator (BBa_B0011). IDT performed the DNA synthesis and delivered the part as gBlock. The construct was cloned by conventional ligation into pSB1C3 plasmid and then transformed into E. coli NM522 strain. In order to study the efficiency of the pTAC promoter for the overproduction of proteins, recombinant clones were grown overnight in LB at 37°C in duplicate in three different induction conditions (IPTG concentrations): 0 mmol.L-1, 1 mmol.L-1 and 5 mmol.L-1. OD600 of each culture was measured every hour over six hours. E. coli NM522 strain was grown overnight in LB at 37°C in the same three induction conditions as control.Results
- Noise of the pTAC: fluorescence in absence of IPTG
We studied the noise of the promoter by comparing the normalized fluorescence between the construction and the NM522 strain without any induction of IPTG.

An ANOVA was made to see if there was a time effect between the two populations. We obtain a p-value of 0.96, suggesting that time had no effect on pTAC-CFP expression in absence of IPTG (α<0.05). Given this result, we gathered data to analyze if there was a significant difference between the two strains. A Student test was performed with the variance not equal. The p-value of 0.10 indicated that the strain carrying pTAC-CFP transcriptional fusion didn’t display a significant fluorescence difference with the control strain.
- pTAC induction by increasing concentration of IPTG
Then we studied the induction of the promoter by comparing the normalized fluorescence of the construction under the induction of [IPTG] = 0 and 1 mmol.L-1.

An ANOVA test was made to see if there was a time effect between the two populations. A p-value of 0.05 indicated that time had no effect on the fluorescence induction (α<0.05). Data was therefore gathered in order to compare the strain fluorescence with 1 mmol.L-1 IPTG and no IPTG. We realized a Student test with the variance not equal and obtained a p-value of 4.64*10-10, showing a significant difference of fluorescence due to the presence of IPTG in the medium.
Finally, we compared the expression of the pTAC-CFP transcriptional fusion in 2 concentrations of IPTG: 1 and 5 mmol.L-1.

An ANOVA was made to see if there was a time effect between the two populations. A p-value of 0.55 indicated that time had no effect (α<0.05). From there, we gathered data to analyze if there was a significant difference between the two concentrations of IPTG. We realized a Student test with the variance not equal and obtained a p-value of 0.54, indicating that no significant difference of fluorescence was observed with the rise of IPTG concentration.
Conclusion
In the case of CFP, the pTAC promoter seemed to enable a gene tune because there wasn’t a differential gene expression in absence of IPTG, so a significant noise wasn’t measured.
Team JUN_China 2019: Fluorescent characterization of the Tac promoter
Tac promoter is a hybrid of two operons: the trp and lac operons and it is inducible by IPTG.In our project, we built a number of parts which also contains a large number of construct intermediates using the tac promoter. The tac promoter is critical in our entire project, so we used green and red fluorescent protein to characterize its effectiveness. At the same time, we explored the changes in expression intensity of green and red fluorescent protein under the control of tac promoter by IPTG inducing at different times.
Experiment design:
We ligated the eGFP and RFP gene into the loop vector pZM1 (Ptac) containing the inducible tac promoter respectively and transfered eGFP and RFP into our chassis microorganism: Corynebacterium glutamicum F343 to carry out fermentation experiment. We added 1mmol/L IPTG to the fermentation broth at 0h, 1h, and 2h respectively and measured the fluorescence intensity and cell growth at different times to investigate whether the inducing capability of tac promoter was different at different induction times.
During the detection process, the cells were washed using PBS and the cells were diluted by appropriate multiples according to the measurement range of the instrument. The growth state and fluorescence intensity of the cells were detected using 96-well plates. The green fluorescent protein has an excitation wavelength of 488 nm, an emission wavelength of 517 nm. The red fluorescent protein excitation wavelength is 560 nm, and the emission wavelength is 630 nm.
Result:
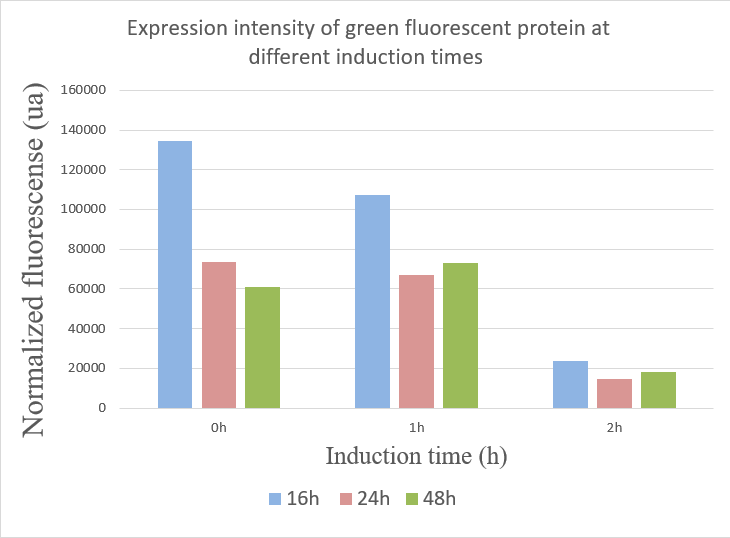
We used the infinite 200Pro instrument to measure green and red fluorescent protein and OD600 measurement. The relevant data are shown below:
The data showed:
1.When inducing at 0 hour, the highest fluorescence value were at 16 hours, and there was a gradual decrease in data from 24 hours to 48 hours. This may due to the degradation of fluorescent protein.
2.The fluorescence values induced at 1 hour were relatively stable, but when at 16 hours the fluorescence value was best.
3.The expression of GFP was the worst when inducing at 2 hour.
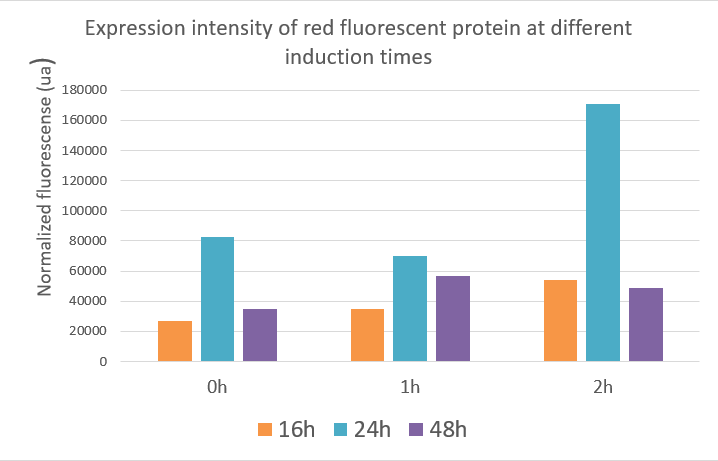
Red fluorescent protein data showed that the expression of the red fluorescent protein was best when inducing at two hours and the value reached about 17,000 ua at 24 hours. The optimal induction time of red fluorescent protein expression was the same as the optimal induction time of polymerase in our project, which was induced at two hours after fermentation, and the product expression was highest. The data indicated that induction at different times, the expression of the target gene and the yield of the product were different.
//direction/forward
//promoter
//regulation/positive
//rnap/prokaryote/ecoli/sigma70
| control | trp, lac |
| direction | forward |
| n/a | Hybrid promoter (trp & lac regulated -- tac pR) |