Part:BBa_K863006
ecol laccase from E. coli
E.coli laccase ORF
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 225
- 1000COMPATIBLE WITH RFC[1000]
Usage and Biology
In the last few years a lot of attention has been drawn to laccases due to their ability to oxidize both phenolic and nonphenolic lignin related compounds as well as highly recalcitrant environmental pollutants. This makes them very useful for applications concerning several biotechnological processes. This includes the detoxification of industrial effluents, for example from the paper and pulp, textile and petrochemical industries. Laccases are also valuable as a tool as a tool for medical diagnostics and as a bioremediation agent to clean up herbicides, pesticides and certain explosives in soil. Furthermore these enzymes are also used as catalysts for the manufacture of anti-cancer drugs and even as ingredients in cosmetics[1].
Their capacity to remove xenobiotic substances and produce polymeric products makes them a useful tool for bioremediation purposes. In our project laccases are used as cleaning agents for a water purification system.
Laccases are copper-containing polyphenol oxidase enzymes (EC 1.10.3.2) that can be found in many plants, insects, microorganisms and mainly in fungi. These enzymes fulfill several functions in different metabolic pathways.
Laccases are able to oxidize a broad range of substrates due to the contained copper-cluster, by reducing oxygen to water. The active site of the enzyme includes a four-copper-ion-cluster, which can be distinguished by spectroscopic analyses. This cluster consists of one blue copper-ion (type 1), one type 2 and two type 3 copper-ions. Because of the blue copper-ion, the laccases belong to the big family of the blue copper proteins. This specific blue copper ion is essential for the enzyme mediated radical oxidation of the phenolic groups. In this reaction the electron from the oxidation is transferred to the other three copper ions. These ions form a trinuclearic cluster, which transfers electrons to the terminal electron acceptor oxygen. By receiving four electrons the molecular oxygen is finally reduced to water.
[1] Susana Rodríguez Couto & José Luis Toca Herrera;Industrial and biotechnological applications of laccases: A review; 2006; Biotechnology Advances 24 500–513
Contents
Cultivation, Purification and SDS-PAGE
Shaking Flask Cultivations
The first trials to produce ECOL were produced in shaking flask with various designs (from 100 mL-1 to 1 L flasks, with and without baffles) and under different conditions. The parameters tested during our screening experiments were temperature (27 °C,30 °C and 37 °C), concentrations of chloramphenicol (20-170 µg mL-1), various induction strategies ([http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Materials#Autoinduction_medium autoinduction] and manual induction) and cultivation time (6 - 24 h). Furthermore it was cultivated with and without 0.25 mM CuCl2 to provide a sufficient amount of copper, which is needed for the active center of the laccase. Based on the screening experiments we identified the best conditions under which ECOL was expressed. The addition of CuCl2 did not increase the activity, so it was omitted.
