Difference between revisions of "Part:BBa K4632024"
(→Construction and Characterization) |
|||
| (21 intermediate revisions by the same user not shown) | |||
| Line 18: | Line 18: | ||
<p>To achieve control over toxicant concentration, we introduced components labeled with '''BBa_K4632016''' [https://parts.igem.org/Part:BBa_K4632016] and '''BBa_K112806'''[https://parts.igem.org/Part:BBa_K112806] to create the T4-T4 lysis device. '''BBa_K4632016''' was originally from '''BBa_K112805'''[https://parts.igem.org/Part:BBa_K112805]. Its Codon was optimized for ''Escherichia coli'' expression.</p> | <p>To achieve control over toxicant concentration, we introduced components labeled with '''BBa_K4632016''' [https://parts.igem.org/Part:BBa_K4632016] and '''BBa_K112806'''[https://parts.igem.org/Part:BBa_K112806] to create the T4-T4 lysis device. '''BBa_K4632016''' was originally from '''BBa_K112805'''[https://parts.igem.org/Part:BBa_K112805]. Its Codon was optimized for ''Escherichia coli'' expression.</p> | ||
| − | <p> | + | <p>In our initial validation experiments, we utilized a dual-plasmid system consisting of pBAD24M and pBAD33 to test our device. (Plasmid maps can be found in Figure 1)</p> |
| − | + | ||
| − | + | ||
| + | ''https://static.igem.wiki/teams/4632/wiki/wiki/registry-part/part-1-1-6-1.png'' | ||
| + | <p><strong>Figure 1. </strong>Diagram of Quorum sensing-based T4 lysis device verification systems circuit design</p> | ||
<p>We characterized the component to demonstrate its effectiveness, as detailed in the construction and characterization section.</p> | <p>We characterized the component to demonstrate its effectiveness, as detailed in the construction and characterization section.</p> | ||
| + | |||
| + | <p>The Quorum sensing-based T4 lysis device verification systems were transformed into Top10 cells and conducted induction experiments using Ara. At regular intervals, the OD600 values and eGFP fluorescence intensity were measured using a microplate reader. A curve with OD600 were plotted as the vertical axis and induction time were plotted as the horizontal axis. If the OD600 value were observed an significantly decreases after the initially increaseing, the expression of the lysis gene could be confrimed. We assessed whether the pathway could limit the maximum expression level based on the trend in eGFP signal changes. If the eGFP expression rate significantly decreases or ceases to increase after the appearance of lysis, the verification is successful. The limitation of the maximum expression level was assessed by the trend of eGFP signal change.</p> | ||
===Sequence and Features=== | ===Sequence and Features=== | ||
| Line 35: | Line 37: | ||
===Construction and Characterization=== | ===Construction and Characterization=== | ||
---- | ---- | ||
| − | |||
| − | |||
| − | |||
| − | |||
'''1. Construction''' | '''1. Construction''' | ||
| − | <p> | + | <p>The complete sequences, synthesized (ordered from Guangzhou IGE Biotechnology Co.,Ltd.) and assembled onto respective plasmids, have been validated through sequencing. Both of the plasmids were simultaneously introduced into E.coli Top10 using the KCM ice method. Successful transformation was confirmed by clone PCR. Universal primers for the pBAD series plasmids were employed to confime the transformation of pBAD24M-paraBAD-lasⅠ-lasR. Amplification of bands corresponding to the size in pBAD24M for transformation plasmids resulted in bands of 1845 bp. This confirms the successful introduction of the pBAD24M plasmids.(Figure 2)</p> |
| − | <p> | + | ''https://static.igem.wiki/teams/4632/wiki/wiki/registry-part/77777777777.png'' |
| + | <p><strong>Figure 2. </strong> Colony PCR of co-transformation by universal primers. </p> | ||
| + | <p>Lane 3: Co-transformation of pBAD24M-paraBAD-lasⅠ-lasR and pBAD33-paraBAD-eGFP-plas-T4 holin-T4 endolysin</p> | ||
