Part:BBa_K895005
Salty_VgrG-CD27_endolysin
This is a coding part comprising the full open reading frame of Salmonella enterica LT2 vgrG gene (also known as STM0289 or tssI) linked via an haemagglutinin eptitope tag to a synthetic gene encoding an endolysin from the CD27 bacteriophage. The CD27 endolysin is believed to have high specificity against the cell wall of Clostridium difficile.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 474
Illegal BglII site found at 1806 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 1048
Illegal AgeI site found at 1195 - 1000COMPATIBLE WITH RFC[1000]
Part Improvement by Oxford 2019
A fundamental crux of the ProQuorum system is the expression of the endolysin which is responsible for the actual lysis and consequent killing of the pathogenic C. difficile. Thus, there is a clear need for an endolysin that fulfills the criteria of being:
1. optimally expressed
2. optimally secreted and
3. optimally effective against C.difficile
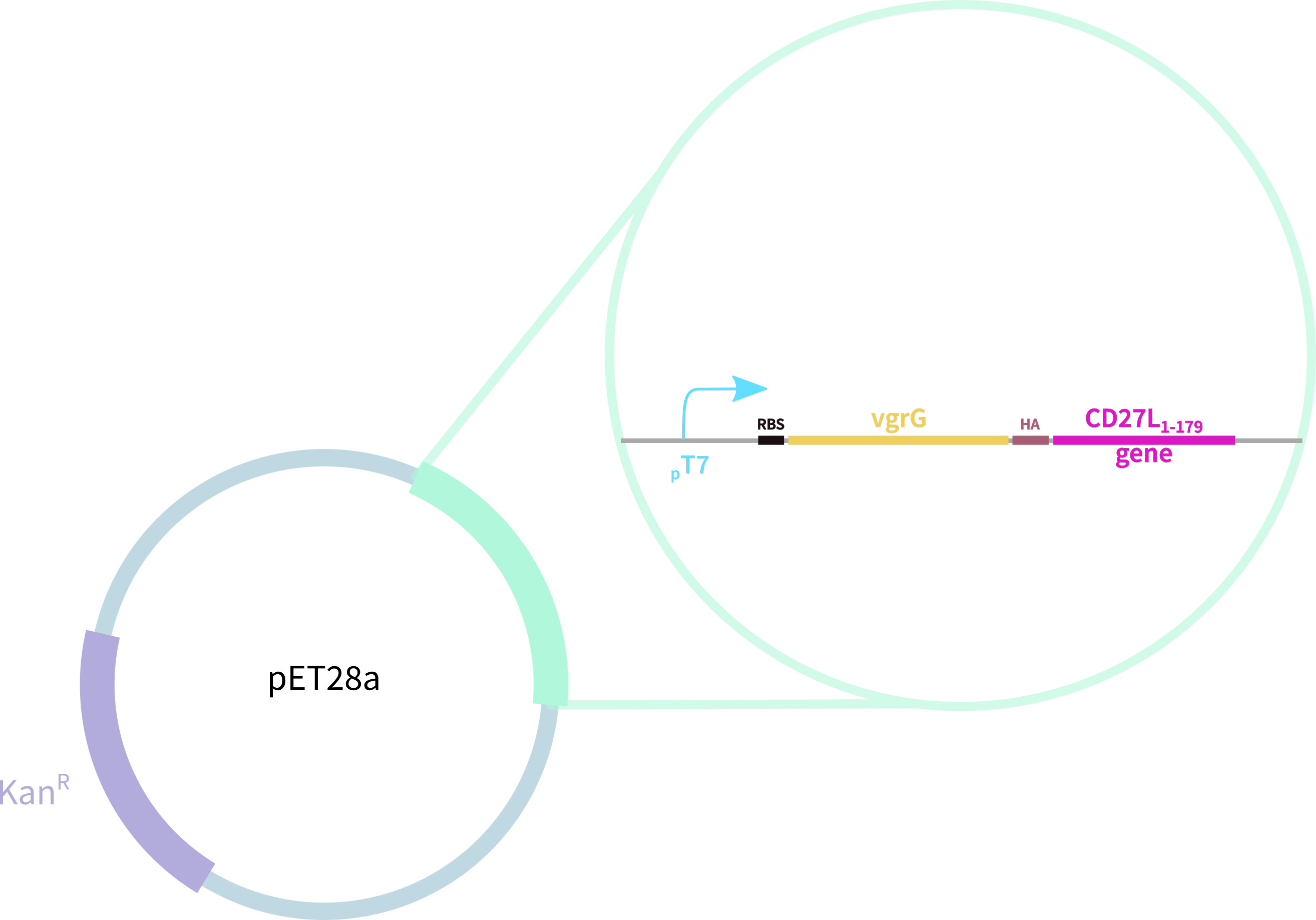
Thus, we redesigned the BBa_K895005 part submitted by the 2012 Dundee iGEM team, which consists of:
1. truncated form of the CD27L endolysin, shown to increase both efficacy and host range2
2. Type VI secretion protein derived from S. typhimurium
3. Hemagglutinin (HA) Tag for Western Blotting</b>
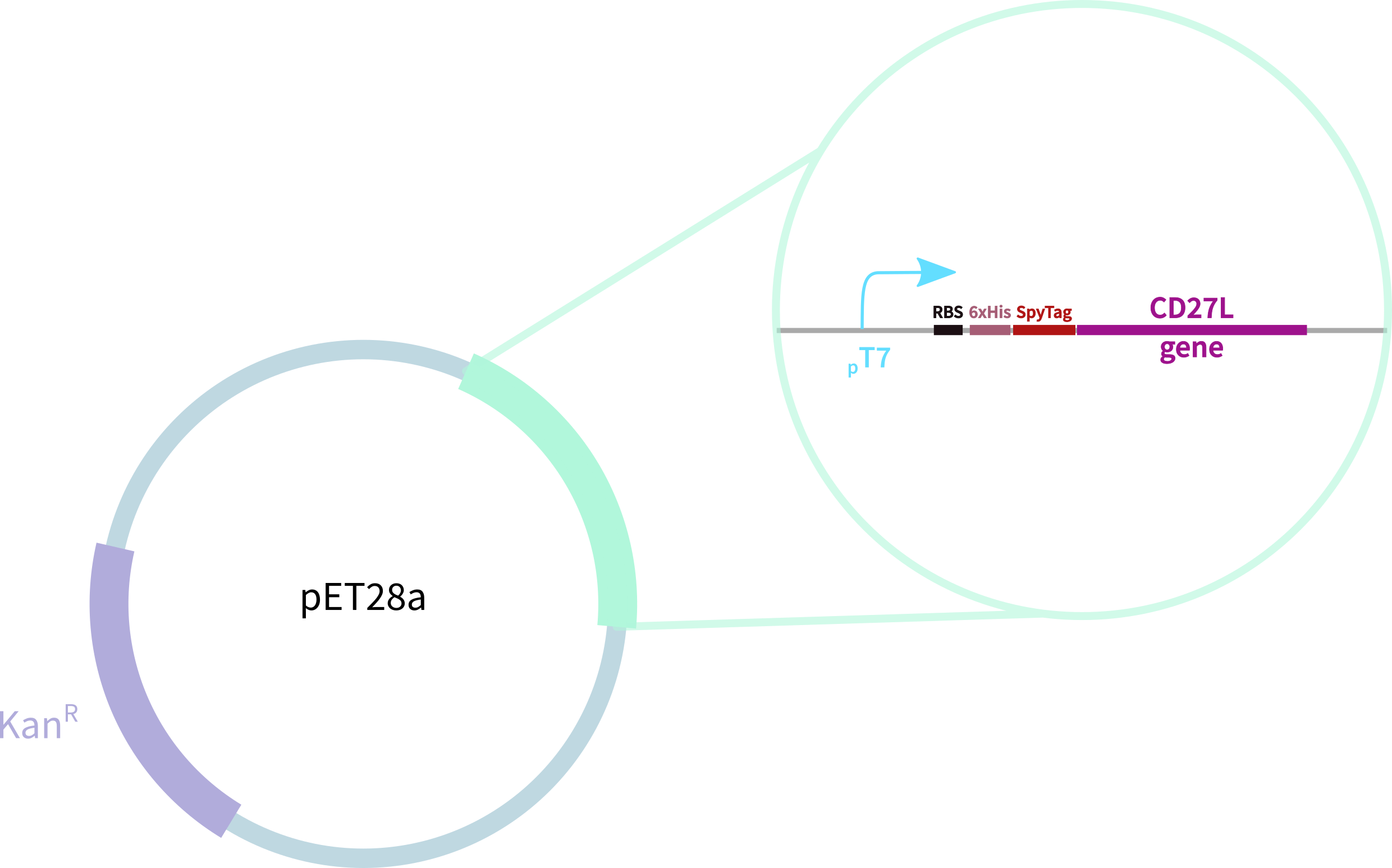
In contrast, our novel part BBa_K3183200 (CD27L) encodes:
1. full-length form of the CD27L endolysin
2. SpyTag for purification, oligomerisation, and other SpyCatcher applications
3. 6-His tag for easy purification
