Difference between revisions of "Part:BBa K592010"
Georgedashen (Talk | contribs) (→Usage and Biology) |
|||
| Line 22: | Line 22: | ||
'''iGEM12_Uppsala_University:''' The Uppsala chromoprotein collection and RFP. The image shows pellets of ''E coli'' expressing chromoproteins eforRed <partinfo>BBa_K592012</partinfo>, RFP <partinfo>BBa_E1010</partinfo>, cjBlue <partinfo>BBa_K592011</partinfo>, aeBlue <partinfo>BBa_K864401</partinfo>, amilGFP <partinfo>BBa_K592010</partinfo> and amilCP <partinfo>BBa_K592009</partinfo>. | '''iGEM12_Uppsala_University:''' The Uppsala chromoprotein collection and RFP. The image shows pellets of ''E coli'' expressing chromoproteins eforRed <partinfo>BBa_K592012</partinfo>, RFP <partinfo>BBa_E1010</partinfo>, cjBlue <partinfo>BBa_K592011</partinfo>, aeBlue <partinfo>BBa_K864401</partinfo>, amilGFP <partinfo>BBa_K592010</partinfo> and amilCP <partinfo>BBa_K592009</partinfo>. | ||
| + | |||
| + | '''iGEM17_SJTU_BioX_Shanghai:'''<br> | ||
| + | '''J23119+Target3+amilGFP '''<partinfo>BBa_K2285013</partinfo><br> | ||
| + | '''J23119+Target1+amilGFP ''' <partinfo>BBa_K2285017</partinfo><br> | ||
| + | |||
| + | An improved part has been constructed. Since this part is a coding sequence, we added a RBS which is on the upstream of sfGFP <partinfo>BBa_K515005</partinfo> and terminator <partinfo>BBa_B1006</partinfo> by PCR. What's more, our device have a stem-loop structure(called Target) following the constitutive promoter J23119, to achieve further control of amilGFP expression. | ||
===References=== | ===References=== | ||
Revision as of 01:34, 28 October 2017
amilGFP, yellow chromoprotein
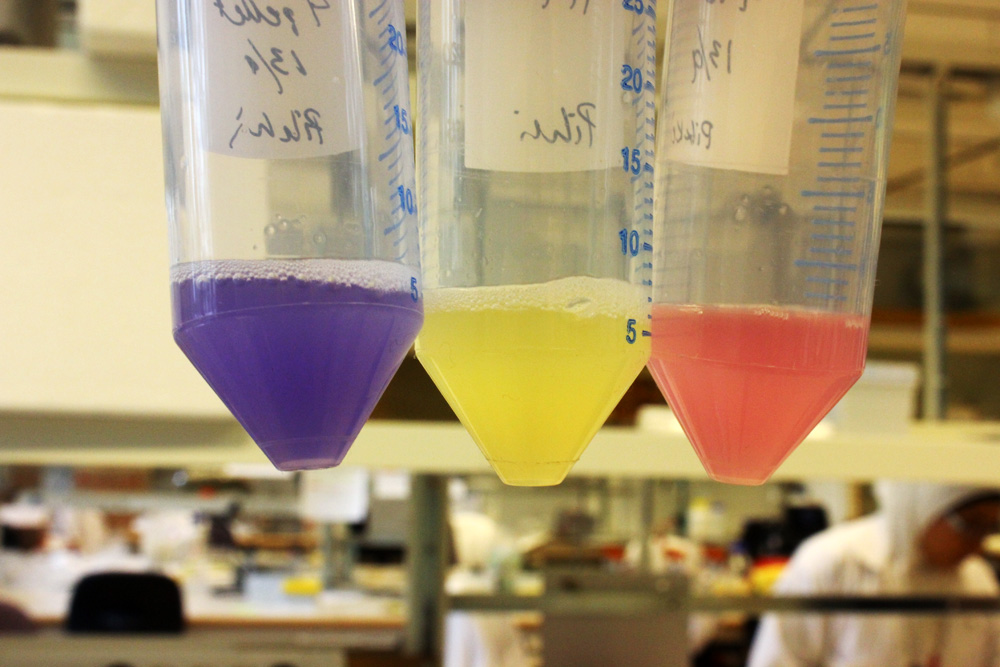
This chromoprotein from the coral Acropora millepora, amilGFP, naturally exhibits strong yellow color when expressed. The color is readily visible to naked eye both in LB-culture and on agar plates. Color development can be seen in less than 24 hours of incubation.
