Part:BBa_K525562
Fusion protein of NADP+ Oxidoreductase and BisdA and BisdB with middle strong promoter,RBS
Fusion protein of ferredoxin-NADP+ oxidoreductase, BisdA and BisdB; RFC 25 (Freiburg BioBrick assembly standard) for characterization of intra- and extracellular BPA degradation.
Usage and Biology
Expressing this BioBrick in E. coli enables the bacterium to degrade the endocrine disruptor bisphenol A (BPA).
BPA is mainly hydroxylated into the products 1,2-Bis(4-hydroxyphenyl)-2-propanol and 2,2-Bis(4-hydroxyphenyl)-1-propanol. In S. bisphenolicum AO1, a total of three genes are responsible for this BPA hydroxylation: a cytochrome P450 (CYP, bisdB), a ferredoxin (Fd, bisdA) and a ferredoxin-NAD+ oxidoreductase (FNR) Sasaki05a. The three gene products act together to reduce BPA while oxidizing NADH + H+. The cytochrome P450 (BisdB) reduces the BPA and is oxidized during this reaction. BisdB in its oxidized status is reduced by the ferredoxin (BisdA) so it can reduce BPA again. The oxidized BisdA is reduced by a ferredoxin-NAD+ oxidoreductase consuming NADH + H+ so the BPA degradation can continue Sasaki05a. This electron transport chain between the three enzymes involved in BPA degradation and the BioBricks needed to enable this reaction in vivo and in vitro are shown in the following figure (please have some patience, it's an animated .gif file):
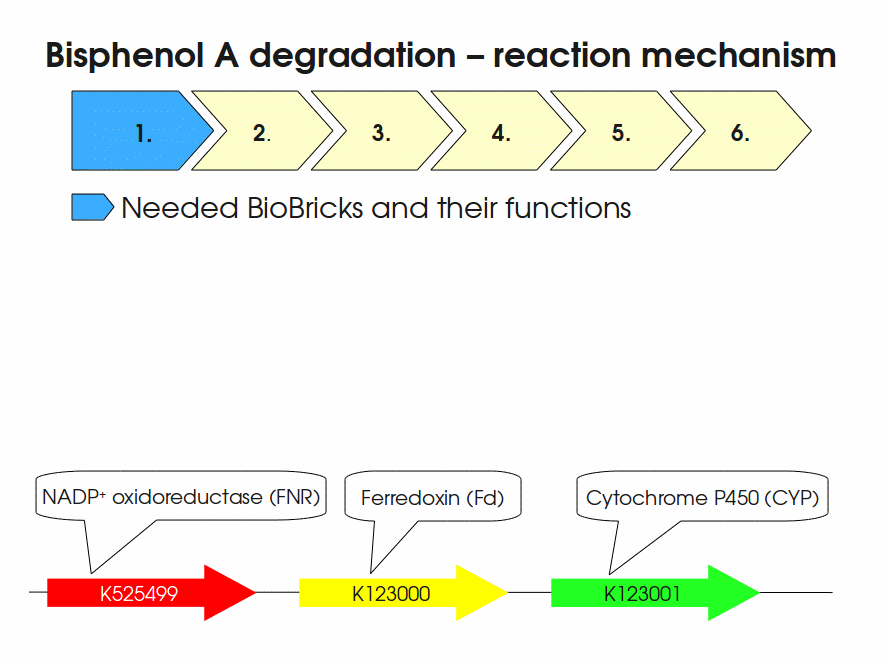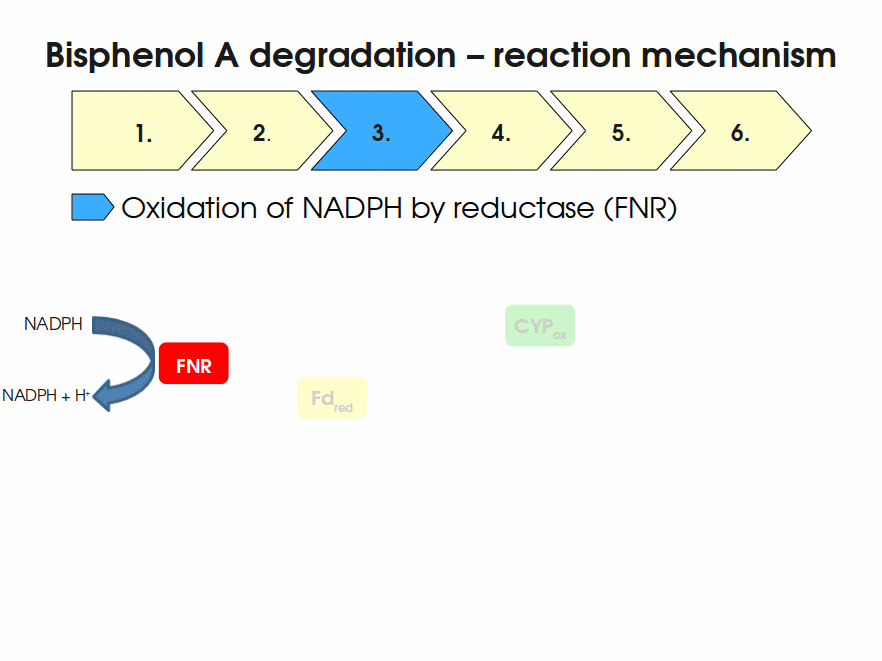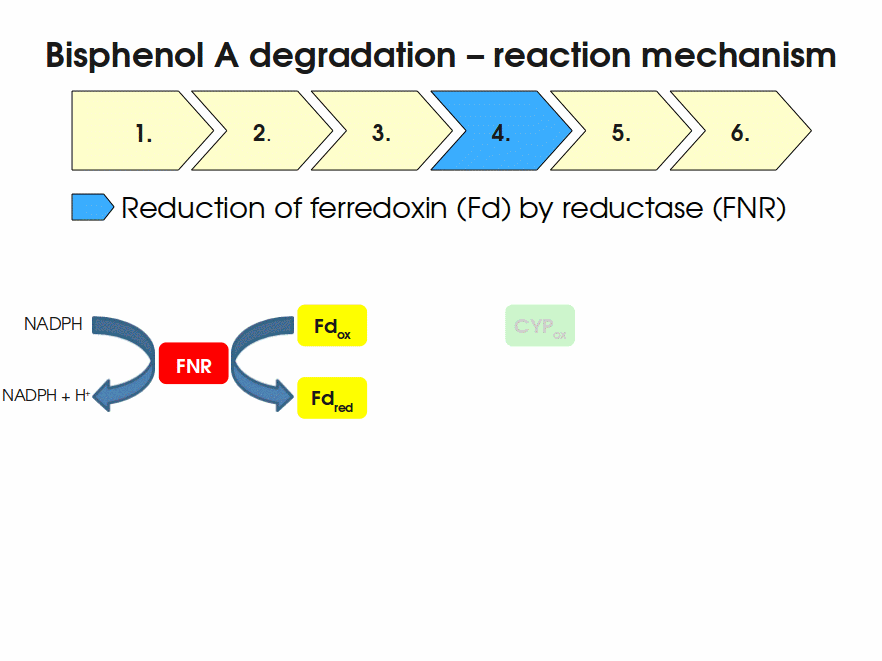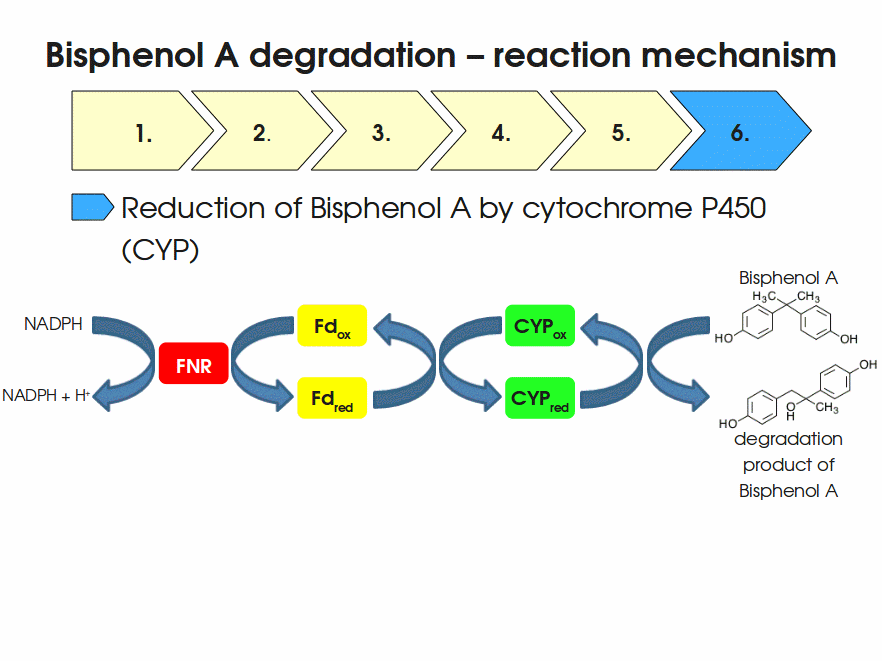
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 1411
Illegal BamHI site found at 2149 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 65
Illegal AgeI site found at 2402 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 746
//function/degradation/bisphenol
//proteindomain/internal
| chassis | E. coli |
| origin | Sphingomonas bisphenolicum AO1 |