Incorporating both of these parts into pET28A, our expression vector, we planned on testing the relative efficacies of B.subtilis killing as per our killing assays. Thus, we followed our generic pipeline of:
1. transformation of both constructs into E.coli (BL21 (DE3)-RIPL Competent E.coli)
2. miniprep + sequencing to verify successful transformation
3. induction of expression with IPTG
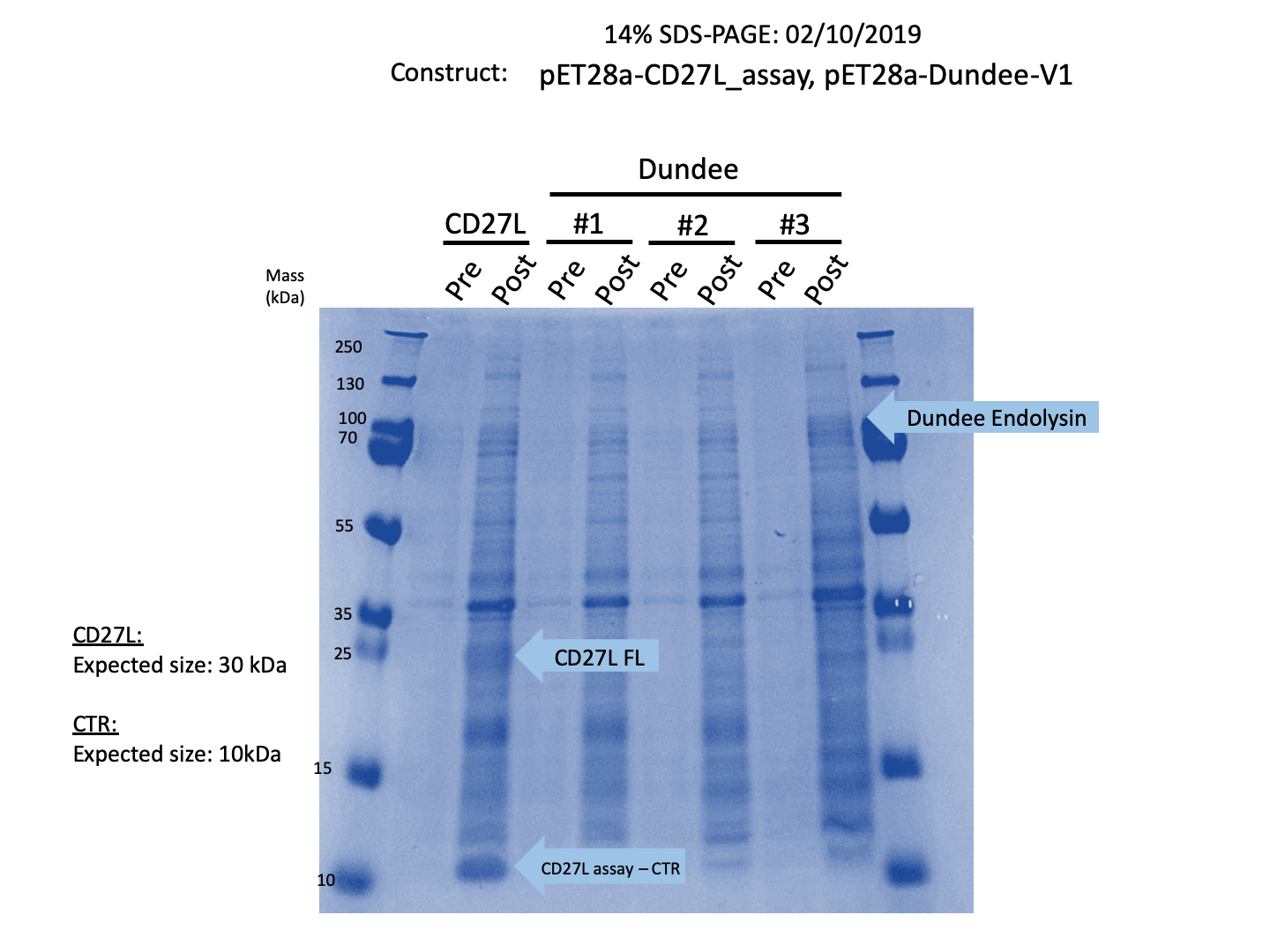
Thus, we sought out the endolysin in an existing part made by the 2012 iGEM Team Dundee. However, despite identical growth and induction conditions, expression of the Dundee 2012 iGEM endolysin was unsuccessful, even in triplicate, as shown in Figure 2. In contrast, expression of the CD27L_Assay part was successful and quantitatively verified via both mass spectrometry and BCA assay, as in the Results page.
