Difference between revisions of "Part:BBa C0062:Experience"
(→Results) |
(→Results) |
||
| Line 66: | Line 66: | ||
|[[File:ETH_Zurich_2014_qs-table_pLas.png|100px|link=https://parts.igem.org/Part:BBa_R0079:Experience]] | |[[File:ETH_Zurich_2014_qs-table_pLas.png|100px|link=https://parts.igem.org/Part:BBa_R0079:Experience]] | ||
|[[File:ETH_Zurich_2014_qs-table_PlasLuxRLuxAHL.png|200px]] | |[[File:ETH_Zurich_2014_qs-table_PlasLuxRLuxAHL.png|200px]] | ||
| − | |[[File:ETH_Zurich_2014_qs- | + | |[[File:ETH_Zurich_2014_qs-table_PlasLuxRLasAHL.png|200px]] |
|[[File:ETH_Zurich_2014_qs-table_PlasLuxRRhlAHL.png|200px]] | |[[File:ETH_Zurich_2014_qs-table_PlasLuxRRhlAHL.png|200px]] | ||
|- | |- | ||
Revision as of 18:08, 20 October 2014
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_C0062
User Reviews
UNIQ7e7af3f8ea886e91-partinfo-00000000-QINU
|
••••
ETH Zurich 2014 |
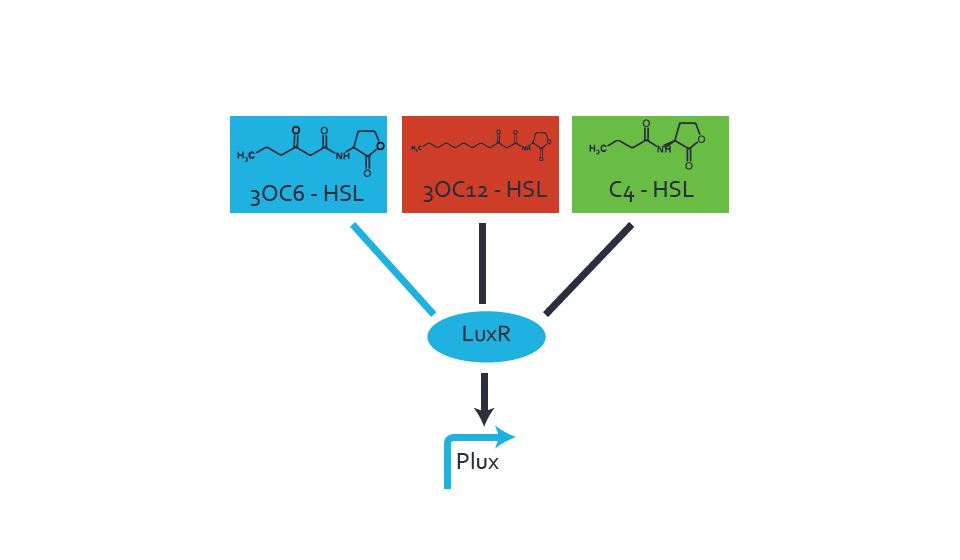
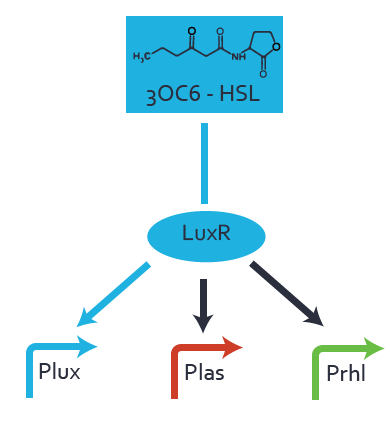
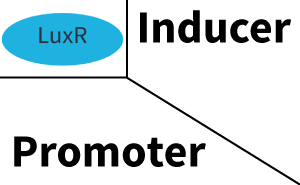
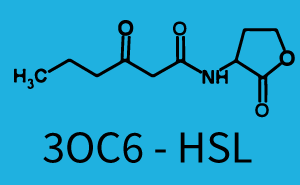
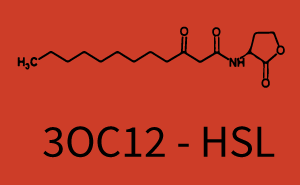
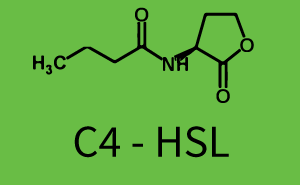
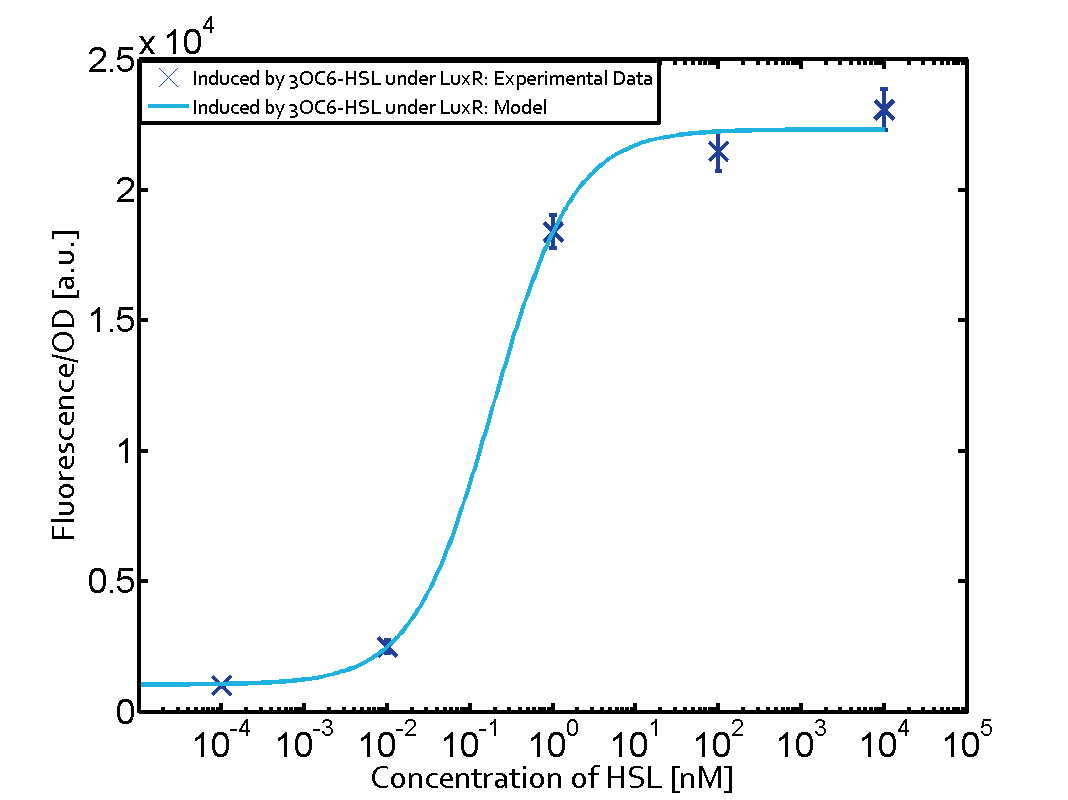
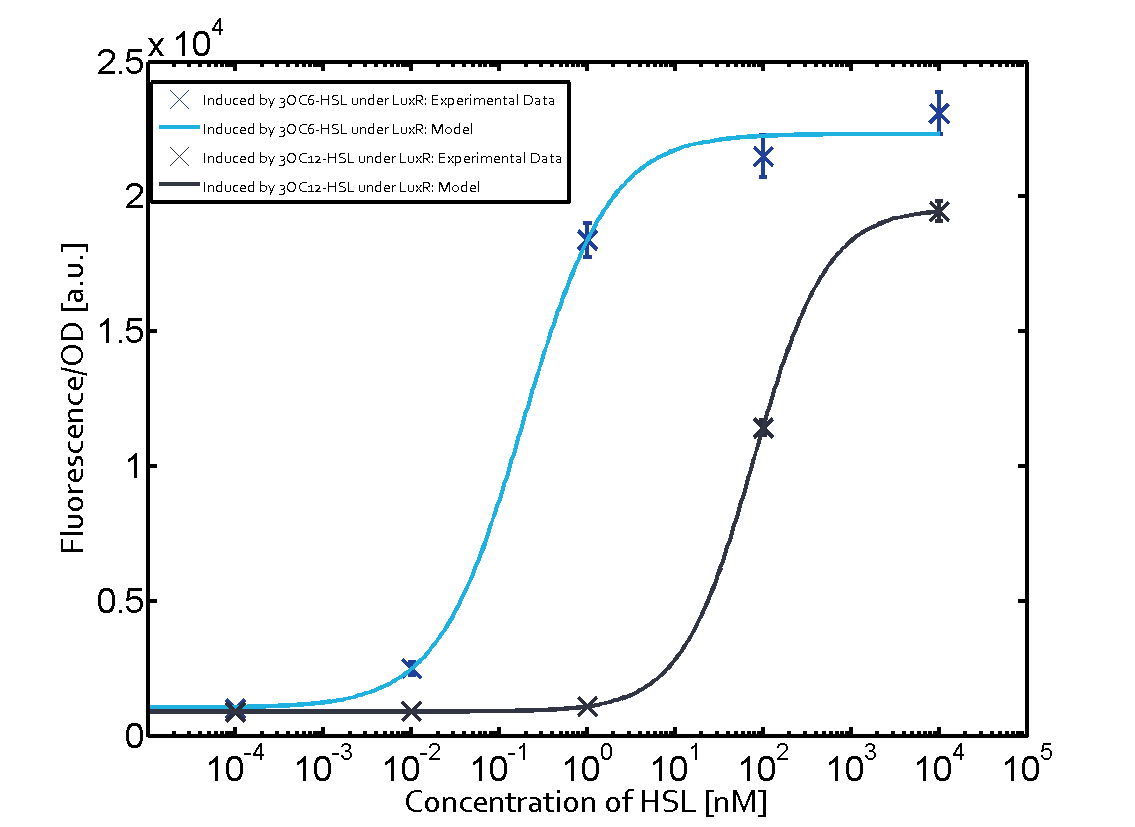
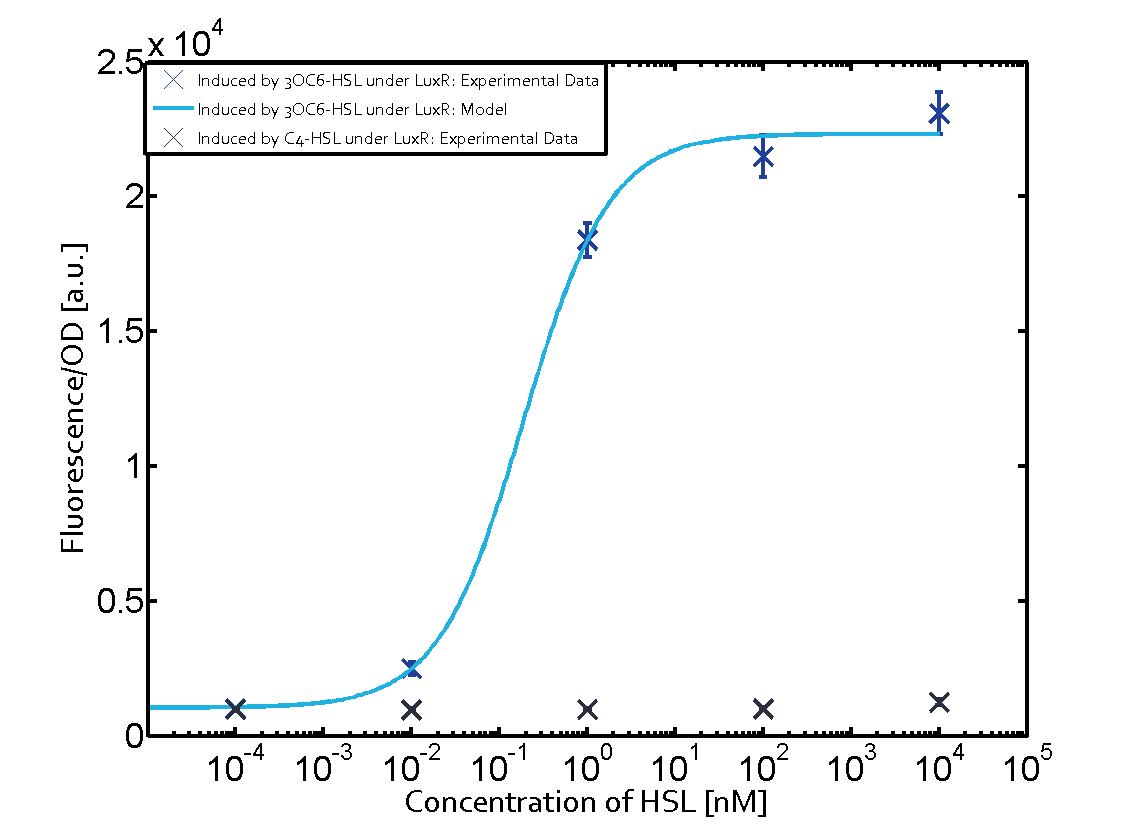
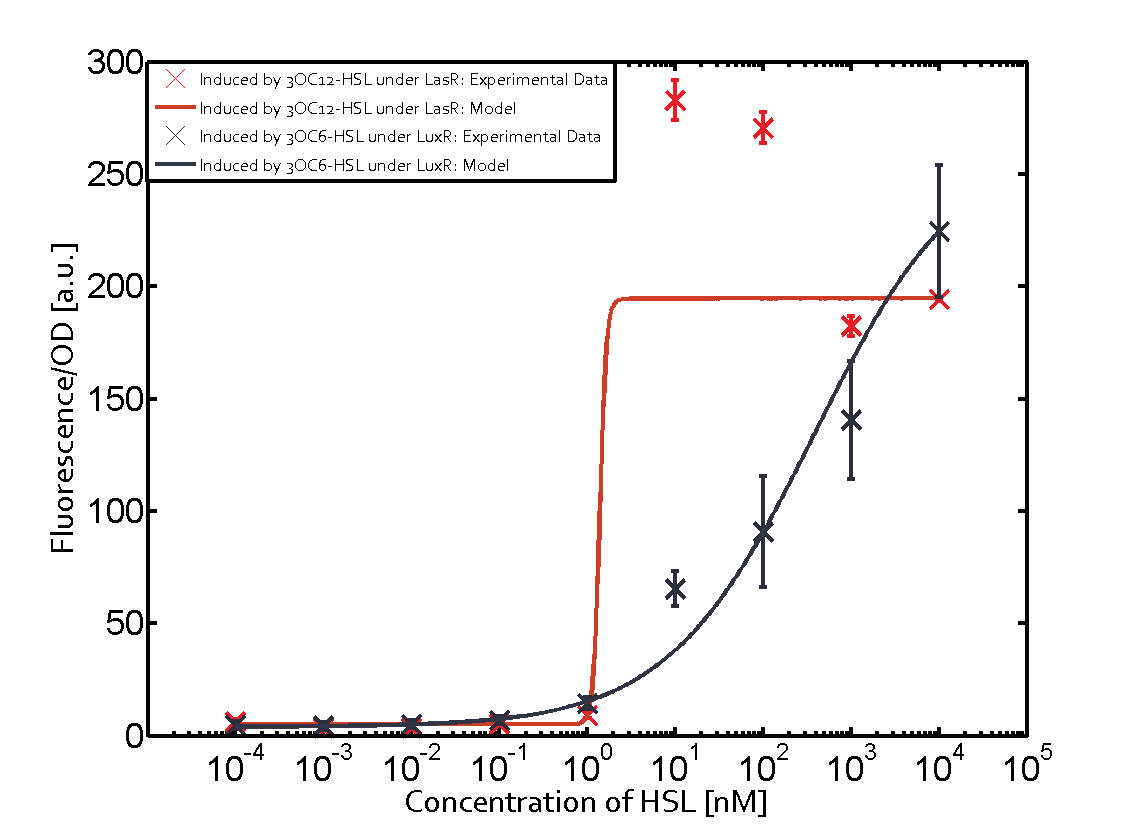
