Difference between revisions of "Part:BBa K325909"
| (51 intermediate revisions by 8 users not shown) | |||
| Line 1: | Line 1: | ||
| + | <partinfo>BBa_K325909 short</partinfo> | ||
<div style="font-family: Arial; padding: 00px; width: 680px; border: 0px solid #000000;"> | <div style="font-family: Arial; padding: 00px; width: 680px; border: 0px solid #000000;"> | ||
{{Template:K325909 page header}} | {{Template:K325909 page header}} | ||
{{Template:K325909 Characterization Navbar}} | {{Template:K325909 Characterization Navbar}} | ||
<div style="padding: 00px; width: 680px; border: 0px solid #000000;"> | <div style="padding: 00px; width: 680px; border: 0px solid #000000;"> | ||
| + | |||
| + | |||
'''Description'''<br> | '''Description'''<br> | ||
| − | G28 | + | This page described the ''lux operon'' from ''Vibrio fischeri''. To relieve LuxR control we placed Lux C, D, A, B, E under the [https://parts.igem.org/Part:BBa_I0500 pBad] promoter. A more complete description can be found on the [http://2010.igem.org/Team:Cambridge/Bioluminescence/G28 Cambridge iGEM 2010 website]. |
| − | + | This Bacterial lux operons encodes five enzymes involved in the light-generating pathway. LuxA and LuxB encode the two subunits of the bacterial luciferase, while the products of LuxC, LuxD and LuxE synthesise the substrate for the light emitting reaction, tetradecanal. The exact function of LuxG is unknown, and it appears to be non-essential for light emission, but its presence increases light output. | |
| + | In nature, the lux genes appear to be repressed by the nucleoid protein, H-NS, and occur under quorum sensing control. We removed the natural quorum sensing control to facilitate use of the part in biosensors under different regulatory inputs. | ||
| − | + | As of October 2010 we believe this is the first and only BioBrick to emit light in normal E. coli strains without the addition of any external substrate. | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | Measured by the [http://2010.igem.org/Team:Cambridge Cambridge iGEM team 2010] | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
<div style="padding: 00px; width: 680px"> | <div style="padding: 00px; width: 680px"> | ||
'''Compatibility'''<br> | '''Compatibility'''<br> | ||
| − | [https://parts.igem.org/cgi/partsdb/pgroup.cgi?pgroup=cell ''Chassis:''] Device has been shown to work in ''Top 10 (Invitrogen)'' | + | [https://parts.igem.org/cgi/partsdb/pgroup.cgi?pgroup=cell ''Chassis:''] Device has been shown to work in ''Top 10 (Invitrogen)'', [http://www.ecoliwiki.net/colipedia/index.php/BW25113 BW25113 DELTA H-NS::kan] and H-NS mutant JM 230 H-NS -205::tn10.br> |
[[Plasmid backbones|''Plasmids:'']] Device has been shown to work on ''<partinfo>pSB1C3</partinfo>'' <br> | [[Plasmid backbones|''Plasmids:'']] Device has been shown to work on ''<partinfo>pSB1C3</partinfo>'' <br> | ||
| + | |||
| + | <b>Additional characterisation by the [http://2015.igem.org/Team:Penn Penn iGEM team 2015]<br></b> | ||