Important: This part is not available in the registry yet, however, the same part is available from the registry with the standard RBS B0034: BBa_K1033931.
Usage and Biology
This part is useful as a reporter.
Peking iGEM 2016 has fused this part with triple spytag. The fused protein is participate in Peking’s polymer network. By adding this protein, the whole polymer network become visible in most conditions. If you want to learn more about Peking’s polymer network and the role of mRFP in this network, please click here https://parts.igem.org/Part:BBa_K1989003", https://parts.igem.org/Part:BBa_K1989045" or https://parts.igem.org/Part:BBa_K1989048".
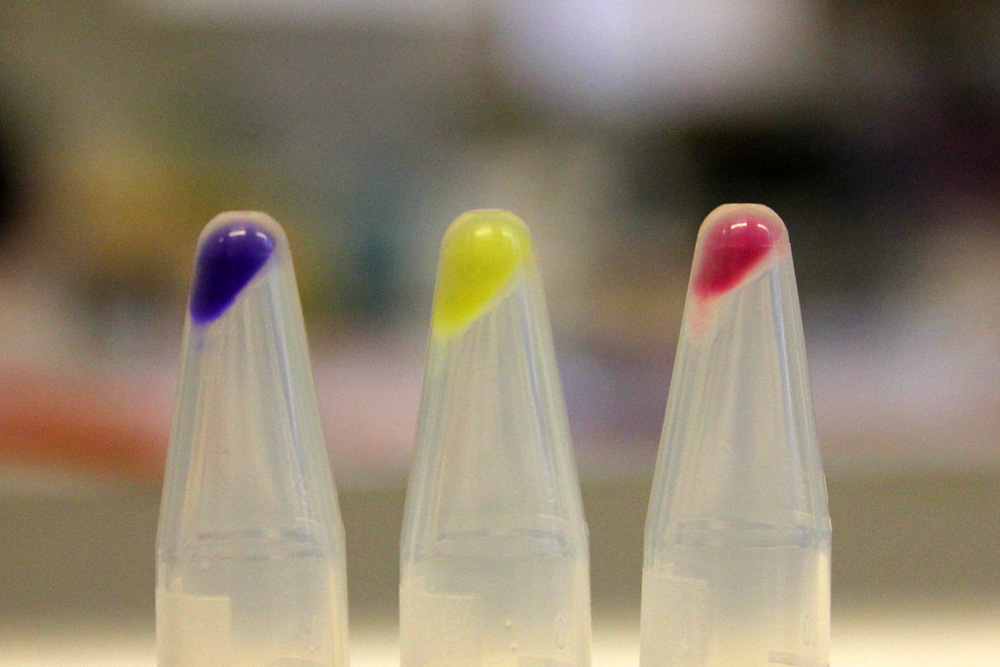
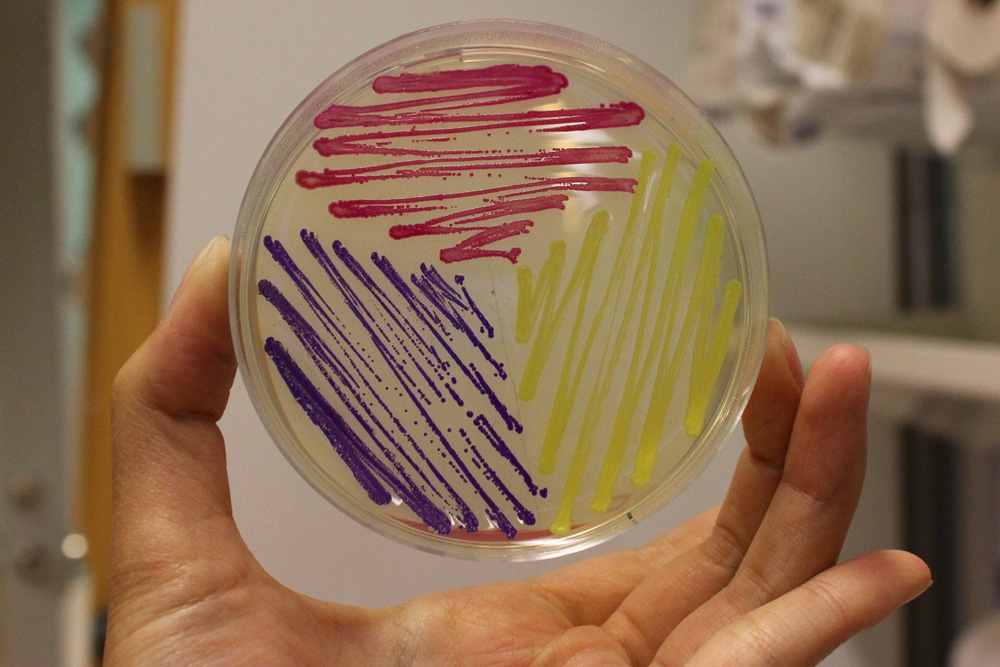
iGEM11_Uppsala-Sweden: Expression of chromoproteins. The images above show E coli constitutively expressing amilGFP BBa_K592010 (yellow), amilCP BBa_K592009 (blue), and RFP BBa_E1010 (red).