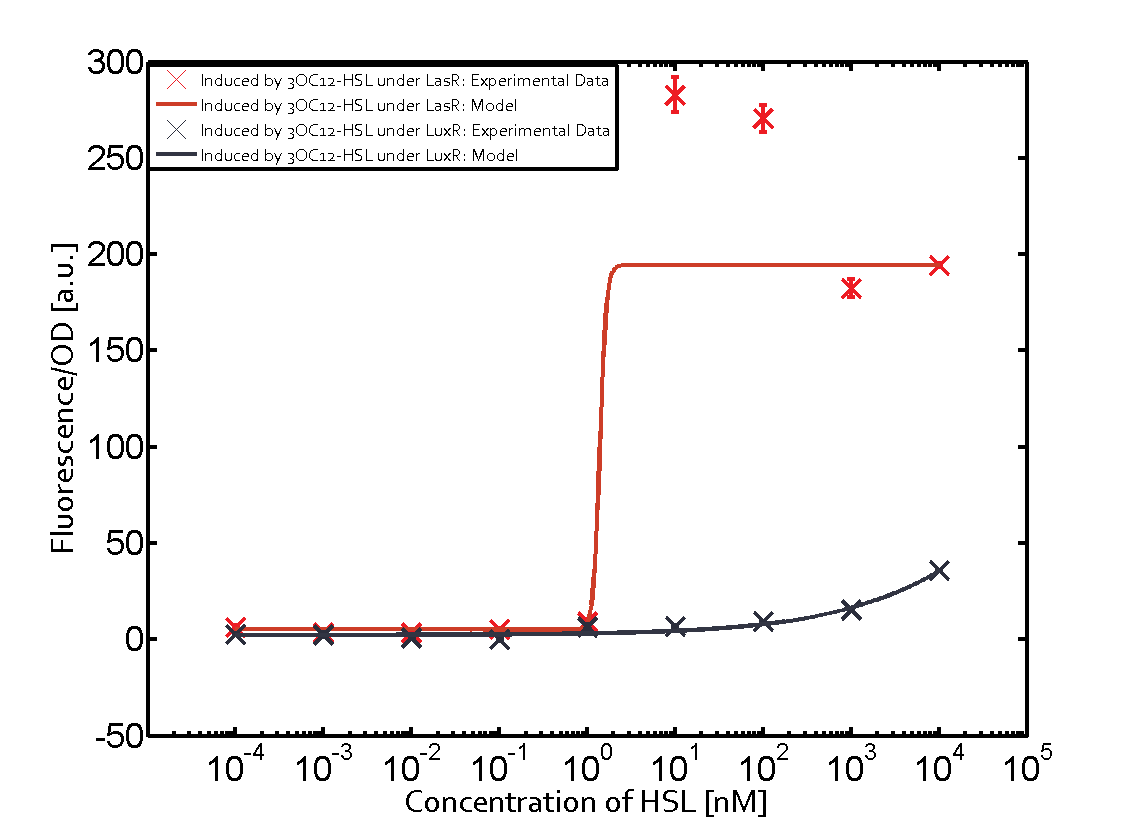
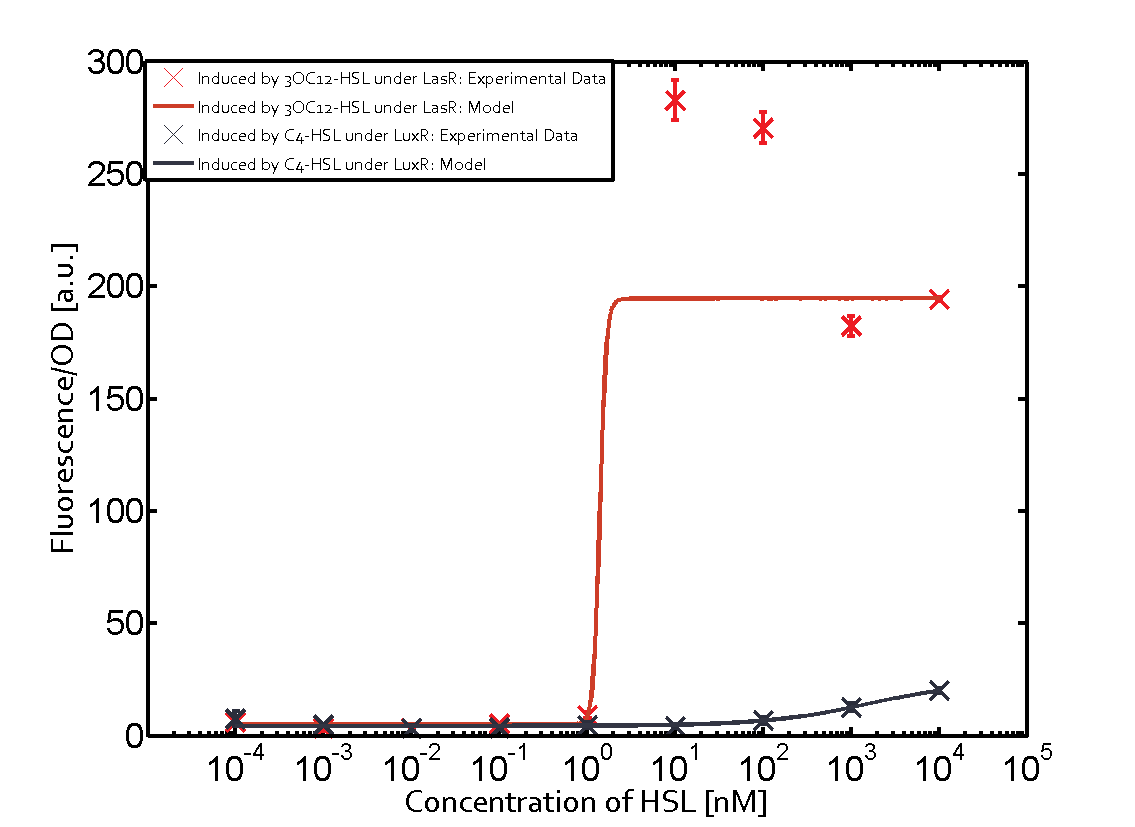
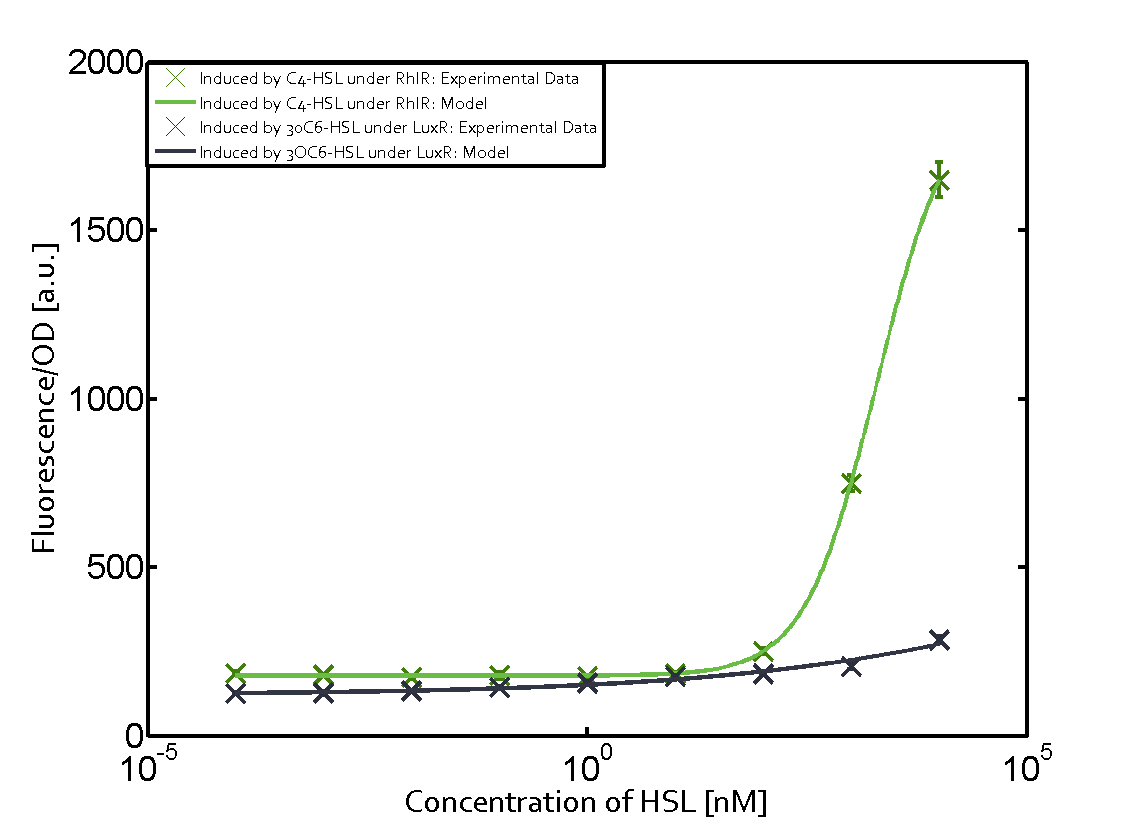
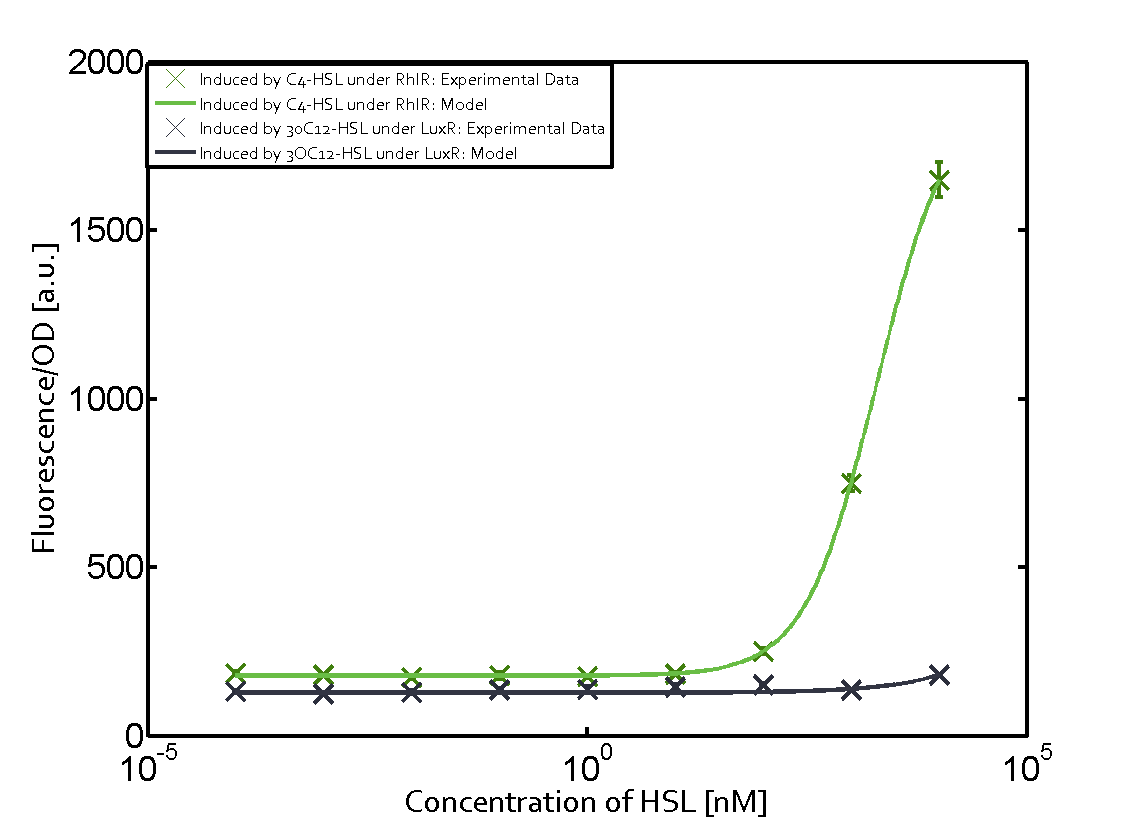
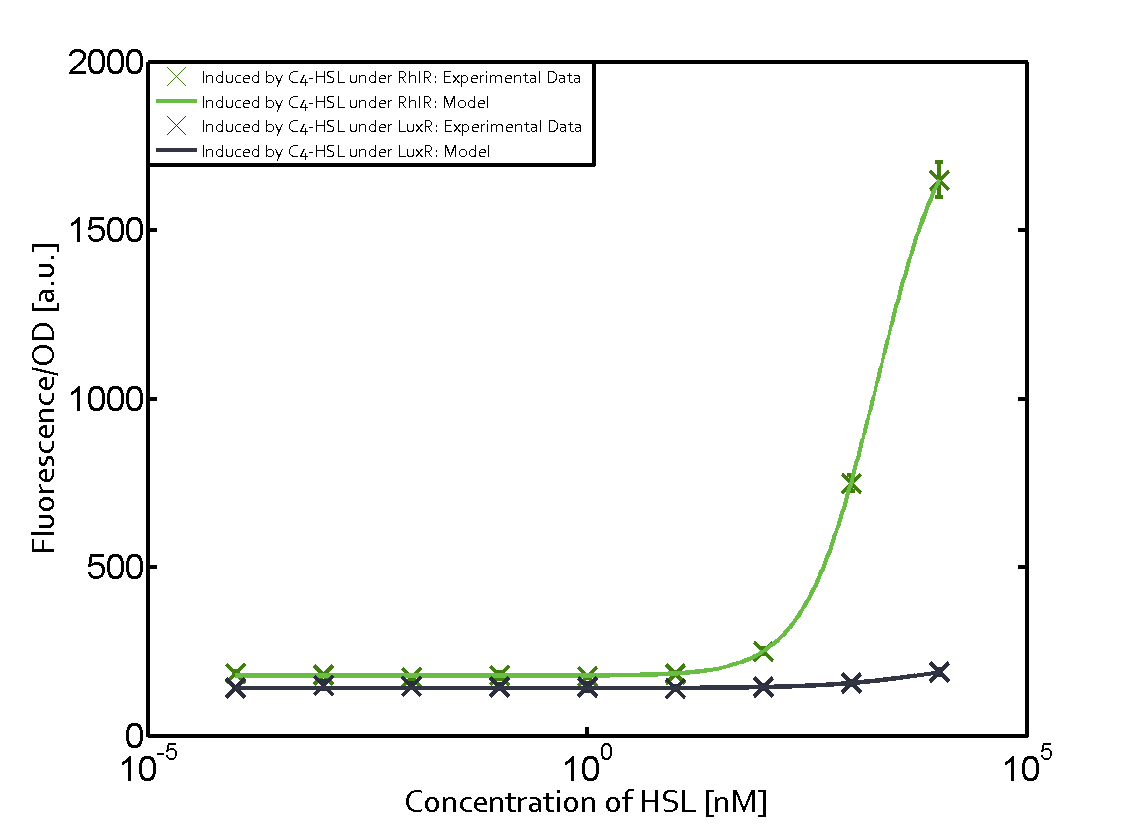
Characterization of two-order crosstalkBackground informationSystem consideredModeling crosstalkFirst-order crosstalkFirst Level crosstalk: LuxR binds to different HSL and activates the promoter PluxSecond Level crosstalk: LuxR binds to 