- flask design: shaking flask without baffles
- medium: [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Materials#Autoinduction_medium autoinduction medium]
- antibiotics: 60 µg mL-1 chloramphenicol
- temperature: 37 °C
- cultivation time: 12 h
The reproducibility of the measured data and results were investigated for the shaking flask and bioreactor cultivation.
3 L Fermentation E. coli KRX with BBa_K863005
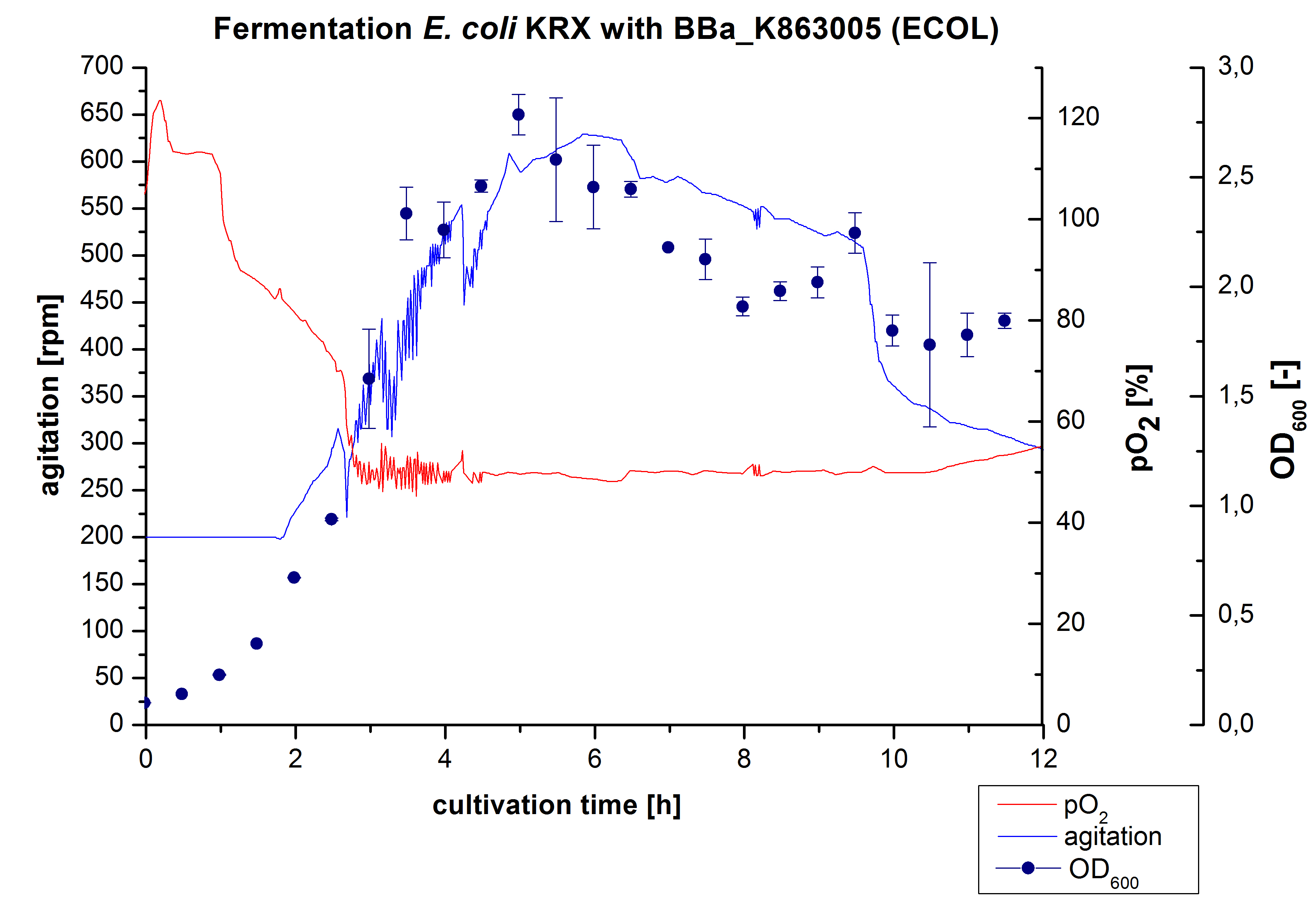
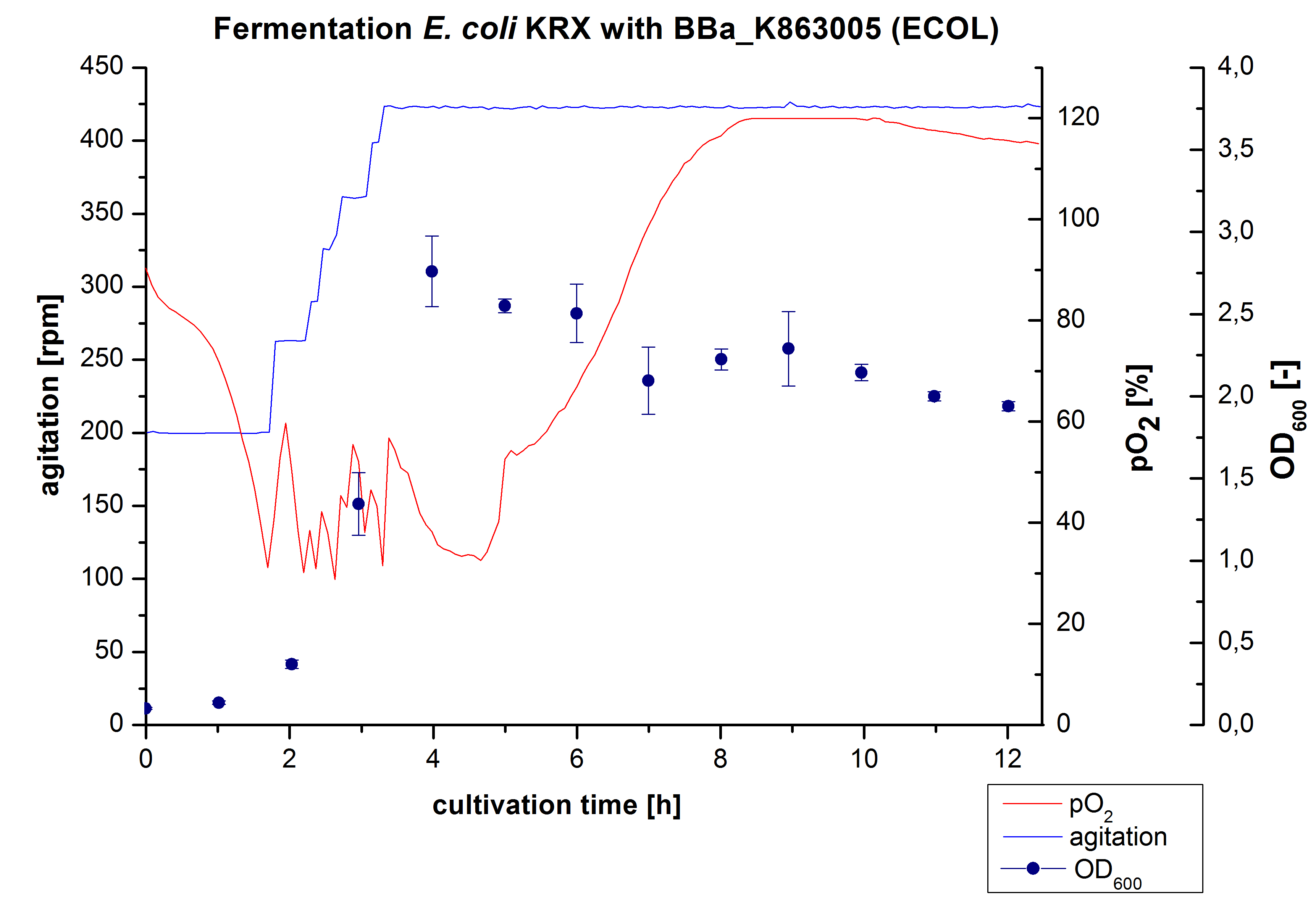
After the positive measurement of activity of ECOL we made a scale-up and fermented E. coli KRX with BBa_K863005 in an Infors Labfors fermenter with a total volume of 3 L. Agitation speed, pO2 and OD600 were determined and illustrated in Figure 1. The exponential phase started after 1.5 hours of cultivation. The cell growth caused a decrease in pO2. After 2 hours of cultivation the agitation speed increased up to 629 rmp (5.9 hours) to hold the minimal pO2 level of 50 %. Then, after 4 hours there was a break in cell growth due to induction of protein expression. The maximal OD600 of 2.78 was reached after 5 hours. In comparison to E. coli KRX (OD600,max =4.86 after 8.5 hours) and to E. coli KRX with BBa_K863000 (OD600,max =3.53 after 10 hours, time shift due to long lag phase) the OD600 max is lower. In the following hours, the OD600 and the agitation speed decreased and the pO2 increased, which indicates the death phase of the cells. This is caused by the cell toxicity of ECOL (reference: [http://www.dbu.de/OPAC/ab/DBU-Abschlussbericht-AZ-13191.pdf DBU final report]). Hence, cells were harvested after 12 hours.
Purification of ECOL
The harvested cells were resuspended in [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA- equilibration buffer], mechanically disrupted by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Production#Mechanical_lysis_of_the_.28bio-reactor.29_cultivation homogenization] and cell debris were removed by centrifugation. The supernatant of the cell lysate was loaded on the Ni-NTA column (15 mL Ni-NTA resin) with a flow rate of 1 mL min-1 cm-2. Then the column was washed with 10 column volumes (CV) [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA equilibration buffer]. The bound proteins were eluted by an increasing [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA elution buffer] step elution from 5 % (equates to 25 mM imidazol) with a length of 50 mL, to 50 % (equates to 250 mM imidazol) with a length of 60 mL, to 80 % (equates to 400 mM imidazol) with a length of 40 mL and finally to 100 % (equates to 500 mM imidazol) with a length of 80 mL. This strategy was chosen to improve the purification caused by a step by step increasing Ni-NTA-elution buffer concentration. The elution was collected in 10 mL fractions. Due to the high UV-detection signal of the loaded samples and to simplify the illustration of the detected product peak only the UV-detection signal of the wash step and the elution are shown. A typical chromatogram of purified laccases is illustrated here. The chromatogram of the ECOL elution is shown in Figure 2:
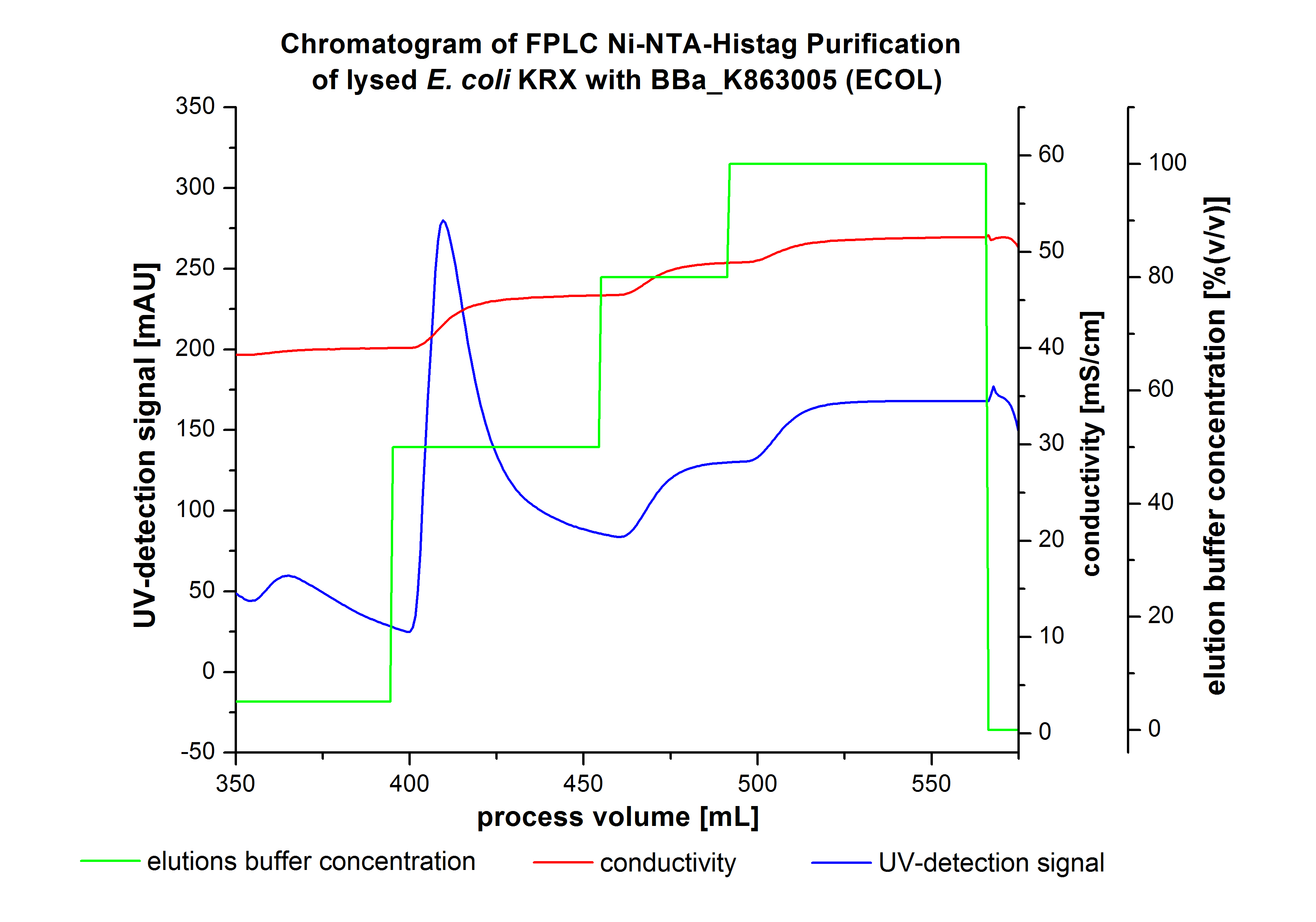
The chromatogram shows two distinguished peaks. The first peak was detected at a [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA-equilibration buffer] concentration of 5 % (equates to 25 mM imidazol) and resulted from the elution of weakly bound proteins. After increasing the Ni-NTA elution buffer concentration to 50 % (equates to 250 mM imidazol), an UV-detection signal peak of 292 mAU was measured. The area of this peak indicates that a high amount of protein was eluted. The corresponding fractions were analyzed by SDS-PAGE to detect ECOL. There were no further peaks detectable. The following increasing UV detection signal results from the rising imidazol concentration of the Ni-NTA elution buffer. The corresponding SDS-PAGES are shown in Figure 3.
SDS-PAGE of ECOL purification
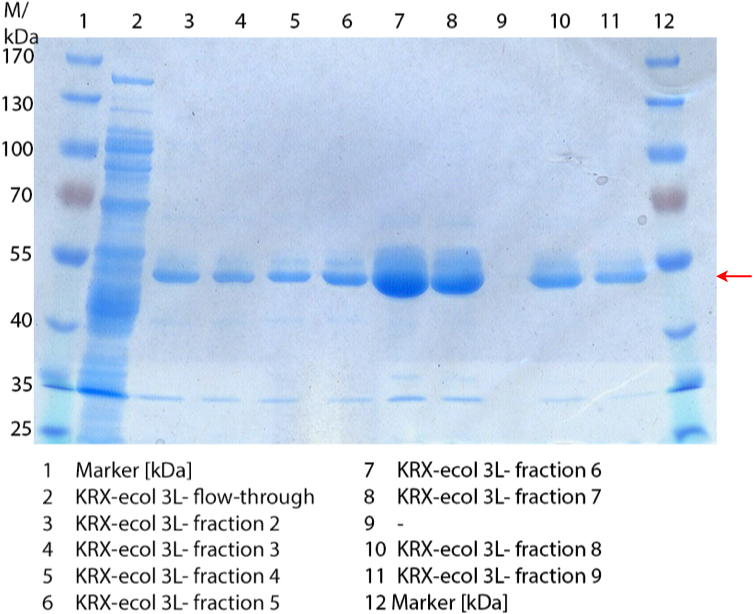
In Figure 3 the SDS-PAGE of the Ni-NTA His tag purification of the lysed culture (E. coli KRX containing BBa_K863005) is shown including the flow-through and the fractions 2 to 9. The red arrow indicates the band of ECOL with a molecular weight of 53.4 kDa, which appears in all fractions. The strongest bands appear in fractions 6 and 7. These were the first two fractions (each 10 mL) eluted with 50 % Ni-NTA elution buffer (equates to 250 mM imidazol), in which the distinguished peak appeared.
These bands were analyzed by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#MALDI MALDI-TOF] and identified as CueO (ECOL). In contrast, the second, faint band with a lower molecular weight could not be identified.
6 L Fermentation of E. coli KRX with BBa_K863005
Another scale-up of the fermentation of E. coli KRX with BBa_K863005 was made up to a final working volume of 6 L in a Bioengineering NFL 22 fermenter. Agitation speed, pO2 and OD600 were determined and illustrated in Figure 3. There was no noticeable lag phase and the cells immediately began to grow. The cells were in an exponential phase between 2 and 4 hours of cultivation, which results in a decrease of pO2 value and therefore in an increase of agitation speed. After 4 hours of cultivation the maximal OD600 of 2.76 was reached, which is comparable to the 3 L fermentation of E. coli KRX with BBa_K863005. Due to induction of protein expression there is a break in cell growth. The death phase started, which is indicated by an increasing pO2 and a decreasing OD600. This demonstrates the cytotoxicity of the laccase for E. coli, which was reported by the [http://www.dbu.de/OPAC/ab/DBU-Abschlussbericht-AZ-13191.pdf DBU]. In comparison to the fermentation of E. coli KRX with BBa_K863000 under the same conditions (OD600,max= 3.53), the OD600,max was lower. Cells were harvested after 12 hours.