| + | <p>The specific primers PCR was also performed to obtain target-sized fragments. The plasmid containing paraBAD-eGFP-plas-T4 holin-T4 endolysin produced a 1296 bp fragment (Figure 3). This confirms the successful introduction of pBAD33 plasmids.</p> | ||
| − | ''https://static.igem.wiki/teams/4632/wiki/wiki/registry-part/ | + | ''https://static.igem.wiki/teams/4632/wiki/wiki/registry-part/8888888.png'' |
| − | <p><strong> | + | <p><strong>Figure 3. </strong> Colony PCR of co-transformation by specific primers. </p> |
| + | <p>Use forward primer (AGGCGGTCGTTGCTAATA) and reverse primer (AAACTCGTGCGGAGGTAA) as specific primer for pBAD33-paraBAD-eGFP-plas-T4 holin-T4 endolysin. lane 3: pBAD33-paraBAD-eGFP-plas-T4 holin-T4 endolysin; lane 4: Co-transformation of pBAD24M-paraBAD-lasⅠ-lasR and pBAD33-paraBAD-eGFP-plas-T4 holin-T4 endolysin</p> | ||
| − | '''2. Validation of Product Expression Level Control''' | + | '''2. Validation of Product Expression Level Control Using Quorum sensing-based T4 lysis device verification systems''' |
| − | + | ||
'''(1)Pre-experiment for Induced Expression of Lysis Effect''' | '''(1)Pre-experiment for Induced Expression of Lysis Effect''' | ||
The successfully transformed engineered bacteria were streaked on plates and grown at 37°C. Single colonies were picked and inoculated into liquid LB medium, followed by the addition of inducers. The cultures were incubated for 6 hours. | The successfully transformed engineered bacteria were streaked on plates and grown at 37°C. Single colonies were picked and inoculated into liquid LB medium, followed by the addition of inducers. The cultures were incubated for 6 hours. | ||
| − | ''https://static.igem.wiki/teams/4632/wiki/wiki/registry-part/ | + | ''https://static.igem.wiki/teams/4632/wiki/wiki/registry-part/100000000000.png'' |
| − | <p><strong>Fig. | + | <p><strong>Fig.4</strong> Verification of lysis effect</p> |
| − | <p>The blank control was LB with 20% Ara. The control group consisted of the engineered bacteria without inducer, while the experimental group consisted of the engineered bacteria with a final concentration of 0.02% Ara. Each group had 3 replicates | + | <p>The blank control was LB with 20% Ara. The control group consisted of the second verification system engineered bacteria without inducer, while the experimental group consisted of the second verification system engineered bacteria with a final concentration of 0.02% Ara. Each group had 3 replicates</p> |
'''Result''' | '''Result''' | ||
| − | <p>After zeroing with the blank control, it was observed that the | + | <p>After zeroing with the blank control, it was observed that the optical density in the experimental group decreased to 34.3% of that in the control group (Fig 4), indicating a significant lysis effect. This confirms the successful expression of quorum sensing and initiation of downstream lysis gene expression. Lysis gene expression was successful without leakage.</p> |
| Line 69: | Line 70: | ||
The blank control group was LB broth, the control group was wild Top10, and the experimental group was the engineering bacteria consisting of pBAD24M and pBAD33. There were 6 replicates per group.</p> | The blank control group was LB broth, the control group was wild Top10, and the experimental group was the engineering bacteria consisting of pBAD24M and pBAD33. There were 6 replicates per group.</p> | ||
| − | ''https://static.igem.wiki/teams/4632/wiki/wiki/registry-part/ | + | ''https://static.igem.wiki/teams/4632/wiki/wiki/registry-part/part-1-5-1.png'' |
| − | <p><strong>Fig. | + | <p><strong>Fig.5</strong> Growth curve testing of the quorum sensing-based T4 lysis device</p> |