3OC6-HSL, its natural HSL, and activates different promoterSecond order crosstalk: Combination of both cross-talk levelsOther regulatory proteins, like LuxR, bind to different HSL and activates the promoter. Results
| ||||||||||||||||||||
|
•••••
SUN(Tsinghua) |
Part was sequenced and functional. LuxR was used in our Portable Pathogen Detector. |
|
•••••
wmholtz |
Using this part, I have successfully constructed and tested a quorum sensing circuit in E. coli. |
|
•••••
Youri |
This part was used and tested as a subpart in K546000, K546001, K546002, K546003, K546005 and K546546. This part functioned in all cases. |
|
Kevin (iGEM Braunschweig 2013) |
The plasmid pSB1C3 BBa_C0062 from the 2013 distribution Kit was transformed in E. coli XL1 BlueMRF. Sequencing with standard verification primer VF2 confirmed matching sequence of backbone DNA up to the EcoRI restriction site. The rest of the sequence (not shown) does not match the registry entry.
96 145
pSB1C3 LuxR (96) GAGGCAGAATTTCAGATAAAAAAAATCCTTAGCTTTCGCTAAGGATGATT
C0062 VF2 (1) GAGGCAGAATTTCAGATAAAAAAAATCCTTAGCTTTCGCTAAGGATGATT
146 195
pSB1C3 LuxR (146) TCTGGAATTCGCGGCCGCTTCTAGAGATGAAAAACATAAATGCCGACGAC
C0062 VF2 (51) TCTGGAATTCGACGCAA-TGGGTGCGCTGTCTACTAAATACAACGACACC
196 245
pSB1C3 LuxR (196) ACATACAGAATAATTAATAAAATTAAAGCTTGTAGAAGCAATAATGATAT
