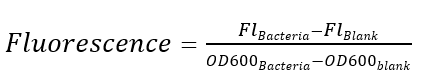Part:BBa_R0051
promoter (lambda cI regulated)
The cI regulated promoter is based on the pR promoter from bacteriophage lambda. The promoter has two DNA binding sites for lambda cI repressor BBa_C0051. cI binding results in repression of transcription. The specific sequence used here is based on the cI repressible promoter used in the Elowitz repressilator (and references therein). [http://google.com f] Intrinsic noise value: 0.0869 (compare with R0010: 0.0707; R0011: 0.0040). See [http://2015.igem.org/Team:William_and_Mary William_and_Mary iGEM 2015]
>Internal Priming Screening Characterization of BBa_R0051: Has no possible internal priming sites between this BioBrick part and the VF2 or the VR primer.
The 2018 Hawaii iGEM team evaluated the 40 most frequently used BioBricks and ran them through an internal priming screening process that we developed using the BLAST program tool. Out of the 40 BioBricks we evaluated, 10 of them showed possible internal priming of either the VF2 or VR primers and sometime even both. The data set has a range of sequence lengths from as small as 12 bases to as large as 1,210 bases. We experienced the issue of possible internal priming during the sequence verification process of our own BBa_K2574001 BioBrick and in the cloning process to express the part as a fusion protein. BBa_K2574001 is a composite part containing a VLP forming Gag protein sequence attached to a frequently used RFP part (BBa_E1010). We conducted a PCR amplification of the Gag-RFP insert using the VF2 and VR primers on the ligation product (pSB1C3 ligated to the Gag + RFP). This amplicon would serve as template for another PCR where we would add the NcoI and BamHI restriction enzyme sites through new primers for ligation into pET14b and subsequent induced expression. Despite gel confirming a rather large, approximately 2.1 kb insert band, our sequencing results with the VR primer and BamHI RFP reverse primer gave mixed results. Both should have displayed the end of the RFP, but the VR primer revealed the end of the Gag. Analysis of the VR primer on the Gag-RFP sequence revealed several sites where the VR primer could have annealed with ~9 - 12 bp of complementarity. Internal priming of forward and reverse primers can be detrimental to an iGEM project because you can never be sure if the desired construct was correctly inserted into the BioBrick plasmid without a successful sequence verification.
Usage and Biology
Strong promoter. [jb, 5/24/04]
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
HZAU-China 2020's contribution
(The following experimental data are from the literature, not the actual data made by the team)
Due to the compression of experimental time this year, we are not able to obtain the actual data of the characterization of PR promoter, so we consulted the literature and found that it depicts some properties of PR promoter. In the literature, the reporter gene LacLM is expressed by PR promoter, and the activity of LacLM is used to characterize the strength of PR promoter. The synthetic promoter expresses CI protein and Pepl reporter gene, and the Pepl activity is used to characterize the expression of CI protein. The LacLM and PepI activities are measured by catalyzing their substrates ortho-nitrophenol-b-galactoside or L-proline-para-nitroanilide, respectively.
In this paper, different synthetic promoters are used to obtain different expression levels of CI protein, and then explore the effect of different concentrations of CI protein on PR promoter. The PL promoter is used to express MOR protein. The active form MOR and its inactive form MORI are used to explore the effect of MOR protein on the inhibition effect of CI protein on PR promoter by CI protein.

* Alsing et al. Biophysical journal, 2011.
Result
Quantitative study on inhibition of PR promoter by CI protein
Some literatures wanted to quantify the inhibitory effect of CI protein on PR promoter, so they established a model of in vitro transcription which use the super helix DNA with wild-type PR promoter as template. With the concentration of CI protein gradually increased in vitro, the inhibitory effect of CI protein on PR promoter is quantitatively explored by measuring the transcription quantity of PR promoter.

* Lewis et al. Proceedings of the National Academy of Sciences, 2011.
Result
In the literature, we can see that in the in vitro transcription system, when the concentration of CI protein added reaches 50 nM , the transcription amount of PR promoter is half of the initial value, and when the concentration of CI protein added reaches 100 mM, the inhibition of PR transcription reaches the maximum.
[1]Alsing A, Pedersen M, Sneppen K, et al. Key players in the genetic switch of bacteriophage TP901-1[J]. Biophysical journal, 2011, 100(2): 313-321.
[2] Lewis D, Le P, Zurla C, et al. Multilevel autoregulation of λ repressor protein CI by DNA looping in vitro[J]. Proceedings of the National Academy of Sciences, 2011, 108(36): 14807-14812.
===Tongji_China 2020===

Measured by excitation wavelength of 588 nm, emission wavelength of 635 nm. Relative fluorescence is calculated by the formula below, LB medium as blank. As shown in the figure, the expression strength of R0051 (without cI repression) is 4.16 times to J04450 (Lac promoter).

Contribution from iGEM 2021 SZPT-China
Group: iGEM 2021 SZPT-China
Author: Yuetong Chen
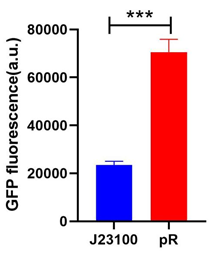
To compare the transcriptional strength between the pR promoter and a constitutive promoter J23100 (BBa_J23100), two plasmids E. coli-DH5α-pR-B0034-sfGFP and E. coli-DH5α-J23100-B0034-sfGFP were firstly constructed, and the reporter gene transcription (Superfolder GFP) controlled by either of two plasmids was determined by measuring the average fluorescence intensity. To simplify the comparison, we set the strength of J23100 promoter as 1 and normalized the strength of pR to J23100.
Excitation is induced at wavelength of 488 nm and the intensity of the emitted fluorescence of wavelength at 520 nm is measured as a function of transcription strength. Fluorescence is calculated by the formula below, LB medium is set as the blank control.
Results
As the figure shows, the expression strength of pR is 3.00 times to that of J23100.
//direction/forward
//promoter
//regulation/negative
//rnap/prokaryote/ecoli/sigma70
| control | lambda cI |
| direction | Forward |
| negative_regulators | 1 |
| o_h | |
| o_l | |
| positive_regulators |

 1 Registry Star
1 Registry Star