| + | Penn iGEM 2015 used this part to make the sender cells in light-mediated cell communication. The lux operon BioBrick on PSB1C3 was transformed into NEB 10-beta competent cells (since it is a commonly used E. coli strain in iGEM) and an H-NS mutant E. coli strain (since the protein has been shown to reduce lux gene expression). The luminescent signal of the two sender strains was compared over time to determine what luminescent output trend would be best for inducing our light-sensitive receiver. Furthermore, using a calibrated conversion system designed by the team, Penn IGEM was about to convert the output from relative lights units (a measure typically used to calculate bioluminescent signals) to an absolute measure in uW/cm^2. Using the absolute units will assist in making the characterization of bioluminescent parts such as BBa_K325909 more consistent across luminometer devices and across projects. For more information, please follow this [http://2015.igem.org/Team:Penn/Sender link]. | ||
| + | |||
| + | <br /> | ||
| + | <b>Additional characterisation by the [http://2017.igem.org/Team:NTNU_Trondheim NTNU Trondheim iGEM team 2017]</b> | ||
| + | <br> | ||
| + | Team NTNU Trondheim 2017 has improved the characterisation of the biobrick BBa_K325909, which is an operon consisting of Lux C, D, A, B, E under the arabinose-induced promoter pBAD. | ||
| + | <br><br> | ||
| + | By looking at previous characterisation results of the biobrick, we thought it would work reasonably to use DH5α competent E. coli cells and an incubation temperature of 37 °C. However, when testing with 0.01M arabinose concentrations, no observable luminescence was detected at any time after induction. Therefore, we decided to characterise the biobrick by measuring the luminescence from the lux operon at different arabinose concentrations and temperatures. We measured the luminescence with a Tecan M200 plate reader, using four different temperatures and six different arabinose concentrations. As the machine could only perform heating and not cooling, it was challenging to measure at temperatures below 29 °C. | ||
| + | <br><br> | ||
| + | Results from our experiments are shown in the graphs below. As no luminescence was detected at 37 °C, these graphs are not included. | ||
| + | <br><br> | ||
| + | [[File:Lux_graph_24.png|500px|thumb|center|Luciferase luminescence of the lux operon in supercompetent DH5 E. coli cells, detected by a Tecan M200 plate reader at 24.0 °C and six arabinose concentrations over five hours.]] | ||
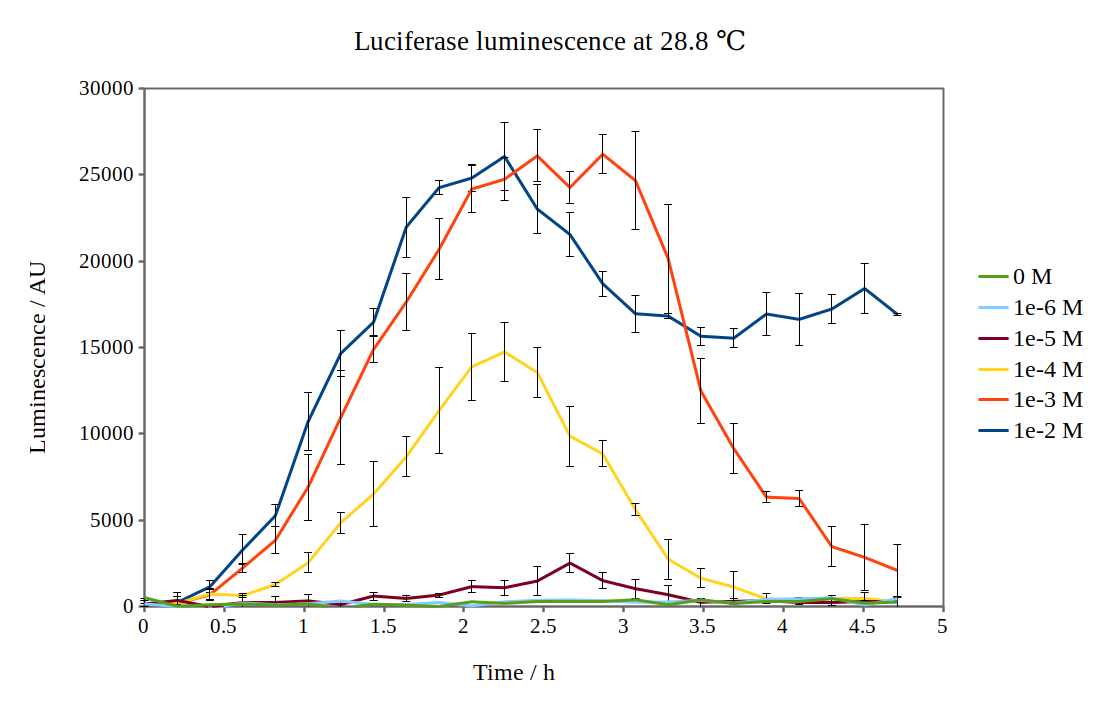
| + | [[File:Lux_graph_28-8.png|500px|thumb|center|Luciferase luminescence of the lux operon in supercompetent DH5 E. coli cells, detected by a Tecan M200 plate reader at 28.8 °C and six arabinose concentrations over five hours.]] | ||
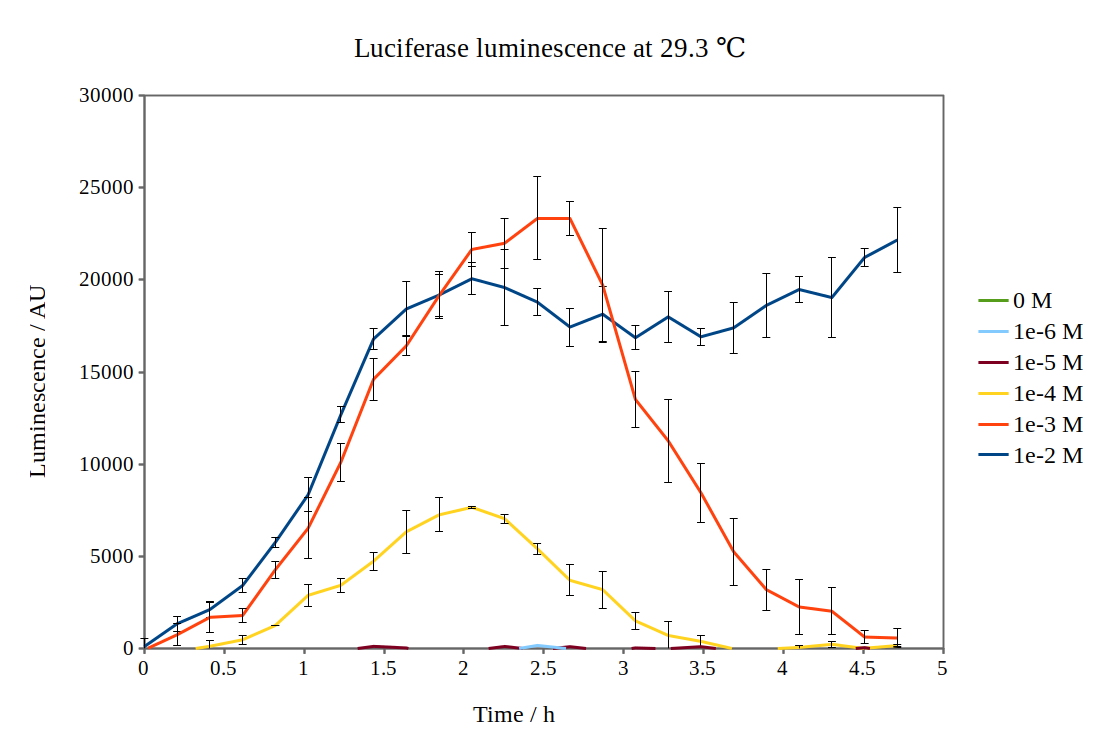
| + | [[File:Lux_graph_29 3.png|500px|thumb|center|Luciferase luminescence of the lux operon in supercompetent DH5 E. coli cells, detected by a Tecan M200 plate reader at 29.3 °C and six arabinose concentrations over five hours.]] | ||