Purification of ECOL
The harvested cells were resuspended in [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA-equilibration buffer], mechanically disrupted by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Production#Mechanical_lysis_of_the_.28bio-reactor.29_cultivation homogenization] and cell debris were removed by centrifugation. The supernatant of the cell lysate was loaded on the Ni-NTA column (15 mL Ni-NTA resin) with a flow rate of 1 mL min-1 cm-2. The column was washed by 10 column volumes (CV) [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA- equilibration buffer]. The bound proteins were eluted by an increasing [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA- elution buffer] gradient from 0 % to 100 % with a length of 200 mL and the elution was collected in 10 mL fractions. Due to the high UV-detection signal of the loaded samples and to simplify the illustration of the detected product peak only the UV-detection signal of the wash step and the eluate are shown. A typical chromatogram of purified laccases is shown here. The chromatogram of the ECOL elution is shown in Figure 5:
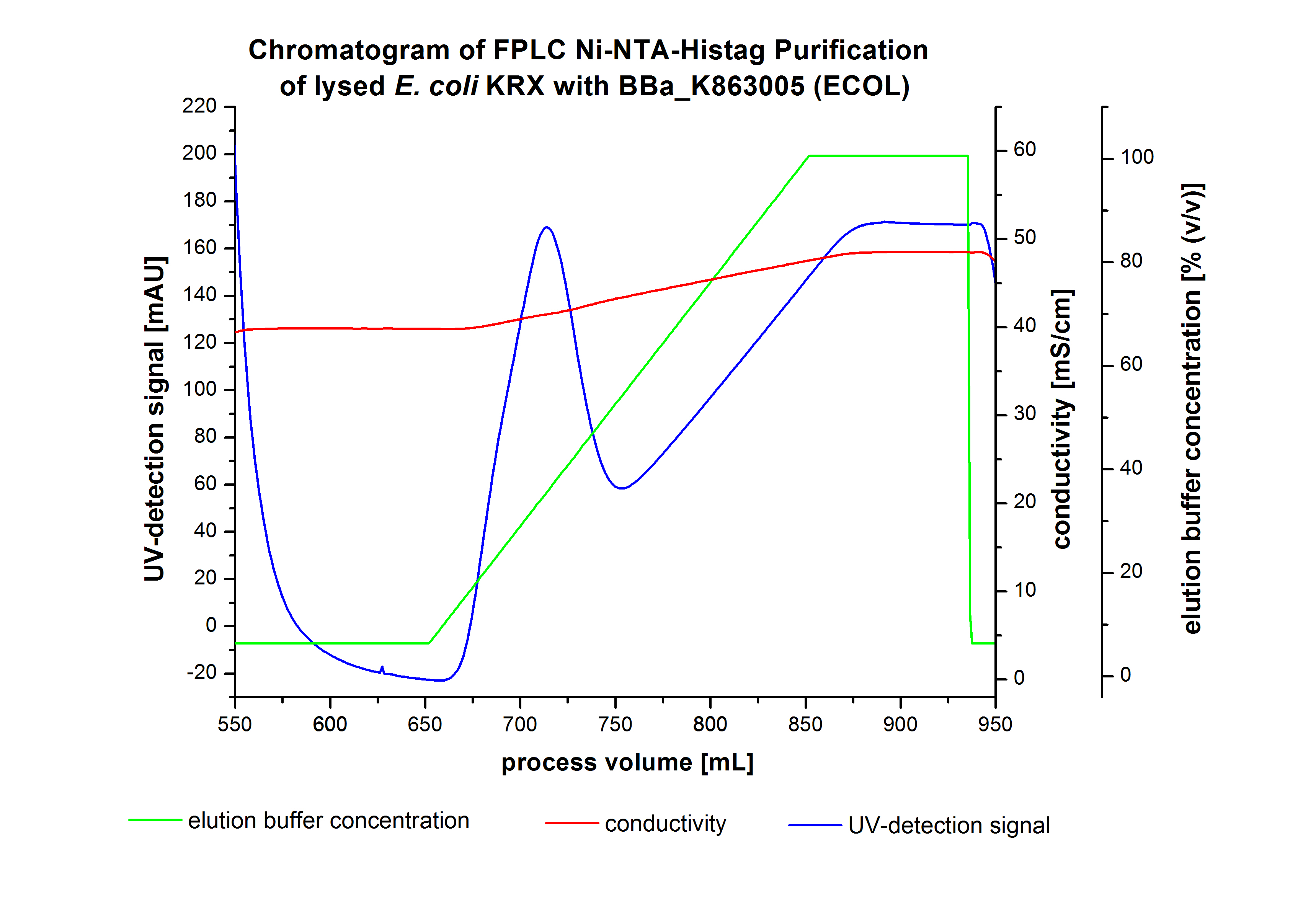
After washing the column with 10 CV [http://2012.igem.org/wiki/index.php?title=Team:Bielefeld-Germany/Protocols/Materials#Buffers_for_His-Tag_affinity_chromatography Ni-NTA-elution buffer] the elution process was started. At a process volume of 670 mL to 750 mL the chromatogram shows a remarkable widespread peak (UV-detection signal 189 mAU) caused by the elution of a high amount of proteins. The run of the curve show a fronting. This can be explained by the elution of weakly bound proteins, which elutes at low imidazol concentrations. A better result could be achieved with a step elution strategy ([http://2012.igem.org/Team:Bielefeld-Germany/Results/Summary#Purification_of_ECOL see purification of the 3 L Fermentation above]). To detect ECOL the corresponding fractions were analyzed by SDS-PAGE.
SDS-PAGES of ECOL purification
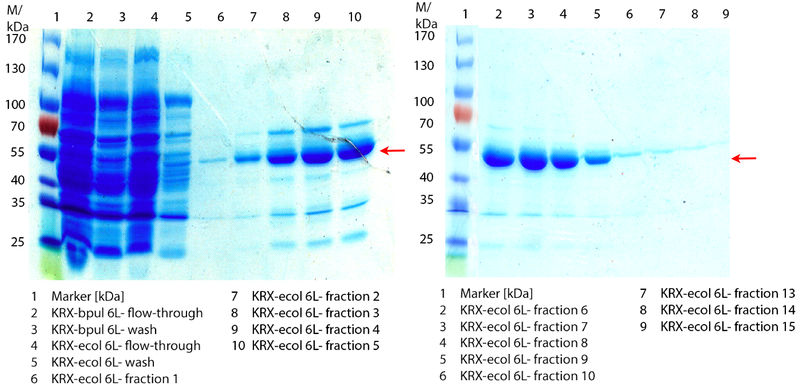
In Figure 6 the SDS-PAGE of the Ni-NTA His tag purification of the lysed culture E. coli KRX containing BBa_K863005 (6 L fermentation) including the flow-through, wash and the fractions 1 to 15 (except from fraction 11/12) is shown. The red arrow indicates the band of ECOL with a molecular weight of 53.4 kDa, which appears in all fractions. The strongest bands appear from fractions 3 and 8 with a decreasing amount of other non-specific bands. In summary, the scale up was successful, improving protein production and purification once again.
Furthermore the bands were analyzed by [http://2012.igem.org/Team:Bielefeld-Germany/Protocols/Analytics#MALDI MALDI-TOF] and identified as CueO (ECOL).