| + | <p>The growth curve revealed that the bacterial density continued to rise in the first 4 hours and began to decrease after 4 hours, stabilizing around the 9th hour. In contrast, the control group's bacterial density continued to rise. At the 4th hour, the quorum sensing signal reached the threshold, initiating lysis gene expression. The engineered bacteria lysed, resulting in a significant decrease in bacterial density, which stabilized around the 9th hour (Fig 5).</p> | ||
| + | '''(3) Characterization of Reporter Gene eGFP''' | ||
| − | + | <p>Single colonies of engineered bacteria were selected and inoculated into LB broth. After overnight growth, the OD600 was adjusted to 0.6. Ara at a final concentration of 0.02% was added, and the cultures were incubated for 6 hours. eGFP fluorescence intensity was measured using a microplate reader (excitation wavelength 488 nm, emission wavelength 506 nm).</p> | |
| − | <p> | + | <p>Blank control group: sterile LB, control group: engineered bacteria without 20% ara added, experimental group: engineered bacteria with inducer added, 3 replicates per group. There was no significant difference in fluorescence intensity between the control group and the experimental group, indicating no expression of eGFP</p> |
| + | '''(4) eGFP Protein Expression Experiment''' | ||
| + | |||
| + | <p>Single colonies of engineered bacteria were selected and inoculated into LB medium with inducer for overnight incubation. Top10 subjected to the same procedure served as the control group. Bacterial pellets were collected, and SDS-PAGE experiments were conducted, revealing no expression of eGFP at the protein level.</p> | ||
| + | |||
| + | ''''(5) q-RT-PCR Experiment'''' | ||
| + | <p>q-RT-PCR experiments were conducted under various conditions, confirming no expression at the mRNA level.</p> | ||
| + | |||
| + | <p>The reporter genes could not be expressed. The characterization of reporter genes eGFP and RFP was performed simultaneously with the second verification system. Consistent results were obtained from SDS-PAGE experiments and q-RT-PCR experiments.</p> | ||
| + | |||
| + | '''We hypothesize several possible reasons:''' | ||
| + | |||
| + | <p>1. The selected reporter genes eGFP and RFP may not be suitable for expression in Top10.</p> | ||
| + | <p>2. Errors may have occurred in the design of the gene expression module.</p> | ||
| + | <p>3. Interactions between genes may have occurred.</p> | ||
| + | |||
| + | ===Reference=== | ||
| + | ---- | ||
| + | <p>Jiang W, He X, Luo Y et al. Two Completely Orthogonal Quorum Sensing Systems with Self-Produced Autoinducers Enable Automatic Delayed Cascade Control[J]. ACS SYNTH BIOL, 2020,9(9):2588-2599.</p> | ||
<!-- Uncomment this to enable Functional Parameter display | <!-- Uncomment this to enable Functional Parameter display | ||
Latest revision as of 15:00, 12 October 2023
Quorum sensing-based T4 lysis device
Description
This part is a Quorum Sensing-based T4 lysis device constructed based on the LasI-LasR system and the T4-T4 lysis device.
The LasI-LasR system is a Quorum Sensing system from 'Pseudomonas aeruginosa.' As modified by Wei Jiang et al., it can be entirely orthogonal to the TraI-TraR system , another Quorum Sensing system from 'Agrobacterium tumefaciens' (Jiang et al., 2020).
1. How does it work?
In such a scenario, newly introduced engineered bacteria will promptly trigger the lysis gene, preventing the high-level expression of product expression., which could significantly reduce the accumulation rate of toxin product in the environment, creating a situation similar to a "threshold" for its accumulation.
2. What have we done? (SCAU-China 2023)
pLas was from BBa_K2967001[1] We introduced it to achieve control over drug concentration through bacterial density.
To achieve control over toxicant concentration, we introduced components labeled with BBa_K4632016 [2] and BBa_K112806[3] to create the T4-T4 lysis device. BBa_K4632016 was originally from BBa_K112805[4]. Its Codon was optimized for Escherichia coli expression.