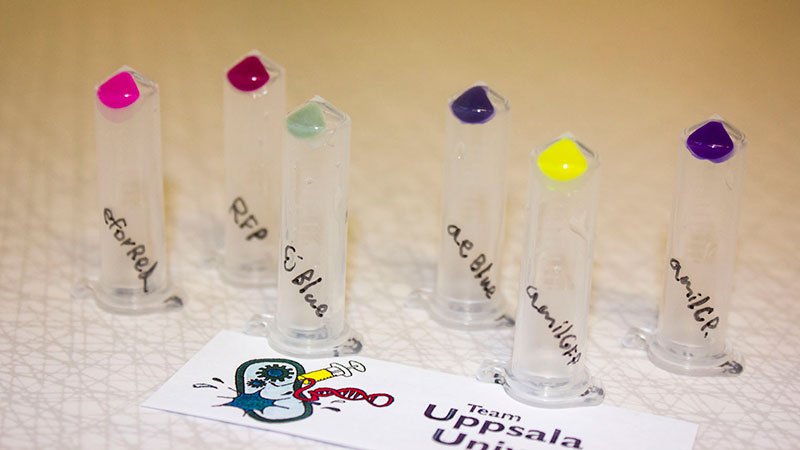
iGEM12_Uppsala_University: The Uppsala chromoprotein collection and RFP. The image shows pellets of E coli expressing chromoproteins eforRed BBa_K592012, RFP BBa_E1010, cjBlue BBa_K592011, aeBlue BBa_K864401, amilGFP BBa_K592010 and amilCP BBa_K592009.
iGEM17_SJTU_BioX_Shanghai:
J23119+Target3+amilGFP BBa_K2285013
J23119+Target1+amilGFP BBa_K2285017
An improved part has been constructed. Since this part is a coding sequence, we added a RBS which is on the upstream of sfGFP BBa_K515005 and terminator BBa_B1006 by PCR. What's more, our device have a stem-loop structure(called Target) following the constitutive promoter J23119, to achieve further control of amilGFP expression.
References
[http://www.ncbi.nlm.nih.gov/pubmed/18648549] Alieva, N. O., et al. 2008. Diversity and evolution of coral fluorescent proteins. PLoS One 3:e2680.
GenBank: AY646067.1
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]