| + | [[File:Lux_graph_33-4.png|500px|thumb|center|Luciferase luminescence of the lux operon in supercompetent DH5 E. coli cells, detected by a Tecan M200 plate reader at 34.4 °C and six arabinose concentrations over five hours.]] | ||
| + | <br /> | ||
| + | By looking at how luciferase luminescence from the lux operon developed with time after induction by arabinose, we established that a temperature of 29 °C was optimal. This can be seen in the graph below: | ||
| + | <br /> | ||
| + | [[File:Lux_trend.png|500px|thumb|center|Average values of luciferase luminescence of the lux operon in supercompetent DH5 E. coli cells, detected by a Tecan M200 plate reader at four different temperatures and six arabinose concentrations with time.]] | ||
| + | <br /> | ||
| + | |||
| + | <b>Additional Characterization by the SHSID_China Team 2018</b> | ||
| + | <p>The SHSID_China Team 2018 has improved upon the work of the Cambridge iGEM team 2010 and NTNU Trondheim iGEM team 2017, by performing additional experiments to determine the optimal conditions of maximum light emission to add further information and build upon the characterization of part BBa_K325909. As this part is provided in the 2018 Distribution Kit, we tested the effectiveness of the arabinose induced <em>lux</em> operon-encoded enzymatic pathway for light emission in part BBa_K325909 by transforming it into E. coli, incubating overnight and subjecting it to different final concentrations of arabinose over a course of five hours. Reflecting upon the previous characterizations, we chose a temperature of 29℃ to perform our experiments, since by looking at the information presented by the NTNU Trondheim iGEM team 2017, it seems as if 29℃ may provide us with the brightest results.</p> | ||
| + | <p>Luminometric and photometric measurements were taken with a Thermo Scientific Varioskan Flask 4.00.53 every hour throughout the experiment, and the data was recorded on the raw data table provided below.</p> | ||
| + | https://static.igem.org/mediawiki/2018/thumb/9/9b/T--SHSID_China--imp1.png/800px-T--SHSID_China--imp1.png | ||
| + | <p>The data is analyzed using the formula: (Average Luminescence/0Hrs Luminescence)/(Average Abs/0Hrs Abs) for each concentration of arabinose, and the graphs are plotted below, with part BBa_K325909 abbreviated as 4L:</p> | ||
| + | https://static.igem.org/mediawiki/2018/thumb/3/33/T--SHSID_China--imp2.png/320px-T--SHSID_China--imp2.png | ||
| + | <p>From the graph, even though the relative luminescence of the bacteria are similar during the first four hours, there is a significant increase in relative luminescence during the fifth hour at a concentration of 0.1M, therefore we have concluded that inducing part BBa_K325909 with 0.1M arabinose will result in the most intense light emission.</p> | ||
| + | |||
| + | ==IMPROVEMENT REFERENCE: SHSID_China 2018 == | ||
| + | |||
| + | The part was linked with a luxG protein to improve the bioluminescence strength in bacteria. The new part is : <html><a href='https://parts.igem.org/Part:BBa_K2836002'>BBa_K2836002</a></html>(Group: iGEM18 SHSID_China), Designed by: Chang Wei-Tse. | ||
| + | |||
| + | ==Pictures== | ||