C0062 VF2 (100) CCGGAAAAAGCCTCCCGTACTTACGACGCTCACCGTGACGGTTTCGTTAT
246
pSB1C3 LuxR (246) TAATCAATGC...
C0062 VF2 (150) CGCTGGCGGC...
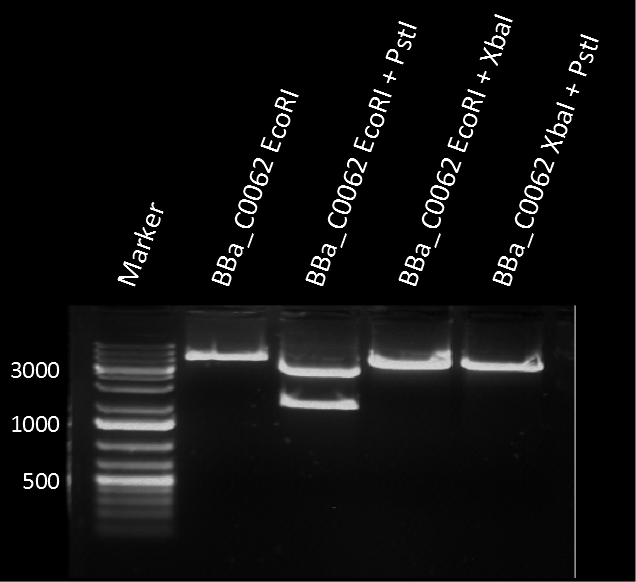
A restriction assay (Figure 1) showed that the sequenced part has no XbaI restriction site following the EcoRI site indicating another part in front of BBa_C0062 with a length of at least 1000 bp. |
UNIQ7e7af3f8ea886e91-partinfo-00000007-QINU