MALDI-TOF Analysis of ECOL
The E. coli laccase was identified using the following software
- FlexControl
- Flexanalysis and
- Biotools
Activity Analysis of ECOL
Initial activity tests of purified fractions
Initial tests were done with elution fractions 2,3,6,7 and 8 to determine the activity of the purified ECOL laccase. The fractions were rebuffered into deionized H2O using [http://www.sigmaaldrich.com/content/dam/sigma-aldrich/docs/Supelco/Product_Information_Sheet/4774.pdf HiTrap Desalting Columns] and incubated with 0.4 mM CuCl2. The reaction setup included 140 µL of a elution fraction, 100 mM sodium acetate buffer (pH 5), and 198 deionized H2O and 0.1 mM ABTS and the absorption was measured at 420 nm to detect oxidization over a time period of 12 hours at 25°C. Each fraction contained active laccase able to oxidize ABTS (see Figure 9). After 1 hour saturation was observed with ~52 µM oxidized ABTS. After 12 hours ~10 µM ABTS got reduced again, if referred to fraction 6. This behavior has been observed in the activity plot of[http://2012.igem.org/Team:Bielefeld-Germany/Results/Summary#7 TVEL0] before, indicating, that the oxidation catalyzed by this laccase is reversible. Additionally protein concentrations of each fraction were identified using the Bradford protocol. The tested fractions showed different amounts of protein after rebuffering, ranging from 0.2 to 0.6 mg mL-1. Fraction 7, containing the most protein and also most of active laccase was chosen for subsequent activity tests of ECOL. The protein concentration was reduced to 0.03 mg mL-1 for each measured sample to allow a comparison between TVEL0 measurements and ECOL measurements.
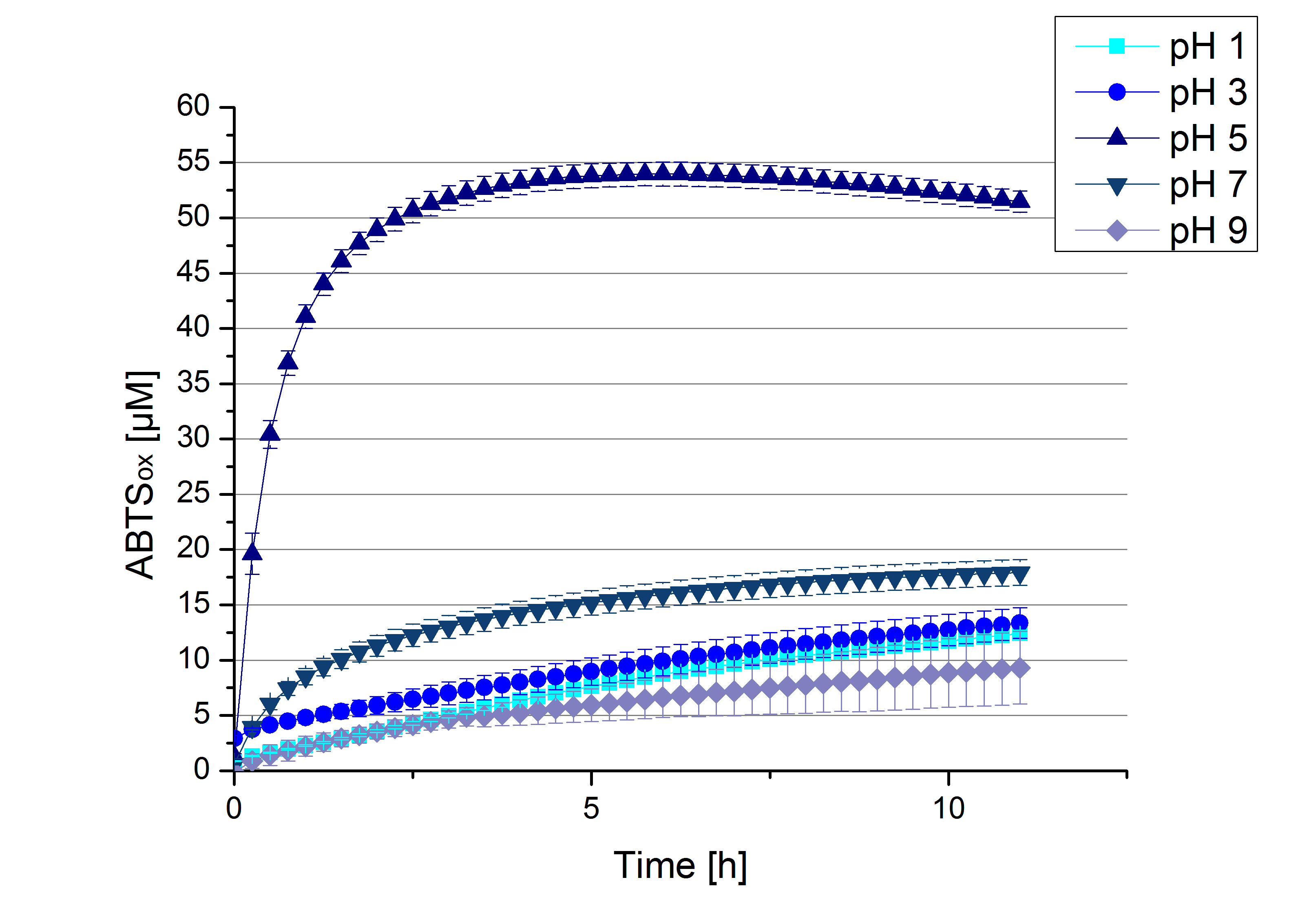
ECOL pH optimum
To determine at which pH the ECOL laccase has its optimum in activity, a gradient of sodium acetate buffer pHs was prepared. Starting with pH 1 to pH 9 ECOL activity was tested using the described conditions above and 0.03 mg mL-1 protein. The results are shown in Figure 10. A distinct pH optimum can be seen at pH 5. Saturation is reached after 2.5 hours with 53% oxidization of ABTS by the ECOL laccase at pH 5 (53 µM oxidized ABTS). The other tested pHs only led to a oxidation of up to 17% of added ABTS, respectively. Figure 11 shows the results of the analog experiments with laccase that was not incubated with CuCl2 before the activity measurements. Again, a pH optimum at pH 5 can be determined with 24 µM ABTS (24%) oxidized by ECOL after 8 hours under these conditions.
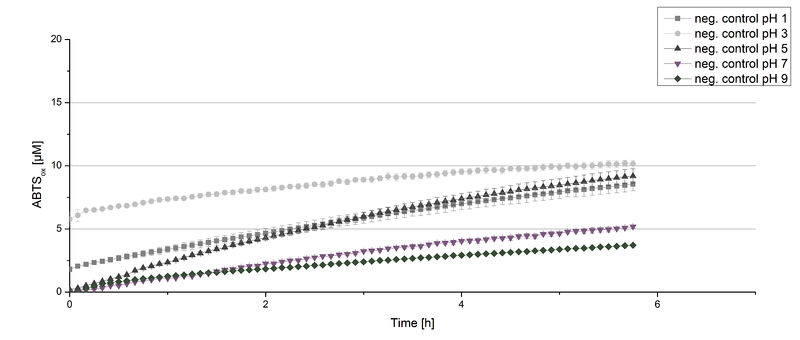
Figure 12 represents the negative control showing the oxidization of ABTS by 0.4 mM CuCl2 at the chosen pHs. The greatest increase in oxidized ABTS can be seen at a pH of 5: after 5 hours 15% ABTS is oxidized by CuCl2 alone. Nevertheless this result does not have an impact on the activity of the ECOL laccase at pH 5, which is still the optimal pH. Therefore it has the same pH optimum as [http://2012.igem.org/Team:Bielefeld-Germany/Results/Summary#7 TVEL0].
With regard to our project knowledge of the optimal pH is useful. Since waste water in waste water treatment plants has an average
pH of 6.9 it has to be kept in mind, that a adjustment of the pH is necessary for optimal laccase activity.