| + | |||
| + | [[Image:250px-Cambridge-iGEMpixels.jpg|thumb|569px|center|'''Figure 1 - E.coli cells (Invitrogen TOP 10) transformed with [https://parts.igem.org/Part:BBa_K325909 BBa K325909] in a 96 well plate. ''']] | ||
</div> | </div> | ||
'''References'''<br> | '''References'''<br> | ||
| − | [http://www.jstor.org/stable/4449975 '''[1]:'''] J. Slock, (1995) Transformation Experiments | + | [http://www.jstor.org/stable/4449975 '''[1]:'''] J. Slock, (1995) Transformation Experiments Using Bioluminescence Genes of ''Vibrio fischeri'',''The American Biology Teacher'', '''57''', 225-227. |
| + | </div> | ||
| + | [http://www.annualreviews.org/doi/pdf/10.1146/annurev.mi.42.100188.001055 '''[2]:'''] E.A. Meighen (1988) Enzymes and genes from the ''lux'' operons of bioluminescent bacteria, ''Annual Reviews in Microbiology'' '''42''', 151-176. | ||
| + | [http://www.annualreviews.org/doi/pdf/10.1146/annurev.ge.28.120194.001001 '''[3]:'''] E.A. Meighen, (1994) Genetics of bacterial bioluminescence, ''Annual Reviews of Genetics'', '''28''', 117-139. | ||
| + | Following is a contribution to this part, by the NUS_Singapore 2019 team. | ||
| + | |||
| + | |||
| + | |||
| + | <br><br><br> | ||
| + | <div style="font-weight:300; font-size:24px; color:blue"><i>Following is a contribution to this part, by the NUS_Singapore 2019 team.</i></div> | ||
| + | <br> | ||
| + | |||
| + | ===Group=== | ||
| + | NUS_Singapore 2019 | ||
| + | |||
| + | ===Summary and Uploads=== | ||
| + | Team NUS Singapore 2019 has contributed additional characterization data for biobrick BBa_K325909 by characterizing this biobrick in two different <i>E. coli</i> strains: MG1655 and 10-beta. To demonstrate the functionality of BBa K325909, the team added arabinose concentrations of 0.2% and 2% proposed by earlier iGEM teams to induce both 10-beta and MG1655 at late exponential phase, at an OD<sub>600</sub> of approximately 0.9. Right after the induction, the 96-well plate was read for 20h continuously at 29°C in a microplate reader. | ||
| + | <br><br>Interestingly, the team observed that for both arabinose concentrations, MG1655 produced luminescence at a higher intensity compared to 10-beta over time. Additionally, the duration of luminescence production was approximately twice as long in MG1655 compared to 10-beta when 0.2% arabinose was added. Looking at the growth curves of both strains induced with 0.2% and 2% arabinose, it appears that the maximum growth of each strain does contribute to the observed luminescence graphs. | ||
| + | <br><br> | ||
| + | <html> | ||
| + | <img style="width:400px" src="https://2019.igem.org/wiki/images/0/0c/T--NUS_Singapore--PartsRegistry_Lux1.png"> | ||
| + | </html> | ||
| + | <br><i>Figure 1: Growth curve of 10-beta and MG1655 right after 0.2% arabinose induction at 0h</i> | ||
| + | <html> | ||
| + | <img style="width:400px" src="https://2019.igem.org/wiki/images/0/0c/T--NUS_Singapore--PartsRegistry_Lux2.png"> | ||
| + | </html> | ||
| + | <br><i>Figure 2: Total luminescence produced by 10-beta and MG1655 right after 0.2% arabinose induction at 0h</i> | ||