In our initial validation experiments, we utilized a dual-plasmid system consisting of pBAD24M and pBAD33 to test our device. (Plasmid maps can be found in Figure 1)

Figure 1. Diagram of Quorum sensing-based T4 lysis device verification systems circuit design
We characterized the component to demonstrate its effectiveness, as detailed in the construction and characterization section.
The Quorum sensing-based T4 lysis device verification systems were transformed into Top10 cells and conducted induction experiments using Ara. At regular intervals, the OD600 values and eGFP fluorescence intensity were measured using a microplate reader. A curve with OD600 were plotted as the vertical axis and induction time were plotted as the horizontal axis. If the OD600 value were observed an significantly decreases after the initially increaseing, the expression of the lysis gene could be confrimed. We assessed whether the pathway could limit the maximum expression level based on the trend in eGFP signal changes. If the eGFP expression rate significantly decreases or ceases to increase after the appearance of lysis, the verification is successful. The limitation of the maximum expression level was assessed by the trend of eGFP signal change.
Sequence and Features
- 10INCOMPATIBLE WITH RFC[10]Illegal XbaI site found at 84
Illegal XbaI site found at 795
Illegal PstI site found at 367 - 12INCOMPATIBLE WITH RFC[12]Illegal PstI site found at 367
- 21COMPATIBLE WITH RFC[21]
- 23INCOMPATIBLE WITH RFC[23]Illegal XbaI site found at 84
Illegal XbaI site found at 795
Illegal PstI site found at 367 - 25INCOMPATIBLE WITH RFC[25]Illegal XbaI site found at 84
Illegal XbaI site found at 795
Illegal PstI site found at 367
Illegal AgeI site found at 1089
Illegal AgeI site found at 1159 - 1000COMPATIBLE WITH RFC[1000]
Construction and Characterization
1. Construction
The complete sequences, synthesized (ordered from Guangzhou IGE Biotechnology Co.,Ltd.) and assembled onto respective plasmids, have been validated through sequencing. Both of the plasmids were simultaneously introduced into E.coli Top10 using the KCM ice method. Successful transformation was confirmed by clone PCR. Universal primers for the pBAD series plasmids were employed to confime the transformation of pBAD24M-paraBAD-lasⅠ-lasR. Amplification of bands corresponding to the size in pBAD24M for transformation plasmids resulted in bands of 1845 bp. This confirms the successful introduction of the pBAD24M plasmids.(Figure 2)

Figure 2. Colony PCR of co-transformation by universal primers.
Lane 3: Co-transformation of pBAD24M-paraBAD-lasⅠ-lasR and pBAD33-paraBAD-eGFP-plas-T4 holin-T4 endolysin
The specific primers PCR was also performed to obtain target-sized fragments. The plasmid containing paraBAD-eGFP-plas-T4 holin-T4 endolysin produced a 1296 bp fragment (Figure 3). This confirms the successful introduction of pBAD33 plasmids.
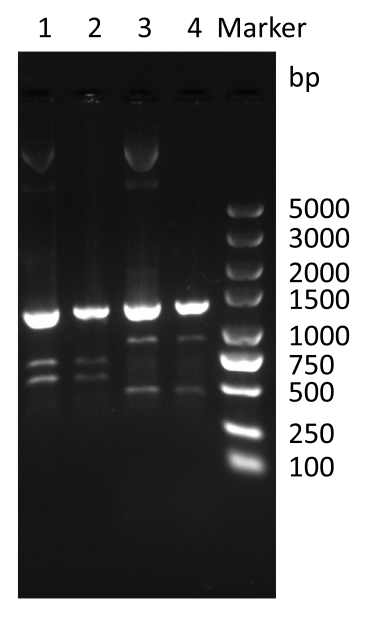
Figure 3. Colony PCR of co-transformation by specific primers.
