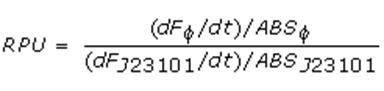Part:BBa_K173007:Experience
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
|
•••••
Michigan iGEM 2013 |
This part design works very well. See K1077002 for our characterization of it in the context of the fim transcriptor. We did not use this part specifically, but used the same parts. |
Applications of BBa_K173007
2018 iGEM team Linkoping Sweden
2018 iGEM team Linkoping Sweden validated this part.
Wells in the gel is named A, B, C and D. In well A contains an induction of GroES with tetracycline 200ng/ml and nothing was added to well B, what we see there is leakage from adjacent. Well C has no induction of GroES, well D shows E.coli (BL21) without the GroES plasmid and finally well E contains a BLUeye Prestained Protein ladder frpm Sigma Aldrich.
Seen in fig 1, well A contains a very clear band in the correct location for the protein induced (GroES) with a much stronger intensity than the both uninduced variants.

BBa_K173007 - aTc inducible device with J23100 promoter - UNIPV-Pavia team (test performed by L.Pasotti, S.Zucca)
Description
This is an aTc sensing device.
BBa_J23100 promoter drives the constitutive production of tetR repressor (BBa_C0040), which inhibits tetR promoter (BBa_R0040) activity. When aTc is added to the medium, it binds tetR and inhibits it. So, the PoPS output is a function of the aTc concentration.
A tight regulation is expected for this inducible system because BBa_J23100 is a strong promoter and so tetR repressor should be produced at extremely high levels.
The data below are referred to BBa_K173009, which is the measurement system of BBa_K173007.
Characterization
Compatibility: E. coli TOP10 in pSB1A2
| aTc concentration [ng/ml] |
LB | M9 supplemented | ||
| Doubling time [minutes] | RPU mean [range] | Doubling time [minutes] | RPU mean [range] | |
| 0 | 36 | 0.05 [~0 ; 0.15] | 59 | 0.03 [ 0.02 ; 0.04] |
| 25 | 32 | 0.12 [~0 ; 0.19] | 61 | 0.03 [0.02; 0.04] |
| 50 | 34 | 0.17 [0.05; 0.25] | 57 | 0.04 [0.03; 0.05] |
| 75 | 43 | 0.15 [~0; 0.23] | 65 | 0.03 [0.02; 0.06] |
| 100 | 40 | 0.21 [0.19; 0.23] | 62 | 0.03 [0.03; 0.05] |
| 200 | 40 | 0.25 [0.20; 0.28] | 70 | 0.04 [0.03; 0.06] |
| 300 | 43 | 0.20 [0.18; 0.21] | 67 | 0.05 [0.04; 0.08] |
 BBa_K173009 Growth curves for BBa_K173009 in M9 |
 BBa_K173009 Growth curves for BBa_K173009 in LB |
 BBa_K173009(dGFP/dt)/OD in LB |
 BBa_K173009(dGFP/dt)/OD in M9 |
 BBa_K173009 Induction curve of BBa_K173009 in M9 |
 BBa_K173009 Induction curve of BBa_K173009 in LB |
Conclusions
We demonstrated that this part works as expected, sensing the aTc concentration provided in the culture medium. The transfer function of this sensor has been characterized in standard units (RPUs) in two different growth media (LB and M9 supplemented with glycerol), as well as the metabolic burden (in terms of doubling time) which affects E. coli bearing this part.
On the other hand, we did not expect to have a higher GFP synthesis rate per cell after the exponential growth phase than in the exponential phase itself (as reported in the 3rd plot for both LB and M9).
This part shows to have a very low leakage rate (about 0.025 RPU) but also a very low transcription rate for high concentrations of aTc. So, it can be used for tight regulation, in those systems who need very low leakage rates (no gene expressed in absence of inducer) but not for massive protein production.
Unfortunately, the estimated transfer function is very noisy (probably because the measured fluorescence values are very close to blank values), and further experiments should be performed in order to validate these induction curves.
Growth conditions
Microplate reader experiments
- 8 ul of long term storage glycerol stock were inoculated in 5 ml of LB + suitable antibiotic in a 15 ml falcon tube and incubated at 37°C, 220 rpm for about 16 hours.
- The grown cultures were then diluted 1:100 in 5 ml of LB or M9 supplemented medium and incubated in the same conditions as before for about 4 hours.
- These new cultures were diluted to an O.D.600 of 0.02 (measured with a TECAN F200 microplate reader on a 200 ul of volume per well; it is not comparable with the 1 cm pathlength cuvette) in a sufficient amount of medium to fill all the desired microplate wells.
- These new dilutions were aliquoted in a flat-bottom 96-well microplate, avoiding to perform dynamic experiments in the microplate frame (see [http://2009.igem.org/Team:UNIPV-Pavia/Methods_Materials/Evaporation Frame effect section] for details). All the wells were filled with a 200 ul volume.
- If required, 2 ul of inducer were added to each single well.
