Assembly standard 23
Pam Silver's lab has developed Assembly standard 23, often called the Silver standard, for assembling protein domains. It relies on shortening the BioBrick prefix and suffix each by 1 base pair such that the resulting SpeI-XbaI scar is only 6 base pairs long and protein domains can be assembled in frame.
As shown in figure 3 (right), Assembly standard 23 results in the inclusion of a Thr-Arg (ACT AGA) two amino acid junction between protein domains.
One disadvantage of the Assembly standard 23 is that the codon AGA which encodes Arginine is a rare codon in E. coli. Rare codons can prevent over-expression of proteins in E. coli.
See [http://openwetware.org/wiki/The_BioBricks_Foundation:Standards/Technical/Formats The BioBricks Foundation wiki] for a discussion and comparison of different technical standards.
| Plasmid backbones (?) | Protein domains (?) |
Or get help on Assembly standard 23 parts.
Plasmid backbones
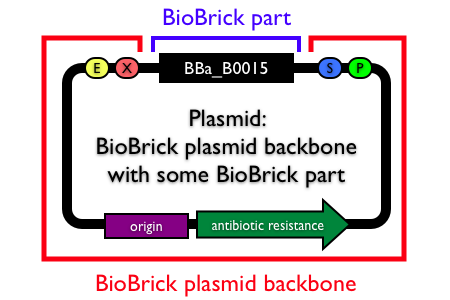
Plasmids are circular, double-stranded DNA molecules typically containing a few thousand base pairs that replicate within the cell independently of the chromosomal DNA. Plasmid DNA is easily purified from cells, manipulated using common lab techniques and incorporated into cells. Most BioBrick parts in the Registry are maintained and propagated on plasmids. Thus, construction of BioBrick parts, devices and systems usually requires working with plasmids.
Note: In the Registry, plasmids are made up of two distinct components:
- the BioBrick part, device or system that is located in the BioBrick cloning site, between (and excluding) the BioBrick prefix and suffix.
- the plasmid backbone which propagates the BioBrick part. The plasmid backbone is defined as the sequence beginning with the BioBrick suffix, including the replication origin and antibiotic resistance marker, and ending with the BioBrick prefix. [Note that the plasmid backbone itself can be composed of BioBrick parts.]
Many BioBrick parts in the Registry are maintained on more than one plasmid backbone!
| Caroline Ajo-Franklin developed the Silver lab protein fusion vectors BBa_J63009 and BBa_J63010 in Pam Silver's lab. |
There are no parts for this table
Protein domains
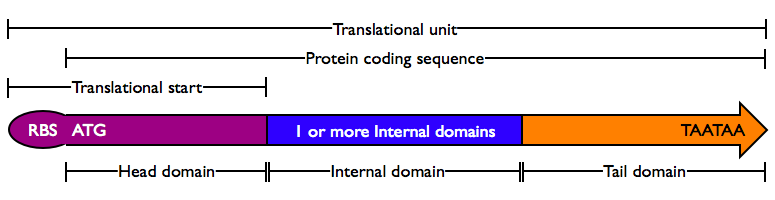
Protein domains encode portions of proteins and can be assembled together to form translational units, a genetic part spanning from translational initiation (the RBS) to translational termination (the stop codon).
There are several different types of protein domains.
- Head Domain: The Head domain consists of the start codon followed immediately by zero or more triplets specifiying an N-terminal tag, such as a protein export tag or lipoprotein binding tag. Head domains should begin with an
ATGstart codon and include codons 2 and 3 of the protein at a minimum. Examples of head domains includeATGstart codonATGstart codon and codons 2-3ATGstart codon and signal sequenceATGstart codon and affinity tag
- Internal Domains: Protein domains consist of a series of codon triplets coding for an amino acid sequence without a start codon or stop codon. Multiple Internal Domains can be fused. Examples of internal domains include
- DNA binding domains
- Dimerization domains
- Kinase domains
- Special Internal Domains: Short Domains with specific function may be separately categorized, but obey the same composition rules as normal internal domains. Examples of special internal domains include
- Linkers
- Cleavage sites
- Inteins
- Tail Domain: The C-terminus of a coding region consists of zero or more triplet codons, followed by a pair of TAA stop codons. In the simplest case, the stop codons terminate the protein with an Stop. More complex Tails may include degradation tags appropriate to the organism (i.e., with different degradation rates). Examples of Tail domain include
TAATAAstop codons- A degradation tag followed by
TAATAAstop codons - An affinity tag followed by
TAATAAstop codon
Unfortunately, the original BioBrick assembly standard, Assembly standard 10, does not support in-frame assembly of protein domains. (Assembly standard 10 creates an 8 bp scar between adjacent parts.) Therefore, it is recommended that you use an alternate approach to assemble protein domains together to make a translational unit. There are several possible approaches to assembling protein domains including direct synthesis (preferred because it creates no scars) as well as various assembly standards. Regardless of which standard you choose, we suggest that the resulting protein coding sequence or translational unit comply with the original BioBrick assembly standard so that your parts can be assembled with most of the parts in the Registry.
Protein coding sequences should be as follows
GAATTC GCGGCCGC T TCTAG [ATG ... TAA TAA] T ACTAGT A GCGGCCG CTGCAG
Note: Although most RBSs are currently specified as separate parts in the Registry, we are now moving to a new design in which the RBS and Head domain are combined into a single part termed a Translational start. The new design has the advantage of encapsulating both ribosome binding and translational initiation within a single part. Our working hypothesis is that the new design will reduce the likelihood of unexpected functional composition problems between the RBS and coding sequence.
There are no parts for this table
References
A New BioBrick Assembly Strategy Designed for Facile Protein Engineering
MIT SBWG Technical Reports, 2006 Apr 20
Ira Philips, Pam Silver
[http://hdl.handle.net/1721.1/32535 URL] (open access!)