ECOL CuCl2 concentration
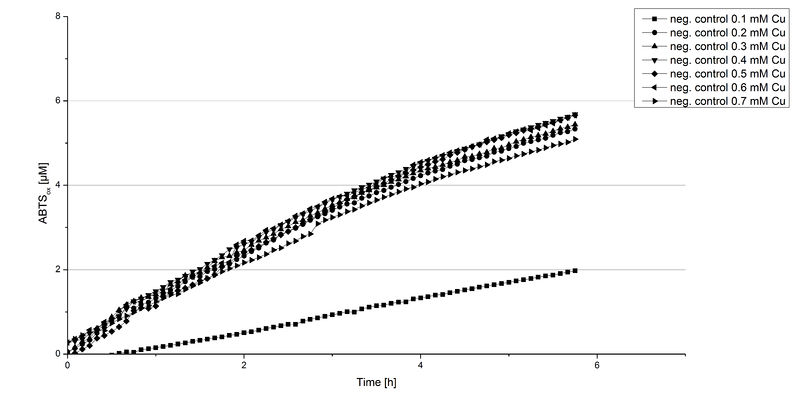
Another test of ECOL was done to survey the best CuCl2 concentration for the activity of the purified ECOL laccase. 0.03 mg mL-1 protein were incubated with different CuCl2 concentration ranging from 0 to 0.7 mM CuCl2. Activity tests were performed with the incubated samples, in 100 mM sodium actetate buffer (pH 5), 0.1 mM ABTS, to a final volume of 200 µL. The activity was measured at 420 nm, 25°C and over a time period of 10 hours. As expected the saturation takes place after 5 hours (see Figure 13). The differences in the activity of ECOL laccase incubated in different CuCl2 differ minimal. The highest activity of ECOL laccase is observed after incubation with 0.4 mM CuCl2 (42% of added ABTS). With a higher concentration of 0.7 mM CuCl2 the activity seems to be reduced (only 41% ABTS got oxidized). This leads to the assumption that CuCl2 supports the ECOL laccase activity but concentrations exceeding this value of CuCl2 may have a negative impact on the ability of oxidizing ABTS. Without any CuCl2 application ECOL laccase show less activity in oxidizing ABTS (see Figure 12). This fits the expectations as laccases are copper reliant enzymes and gain their activity through the incorporation of copper. Additionally negative controls were done using the tested concentrations of CuCl2 but no laccase was added to detect the oxidization of ABTS through copper (see Figure 14). The more CuCl2 was present, the more ABTS was oxidized after 5 hours. Still the maximal change accounts only for ~6% oxidized ABTS after 5 hours.
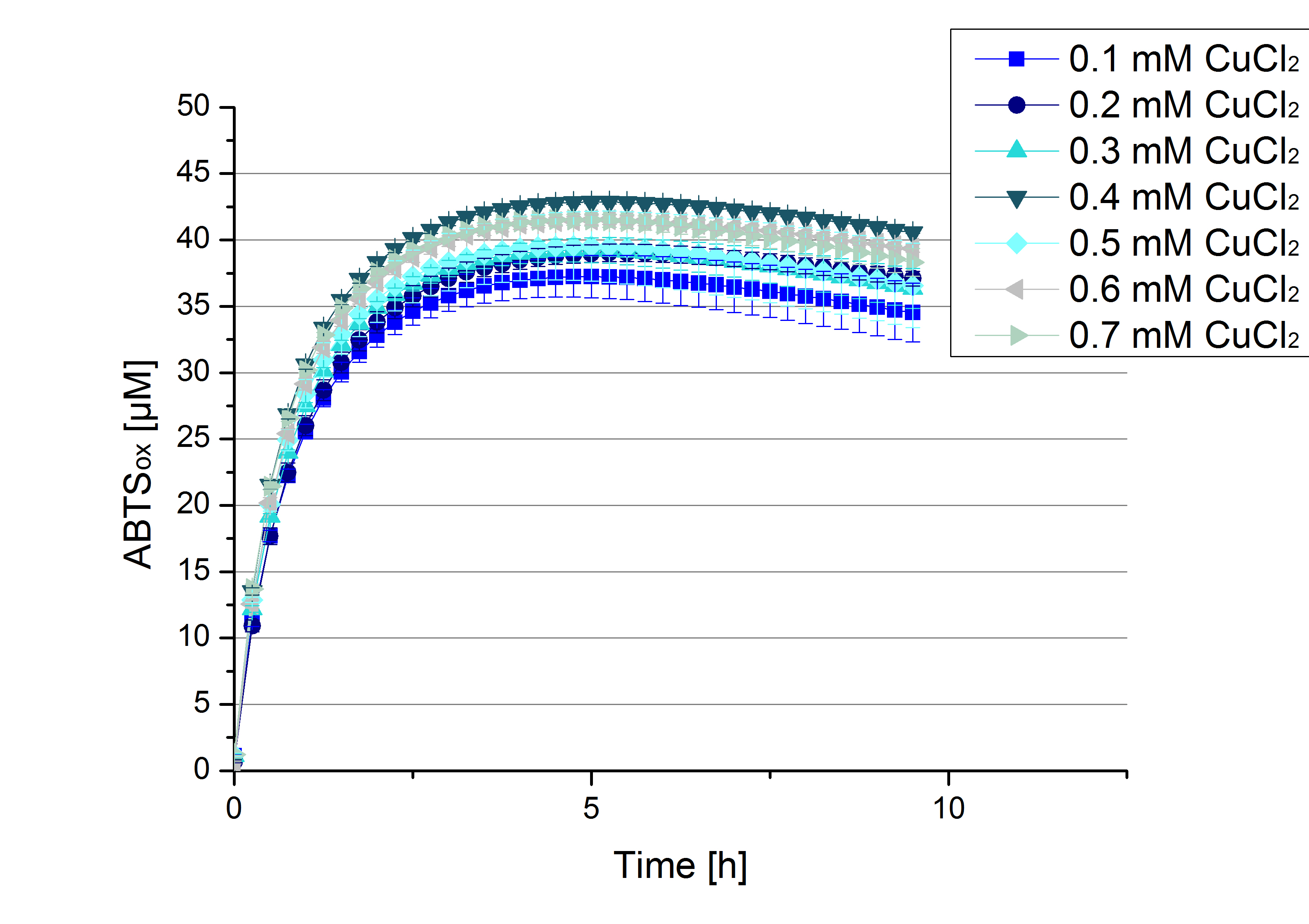
In relation to apply the laccase in waste water treatment plants it is beneficial knowing, that small amounts of CuCl2 are enough to activate the enzymes. This reduces the cost factor for the needed CuCl2 to incubate the laccases before application.