- The microplate was incubated in the Tecan Infinite F200 microplate reader and fluorescence (when required) and absorbance were measured with this automatic protocol:
- 37°C constant for all the experiment;
- sampling time of 5 minutes;
- fluorescence gain of 50;
- O.D. filter was 600 nm;
- GFP filters were 485nm (ex) / 540nm (em);
- 15 seconds of linear shaking (3mm amplitude) followed by 10 seconds of waiting before the measurements in order to make a homogeneous culture.
- Variable experiment duration time (from 3 to 24 hours).
Data analysis
Growth curves
All our growth curves have been obtained subtracting for each time sample the broth O.D.600 measurement from that of the culture; broth was considered in the same conditions of the culture (e.g. induced with the same inducer concentration).
Doubling time
The natural logarithm of the growth curves (processed according to the above section) was computed and the linear phase (corresponding to the bacterial exponential growth phase) was isolated by visual inspection. Then the linear regression was performed in order to estimate the slope of the line m. Finally the doubling time was estimated as d=ln(2)/m [minutes].
In the case of multiple growth curves for a strain, the mean value of the processed curves was computed for each time sample before applying the above described procedure.
Relative Promoter Units (RPUs)
The RPUs are standard units proposed by Kelly J. et al., 2008, in which the transcriptional strength of a promoter can be measured using a reference standard, just like the ground in electric circuits.
RPUs have been computed as:
in which:
- phi is the considered promoter and J23101 is the reference standard promoter (taken from Anderson Promoter Collection);
- F is the blanked fluorescence of the culture, computed subtracting for each time sample fluorescence measure for negative control from that of culture, where the negative control is a non-fluorescent strain (in our experiment it is usually used TOP10 strain bearing BBa_B0032 or BBa_B0033, which are symmply RBSs do not have expression systems for reporter genes);
- ABS is the blanked absorbance (O.D.600) of the culture, computed as described in "Growth curves" section.
RPU measurement has the following advantages (under suitable conditions)
- it is proportional to PoPS (Polymerase Per Second), a very important parameter that expresses the transcription rate of a promoter;
- it uses a reference standard and so measurements can be compared between different laboratories.
The hypotheses on which RPU theory is based can be found in Kelly J. et al., 2008, as well as all the mathematical steps. From our point of view, the main hypotheses that have to be satisfied are the following:
- the reporter protein must have a half life higher than the experiment duration (we use GFPmut3, BBa_E0240, which has an estimated half life of at least 24 hours, and the experiments duration is always less than 7 hours);
- strain, plasmid copy number, antibiotic, growth medium, growth conditions, protein generator assembled downstream of the promoter must be the same in the promoter of interest and in J23101 reference standard.
- steady state must be valid, so (dF/dt)/ABS (proportional to the GFP synthesis rate per cell) must be constant.
Inducible systems
Every experiment is performed on the following cultures:
- the culture of interest (system studied expressing GFP)
- the benchmarck used to evaluate R.P.U. (BBa_K173001 measurement part, that is BBa_J23101 with BBa_E0240 downstream)
- a negative control (generally, BBa_B0033 RBS)
For inducible systems several plots are reported. The first plot is a panel containing 4 subplots, numerated this way:
| (1) | (2) |
| (3) |
Plot (1) contains growth curves of the cultures, after blank value has been removed. Every curve is calculated averaging on three replicates of the same culture and subtracting the blank for each time sample. Blank is calculated averaging the replicates of blank wells.
Plot (2) shows the logarithm of absorbance in exponential phase of bacterial growth, determined by a visual inspection of log-plots. These values are used to evaluate doubling time and R.P.U..
Plot (3) contains (dGFP/dt)/O.D., the value named S_cell in Kelly J. et al., 2008 procedure for RPU evaluation.
In these plots are reported black veritcal lines that define the range of values used to evaluate RPU. It is important to underline, as explained in next paragraph, that RPU are calculated on cultures at the same O.D. level, not at the same time.
The second graphic shows S_cell VS O.D.. This plot allows the conparison of S_cell values between different cultures, that are supposed to reach the same level of growth not at the same time, but at the same O.D. value.
The third graphic shows the induction curve. The RPU value is calculated on S_cell values corresponding to O.D. values in exponential phase (typically, from 0.05 to 0.16). The curve is obtained averaging in time S_cell values corresponding to exponential phase.
Error bars rapresent the minimum and maximum value of R.P.U. belonging to the range of O.D. in exponential phase.
Materials
- Long term glycerol stocks were stored at -80°C with a final glycerol concentration of 20%
- Antibiotics were: Ampicillin (Amp) 100 ug/ml, stored at -20°C in 1000x stocks. Amp was dissolved in water.
- LB medium was prepare with: 1% NaCl, 1% bactotryptone, 0.5% yeast extract. The medium was not buffered with NaOH.
- M9 supplemented medium was prepared according to: [http://openwetware.org/wiki/Knight:M9_supplemented_media Openwetware protocol].
- aTc (Clontech) was dissolved in ethanol 50% and stored at -20°C in a 100 ug/ml stock. All the following dilutions were performed in water.
User Reviews
UNIQ2d9dff3a899cf6d1-partinfo-00000015-QINU UNIQ2d9dff3a899cf6d1-partinfo-00000016-QINU