| + | <html> | ||
| + | <img style="width:400px" src="https://2019.igem.org/wiki/images/d/d7/T--NUS_Singapore--PartsRegistry_Lux3.png"> | ||
| + | </html> | ||
| + | <br><i>Figure 3: Growth curve of 10-beta and MG1655 right after 2% arabinose induction at 0h</i> | ||
| + | <html> | ||
| + | <img style="width:400px" src="https://2019.igem.org/wiki/images/4/40/T--NUS_Singapore--PartsRegistry_Lux4.png"> | ||
| + | </html> | ||
| + | <br><i>Figure 4: Total luminescence produced by 10-beta and MG1655 right after 2% arabinose induction at 0h</i> | ||
| + | <br><br>Taken together, these results show that MG1655, which has never been used to characterize BBa_K325909 activity previously, demonstrates enhanced and prolonged luminescence production compared to the commonly used strain - 10-beta; and this is likely due to a higher maximum growth in the former strain. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ==Functional Parameters: Austin_UTexas== | ||
| + | <html> | ||
| + | <body> | ||
| + | <h3><center>Burden Imposed by this Part:</center></h3> | ||
| + | <figure> | ||
| + | <div class = "center"> | ||
| + | <center><img src = "https://static.igem.org/mediawiki/parts/4/43/T--Austin_Utexas--Low_significant_burden.png" style = "width:200px;height:120px"></center> | ||
</div> | </div> | ||
| − | + | <figcaption><center><b>Burden Value: 8.3 ± 2.2% </b></center></figcaption> | |
| − | + | </figure> | |
| + | <p> Burden is the percent reduction in the growth rate of <i>E. coli</i> cells transformed with a plasmid containing this BioBrick (± values are 95% confidence limits). This part exhibited a significant burden. Users should be aware that BioBricks with a burden of >20-30% may be susceptible to mutating to become less functional or nonfunctional as an evolutionary consequence of this fitness cost. This risk increases as they used for more bacterial cell divisions or in larger cultures. Users should be especially careful when combining multiple burdensome parts, as plasmids with a total burden of >40% are expected to mutate so quickly that they become unclonable. Refer to any one of the | ||
| + | <a href="https://parts.igem.org/Part:BBa_K3174002">BBa_K3174002</a> - <a href="https://parts.igem.org/Part:BBa_K3174007">BBa_K3174007</a> pages for more information on the methods and other conclusions from a large-scale measurement project conducted by the <a href="https://2019.igem.org/Team:Austin_UTexas">2019 Austin_UTexas team.</a></p> | ||
| + | <p>This functional parameter was added by the <a href="https://2020.igem.org/Team:Austin_UTexas/Contribution">2020 Austin_UTexas team.</a></p> | ||
| + | </p> | ||
| + | </body> | ||
| + | </html> | ||
Latest revision as of 00:00, 4 September 2020
Lux Operon (under pBAD)
Vibrio Fischeri
|
|

|
Description
This page described the lux operon from Vibrio fischeri. To relieve LuxR control we placed Lux C, D, A, B, E under the pBad promoter. A more complete description can be found on the [http://2010.igem.org/Team:Cambridge/Bioluminescence/G28 Cambridge iGEM 2010 website].