ECOL activity at different temperatures
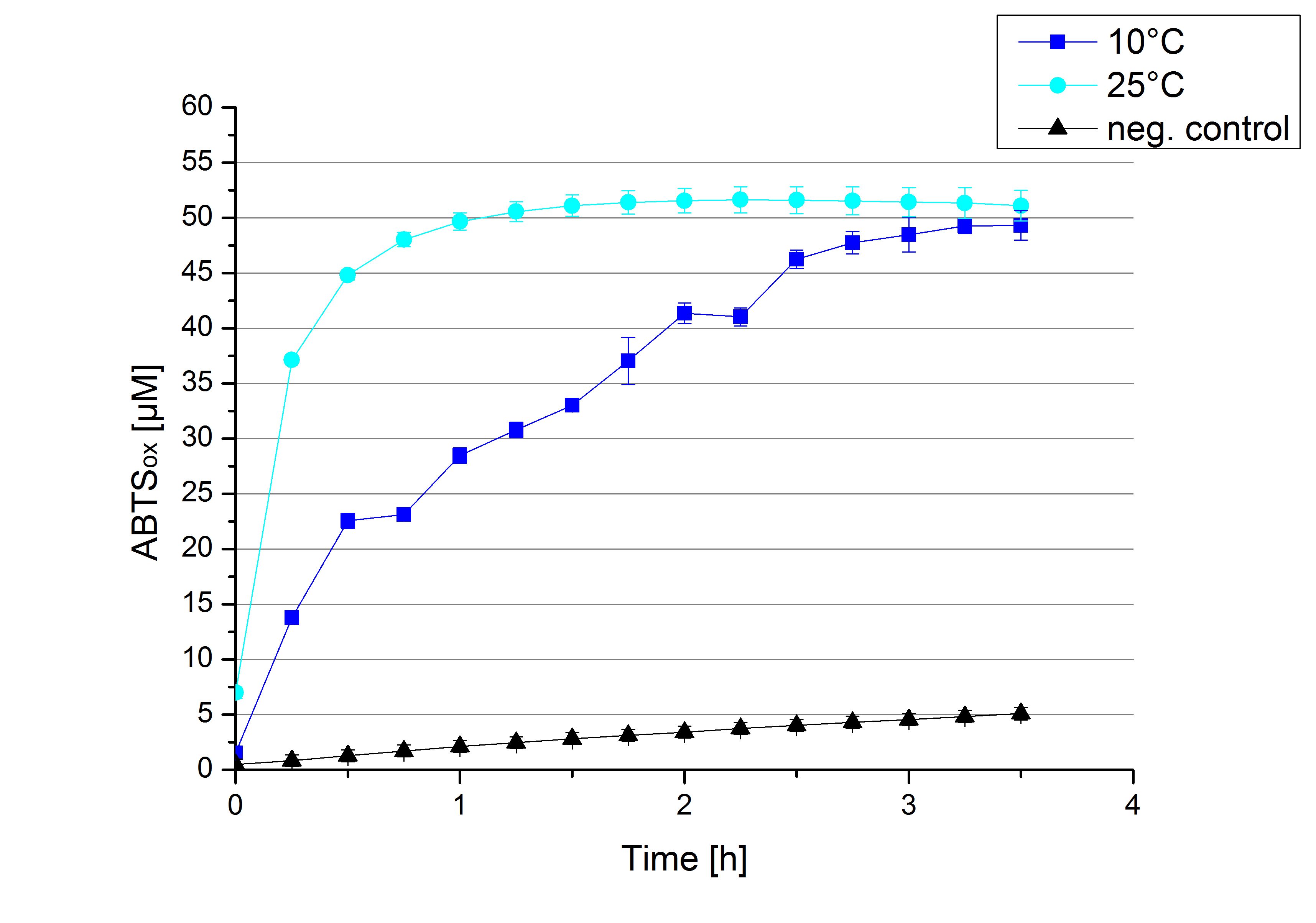
To investigate the activity of ECOL at lower temperatures activity tests as described above were done at 10°C and 25°C (Figure 15). A significant decrease in the activity can be observed upon reducing the temperature from 25°C to 10°C. While the activity at 10 °C is reduced, final saturation levels are still comparable: after 3,5 hours, only 2% difference in oxidized ABTS is observable. The negative control without the ECOL laccase and only 0.4 mM CuCl2 at 10°C shows a negligible oxidation of ABTS. Although a decrease in the activity of ECOL laccase was expected the observed reduction in enzyme activity is problematic for the possible application in waste water treatment plants where the temperature differs from 8.1°C to 20.8°C. A more cryo tolerant enzyme would be preferable.
MALDI-TOF Analysis of ECOL
<p align="justify"> The E. coli laccase was identified using the following software
- FlexControl
- Flexanalysis and
- Biotools
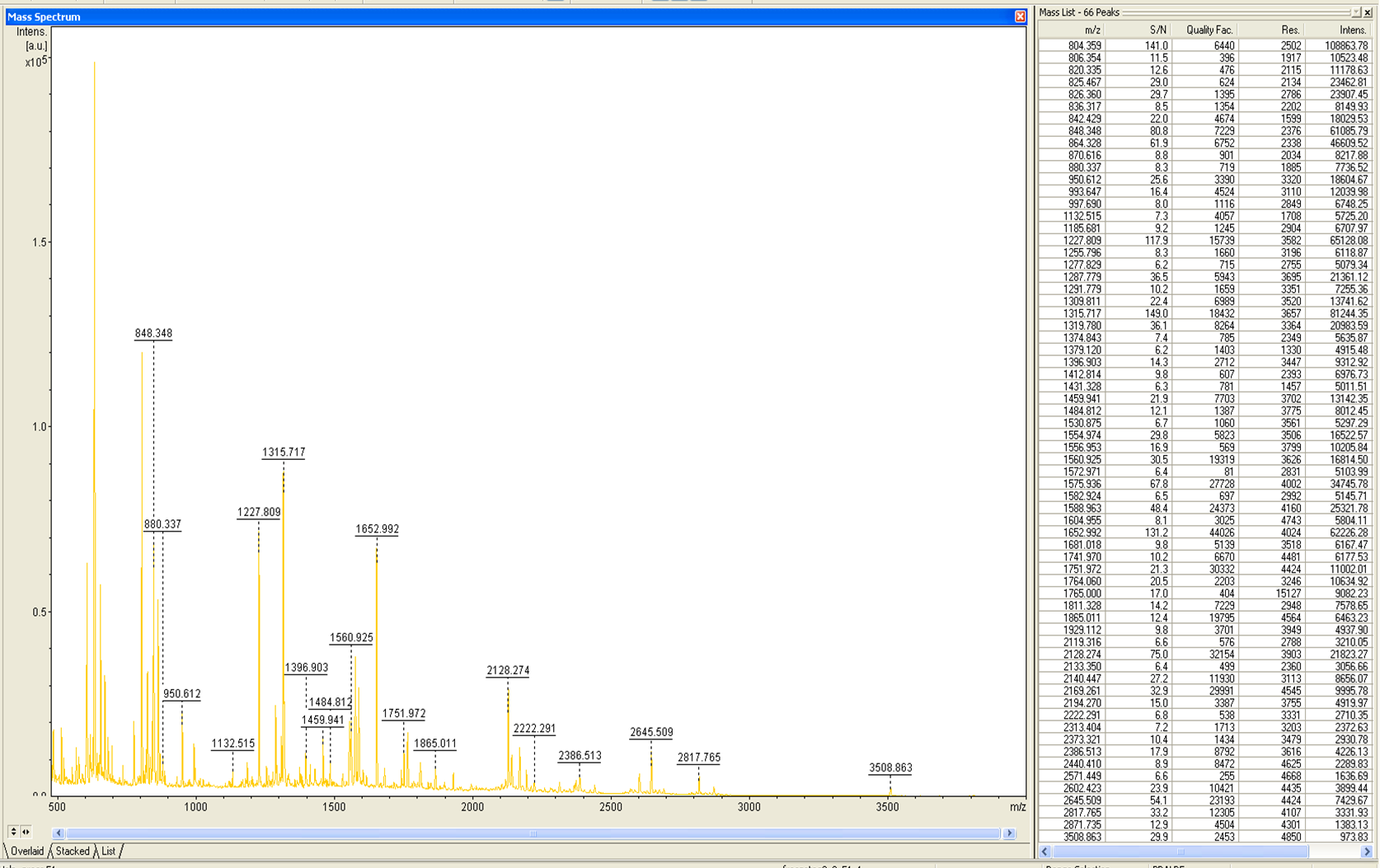
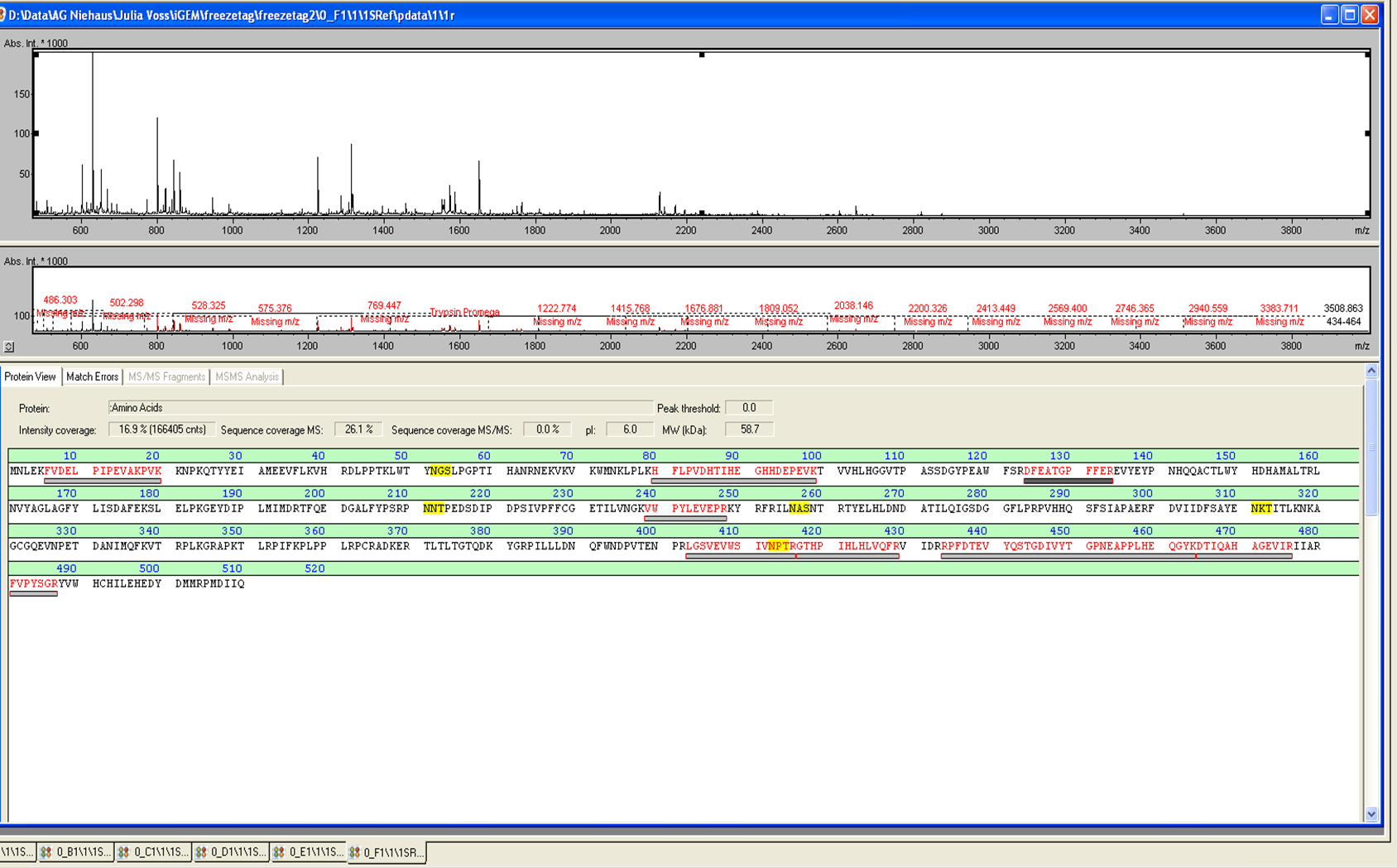
from Brunker Daltronics. TheE. coli laccase P36649 was identified with a mascot-score of 108 with an automatic run. In Figure 7 and 8 the chromatogram of the peptid mass fingerprint and the single masses are shown with a sequence coverage of 26,1%. It can be assumed that the isolated protein is ECOL.
Substrate Analytic
