Part:BBa_K539119
crtY (lycopene cyclase)
Toulouse_INSA-UPS 2020contributed to the characterisation of this part by adding a new documentation learned form literature on the expression and stability of CrtI.
(--antonmykhailiuk 19:10, 08 October 2020 (UTC+2))
This is the coding sequence of crtY from Pantoea ananatis (formerly Erwinia uredovora) (Accession number D90087). It encodes lycopene B-cyclase, part of the carotenoid biosynthesis pathway, which converts lycopene to B-carotene (Misawa, N., Nakagawa, N., Kobayashi, K., Yamano, S., Nakamura, K., and Harashima, K. 1990. Elucidation of the Erwinia uredovora carotenoid biosynthetic pathway by functional analysis of gene products expressed in Escherichia coli. Journal of Bacteriology 172, 6704-612).
We found a point mutation of [BBa_K118008] [1](iGEM08_Edinburgh) at SpeI site that mutate into ACCTAGT. However, We corrected it successfully(ACCTAGT->ACTAGT), and submited to igem.
The following is the gel electrophoresis result of crtY Part:BBa_K118008. The sample included are uncut plasmid, cut by EcoR1, Xba1, Spe1 and Pst1 respctivly.

We can see that the location of plasmid cut by Spe1 is the same as uncut plasmid. We assumed that Spe1 site has mutated.
After the Point Mutation, we correct the Spe1 site and the gel electrophoresis result is shown below.

Please refer to the Experience to surf more:BBa_K118008:Experience
Contribution from other teams
Toulouse_INSA-UPS 2020's contribution
Characterisation
Since the CrtI (phytoene desaturase) is a part of the biosynthesis pathway of carotenoids (fig. 1), it is often co-expressed with the other enzymes of the pathway: such as CrtB or CrtY. Rabeharindranto et al. analyzed the expression of single domain CrtI, CrtY, and CrtB (fig.2). CrtB is unambiguously detected as an intense bands in all strains B, B/I or Y/B/I. On the other hand, CrtI can be observed at the expected size (67kDa) only when coexpressed with CrtY domain (strain Y/B/I). The explanation could be that CrtI protein needs to be co-expressed together with CrtY to have a normal production/stability. A faint band with 50kDa migration pattern could be observed in B/I strain which could further support the idea of low production or stability of CrtI in absence of CrtY.
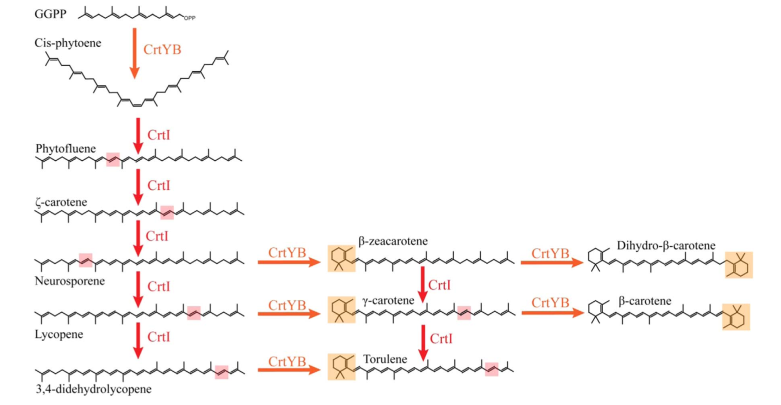
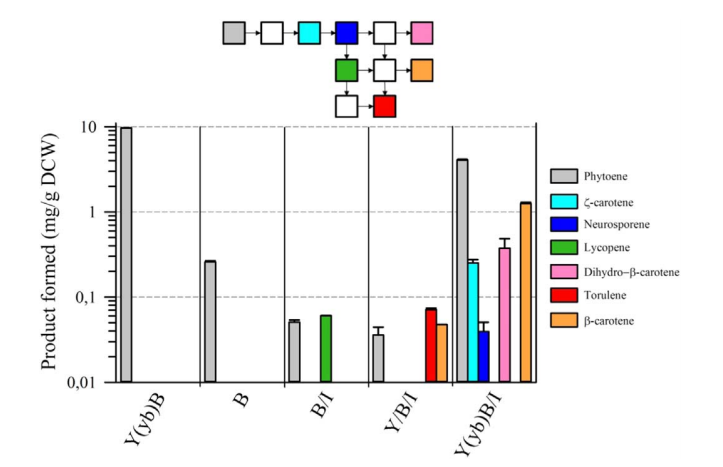
The second point concerns the presence of a natural fusion of CrtY (lycopene cyclase) and CrtB(phytoene synthase). Although, in eucaryotes, CrtB enzyme is predicted to be cytosolic and CrtY enzyme is predicted to be transmembrane [2], surprisingly, there is a natural fusion between CrtY and CrtB which gives CrtYB enzyme[3]. There is a strong opinion on the importance of this natural fusion on the activity of phytoene synthase [4,5]. Rabeharindranto et al. confirmed the impact of CrtY and CrtB splitting on the phytoene production as it has significantly decreased (40 times) in the B strain compared to the Y(yb)B strain (fig. 3).
References for Toulouse_INSA-UPS 2020's contribution
- [1]Rabeharindranto, H., Castaño-Cerezo, S., Lautier, T., Garcia-Alles, L. F., Treitz, C., Tholey, A., & Truan, G. (2019). Enzyme-fusion strategies for redirecting and improving carotenoid synthesis in S. cerevisiae. Metabolic Engineering Communications, 8, e00086. https://doi.org/10.1016/j.mec.2019.e00086
- [2]Schaub, P., Yu, Q., Gemmecker, S., Poussin-Courmontagne, P., Mailliot, J., McEwen, A.G., et al., 2012. On the structure and function of the phytoene desaturase CRTI from Pantoea ananatis, a membrane-peripheral and FAD-dependent oxidase/isomerase. PLoS One 7, e39550. http://dx.doi.org/10.1371/journal.pone.0039550
- [3]Verdoes, J.C., Krubasik, P., Sandmann, G., Van Ooyen, A.J.J., 1999. Isolation and functional characterisation of a novel type of carotenoid biosynthetic gene from Xanthophyllomyces dendrorhous. Mol. Gen. Genet. MGG 262, 453–461.
- [4]Niklitschek, M., Alcaíno, J., Barahona, S., Sepúlveda, D., Lozano, C., Carmona, M., et al., 2008. Genomic organization of the structural genes controlling the astaxanthin biosynthesis pathway of Xanthophyllomyces dendrorhous. Biol. Res. 41, 93–108. http://dx.doi.org/10.4067/S0716-97602008000100011.
- [5]Xie, W., Lv, X., Ye, L., Zhou, P., Yu, H., 2015a. Construction of lycopene-overproducing
Saccharomyces cerevisiae by combining directed evolution and metabolic engineering. Metab. Eng. 30, 69–78. http://dx.doi.org/10.1016/j.ymben.2015.04.009.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Functional Parameters: Austin_UTexas
Burden Imposed by this Part:

Burden is the percent reduction in the growth rate of E. coli cells transformed with a plasmid containing this BioBrick (± values are 95% confidence limits). This BioBrick did not exhibit a burden that was significantly greater than zero (i.e., it appears to have little to no impact on growth). Therefore, users can depend on this part to remain stable for many bacterial cell divisions and in large culture volumes. Refer to any one of the BBa_K3174002 - BBa_K3174007 pages for more information on the methods, an explanation of the sources of burden, and other conclusions from a large-scale measurement project conducted by the 2019 Austin_UTexas team.
This functional parameter was added by the 2020 Austin_UTexas team.
| None |