Use forward primer (AGGCGGTCGTTGCTAATA) and reverse primer (AAACTCGTGCGGAGGTAA) as specific primer for pBAD33-paraBAD-eGFP-plas-T4 holin-T4 endolysin. lane 3: pBAD33-paraBAD-eGFP-plas-T4 holin-T4 endolysin; lane 4: Co-transformation of pBAD24M-paraBAD-lasⅠ-lasR and pBAD33-paraBAD-eGFP-plas-T4 holin-T4 endolysin
2. Validation of Product Expression Level Control Using Quorum sensing-based T4 lysis device verification systems
(1)Pre-experiment for Induced Expression of Lysis Effect The successfully transformed engineered bacteria were streaked on plates and grown at 37°C. Single colonies were picked and inoculated into liquid LB medium, followed by the addition of inducers. The cultures were incubated for 6 hours.

Fig.4 Verification of lysis effect
The blank control was LB with 20% Ara. The control group consisted of the second verification system engineered bacteria without inducer, while the experimental group consisted of the second verification system engineered bacteria with a final concentration of 0.02% Ara. Each group had 3 replicates
Result
After zeroing with the blank control, it was observed that the optical density in the experimental group decreased to 34.3% of that in the control group (Fig 4), indicating a significant lysis effect. This confirms the successful expression of quorum sensing and initiation of downstream lysis gene expression. Lysis gene expression was successful without leakage.
(2)Growth Curve Testing of the Second Verification System
Single clones were selected and inoculated into LB medium. After overnight incubation, the OD600 was adjusted to 0.6, and arabinose was added to a final concentration of 0.02%. The cultures were shaken for 21 hours in a sterile 96-well plate, and a growth curve was plotted. The blank control group was LB broth, the control group was wild Top10, and the experimental group was the engineering bacteria consisting of pBAD24M and pBAD33. There were 6 replicates per group.

Fig.5 Growth curve testing of the quorum sensing-based T4 lysis device
The growth curve revealed that the bacterial density continued to rise in the first 4 hours and began to decrease after 4 hours, stabilizing around the 9th hour. In contrast, the control group's bacterial density continued to rise. At the 4th hour, the quorum sensing signal reached the threshold, initiating lysis gene expression. The engineered bacteria lysed, resulting in a significant decrease in bacterial density, which stabilized around the 9th hour (Fig 5).
(3) Characterization of Reporter Gene eGFP
Single colonies of engineered bacteria were selected and inoculated into LB broth. After overnight growth, the OD600 was adjusted to 0.6. Ara at a final concentration of 0.02% was added, and the cultures were incubated for 6 hours. eGFP fluorescence intensity was measured using a microplate reader (excitation wavelength 488 nm, emission wavelength 506 nm).
Blank control group: sterile LB, control group: engineered bacteria without 20% ara added, experimental group: engineered bacteria with inducer added, 3 replicates per group. There was no significant difference in fluorescence intensity between the control group and the experimental group, indicating no expression of eGFP
(4) eGFP Protein Expression Experiment
Single colonies of engineered bacteria were selected and inoculated into LB medium with inducer for overnight incubation. Top10 subjected to the same procedure served as the control group. Bacterial pellets were collected, and SDS-PAGE experiments were conducted, revealing no expression of eGFP at the protein level.
'(5) q-RT-PCR Experiment'
q-RT-PCR experiments were conducted under various conditions, confirming no expression at the mRNA level.
The reporter genes could not be expressed. The characterization of reporter genes eGFP and RFP was performed simultaneously with the second verification system. Consistent results were obtained from SDS-PAGE experiments and q-RT-PCR experiments.
We hypothesize several possible reasons:
1. The selected reporter genes eGFP and RFP may not be suitable for expression in Top10.
2. Errors may have occurred in the design of the gene expression module.
3. Interactions between genes may have occurred.
Reference
Jiang W, He X, Luo Y et al. Two Completely Orthogonal Quorum Sensing Systems with Self-Produced Autoinducers Enable Automatic Delayed Cascade Control[J]. ACS SYNTH BIOL, 2020,9(9):2588-2599.