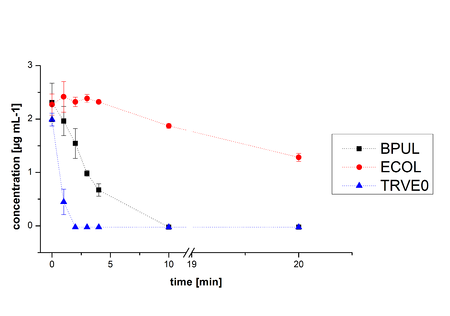
In Figure 19 the degradation reactions of ethinyl estradiole with ABTS of BPUL and ECOL are shown. The data was measured by HPLC. Since it was already determined that both are able to oxidize ABTS by the activity test, the results were as expected. ABTS works as mediator, which explains the ability to degrade ethinyl estradiol and probably other substrates. In comparison, ECOL has a lower potential for degradation of ethinyl with ABTS than TVEL0 and BPUL. Until now, no further experiments were made.
Immobilisation

Figure 20 shows the percentage of laccases in the supernatant after incubation with CPC-beads, relative to the original concentration . The concentration of laccases in the supernatant after incubation was measured using Roti®-Nanoquant. The results show that only 0.2% of ECOL laccases are still present in the supernatant. This illustrates that ECOL was successfully immobilized on the CPC-beads.
In figure 21, the enzymatic activity of ECOL in the supernatant is compared to the activity of nontreated ECOL. Although an activity can already be detected in the supernatant, this activity is low compared to the original.
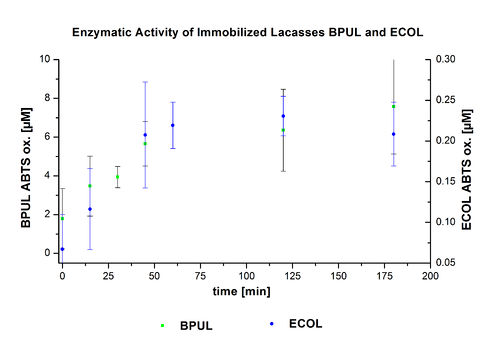
Figure 22 presents the enzymatic activity of immobilized laccases compared to nontreated laccases. The activity of bound ECOL is very low, even lower than that in the supernatant.
//function/degradation
//proteindomain/degradation
| None |