This Bacterial lux operons encodes five enzymes involved in the light-generating pathway. LuxA and LuxB encode the two subunits of the bacterial luciferase, while the products of LuxC, LuxD and LuxE synthesise the substrate for the light emitting reaction, tetradecanal. The exact function of LuxG is unknown, and it appears to be non-essential for light emission, but its presence increases light output.
In nature, the lux genes appear to be repressed by the nucleoid protein, H-NS, and occur under quorum sensing control. We removed the natural quorum sensing control to facilitate use of the part in biosensors under different regulatory inputs.
As of October 2010 we believe this is the first and only BioBrick to emit light in normal E. coli strains without the addition of any external substrate.
Measured by the [http://2010.igem.org/Team:Cambridge Cambridge iGEM team 2010]
Compatibility
Chassis: Device has been shown to work in Top 10 (Invitrogen), [http://www.ecoliwiki.net/colipedia/index.php/BW25113 BW25113 DELTA H-NS::kan] and H-NS mutant JM 230 H-NS -205::tn10.br>
Plasmids: Device has been shown to work on pSB1C3
Additional characterisation by the [http://2015.igem.org/Team:Penn Penn iGEM team 2015]
Penn iGEM 2015 used this part to make the sender cells in light-mediated cell communication. The lux operon BioBrick on PSB1C3 was transformed into NEB 10-beta competent cells (since it is a commonly used E. coli strain in iGEM) and an H-NS mutant E. coli strain (since the protein has been shown to reduce lux gene expression). The luminescent signal of the two sender strains was compared over time to determine what luminescent output trend would be best for inducing our light-sensitive receiver. Furthermore, using a calibrated conversion system designed by the team, Penn IGEM was about to convert the output from relative lights units (a measure typically used to calculate bioluminescent signals) to an absolute measure in uW/cm^2. Using the absolute units will assist in making the characterization of bioluminescent parts such as BBa_K325909 more consistent across luminometer devices and across projects. For more information, please follow this [http://2015.igem.org/Team:Penn/Sender link].
Additional characterisation by the [http://2017.igem.org/Team:NTNU_Trondheim NTNU Trondheim iGEM team 2017]
Team NTNU Trondheim 2017 has improved the characterisation of the biobrick BBa_K325909, which is an operon consisting of Lux C, D, A, B, E under the arabinose-induced promoter pBAD.
By looking at previous characterisation results of the biobrick, we thought it would work reasonably to use DH5α competent E. coli cells and an incubation temperature of 37 °C. However, when testing with 0.01M arabinose concentrations, no observable luminescence was detected at any time after induction. Therefore, we decided to characterise the biobrick by measuring the luminescence from the lux operon at different arabinose concentrations and temperatures. We measured the luminescence with a Tecan M200 plate reader, using four different temperatures and six different arabinose concentrations. As the machine could only perform heating and not cooling, it was challenging to measure at temperatures below 29 °C.
Results from our experiments are shown in the graphs below. As no luminescence was detected at 37 °C, these graphs are not included.
By looking at how luciferase luminescence from the lux operon developed with time after induction by arabinose, we established that a temperature of 29 °C was optimal. This can be seen in the graph below:
Additional Characterization by the SHSID_China Team 2018
The SHSID_China Team 2018 has improved upon the work of the Cambridge iGEM team 2010 and NTNU Trondheim iGEM team 2017, by performing additional experiments to determine the optimal conditions of maximum light emission to add further information and build upon the characterization of part BBa_K325909. As this part is provided in the 2018 Distribution Kit, we tested the effectiveness of the arabinose induced lux operon-encoded enzymatic pathway for light emission in part BBa_K325909 by transforming it into E. coli, incubating overnight and subjecting it to different final concentrations of arabinose over a course of five hours. Reflecting upon the previous characterizations, we chose a temperature of 29℃ to perform our experiments, since by looking at the information presented by the NTNU Trondheim iGEM team 2017, it seems as if 29℃ may provide us with the brightest results.
Luminometric and photometric measurements were taken with a Thermo Scientific Varioskan Flask 4.00.53 every hour throughout the experiment, and the data was recorded on the raw data table provided below.

The data is analyzed using the formula: (Average Luminescence/0Hrs Luminescence)/(Average Abs/0Hrs Abs) for each concentration of arabinose, and the graphs are plotted below, with part BBa_K325909 abbreviated as 4L:

From the graph, even though the relative luminescence of the bacteria are similar during the first four hours, there is a significant increase in relative luminescence during the fifth hour at a concentration of 0.1M, therefore we have concluded that inducing part BBa_K325909 with 0.1M arabinose will result in the most intense light emission.
Contents
IMPROVEMENT REFERENCE: SHSID_China 2018
The part was linked with a luxG protein to improve the bioluminescence strength in bacteria. The new part is : BBa_K2836002(Group: iGEM18 SHSID_China), Designed by: Chang Wei-Tse.
Pictures
References
[http://www.jstor.org/stable/4449975 [1]:] J. Slock, (1995) Transformation Experiments Using Bioluminescence Genes of Vibrio fischeri,The American Biology Teacher, 57, 225-227.
[http://www.annualreviews.org/doi/pdf/10.1146/annurev.mi.42.100188.001055 [2]:] E.A. Meighen (1988) Enzymes and genes from the lux operons of bioluminescent bacteria, Annual Reviews in Microbiology 42, 151-176. [http://www.annualreviews.org/doi/pdf/10.1146/annurev.ge.28.120194.001001 [3]:] E.A. Meighen, (1994) Genetics of bacterial bioluminescence, Annual Reviews of Genetics, 28, 117-139. Following is a contribution to this part, by the NUS_Singapore 2019 team.
Group
NUS_Singapore 2019
Summary and Uploads
Team NUS Singapore 2019 has contributed additional characterization data for biobrick BBa_K325909 by characterizing this biobrick in two different E. coli strains: MG1655 and 10-beta. To demonstrate the functionality of BBa K325909, the team added arabinose concentrations of 0.2% and 2% proposed by earlier iGEM teams to induce both 10-beta and MG1655 at late exponential phase, at an OD600 of approximately 0.9. Right after the induction, the 96-well plate was read for 20h continuously at 29°C in a microplate reader.
Interestingly, the team observed that for both arabinose concentrations, MG1655 produced luminescence at a higher intensity compared to 10-beta over time. Additionally, the duration of luminescence production was approximately twice as long in MG1655 compared to 10-beta when 0.2% arabinose was added. Looking at the growth curves of both strains induced with 0.2% and 2% arabinose, it appears that the maximum growth of each strain does contribute to the observed luminescence graphs.

Figure 1: Growth curve of 10-beta and MG1655 right after 0.2% arabinose induction at 0h

Figure 2: Total luminescence produced by 10-beta and MG1655 right after 0.2% arabinose induction at 0h

Figure 3: Growth curve of 10-beta and MG1655 right after 2% arabinose induction at 0h

Figure 4: Total luminescence produced by 10-beta and MG1655 right after 2% arabinose induction at 0h
Taken together, these results show that MG1655, which has never been used to characterize BBa_K325909 activity previously, demonstrates enhanced and prolonged luminescence production compared to the commonly used strain - 10-beta; and this is likely due to a higher maximum growth in the former strain.
Functional Parameters: Austin_UTexas
Burden Imposed by this Part:

Burden is the percent reduction in the growth rate of E. coli cells transformed with a plasmid containing this BioBrick (± values are 95% confidence limits). This part exhibited a significant burden. Users should be aware that BioBricks with a burden of >20-30% may be susceptible to mutating to become less functional or nonfunctional as an evolutionary consequence of this fitness cost. This risk increases as they used for more bacterial cell divisions or in larger cultures. Users should be especially careful when combining multiple burdensome parts, as plasmids with a total burden of >40% are expected to mutate so quickly that they become unclonable. Refer to any one of the BBa_K3174002 - BBa_K3174007 pages for more information on the methods and other conclusions from a large-scale measurement project conducted by the 2019 Austin_UTexas team.
This functional parameter was added by the 2020 Austin_UTexas team.