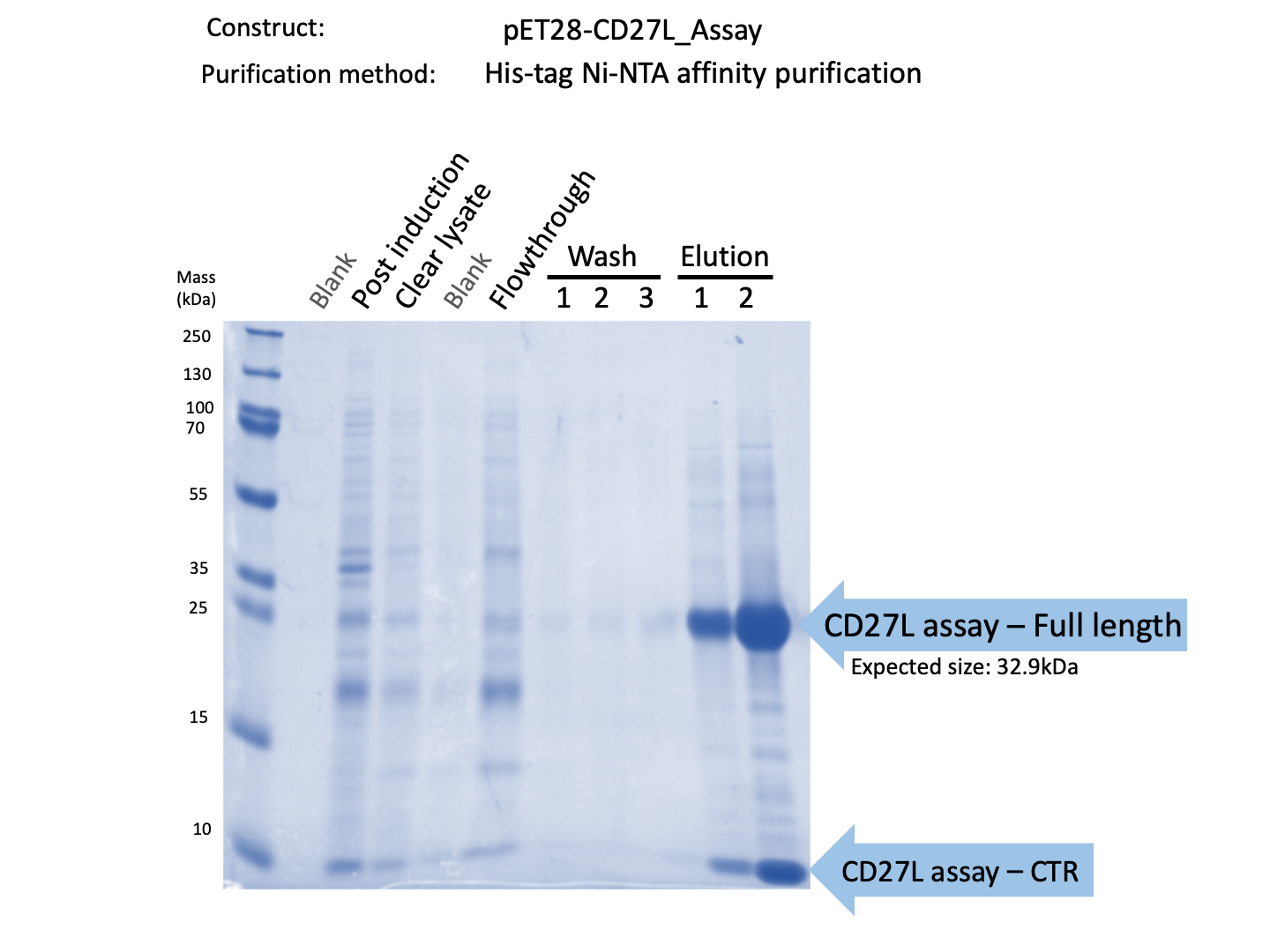
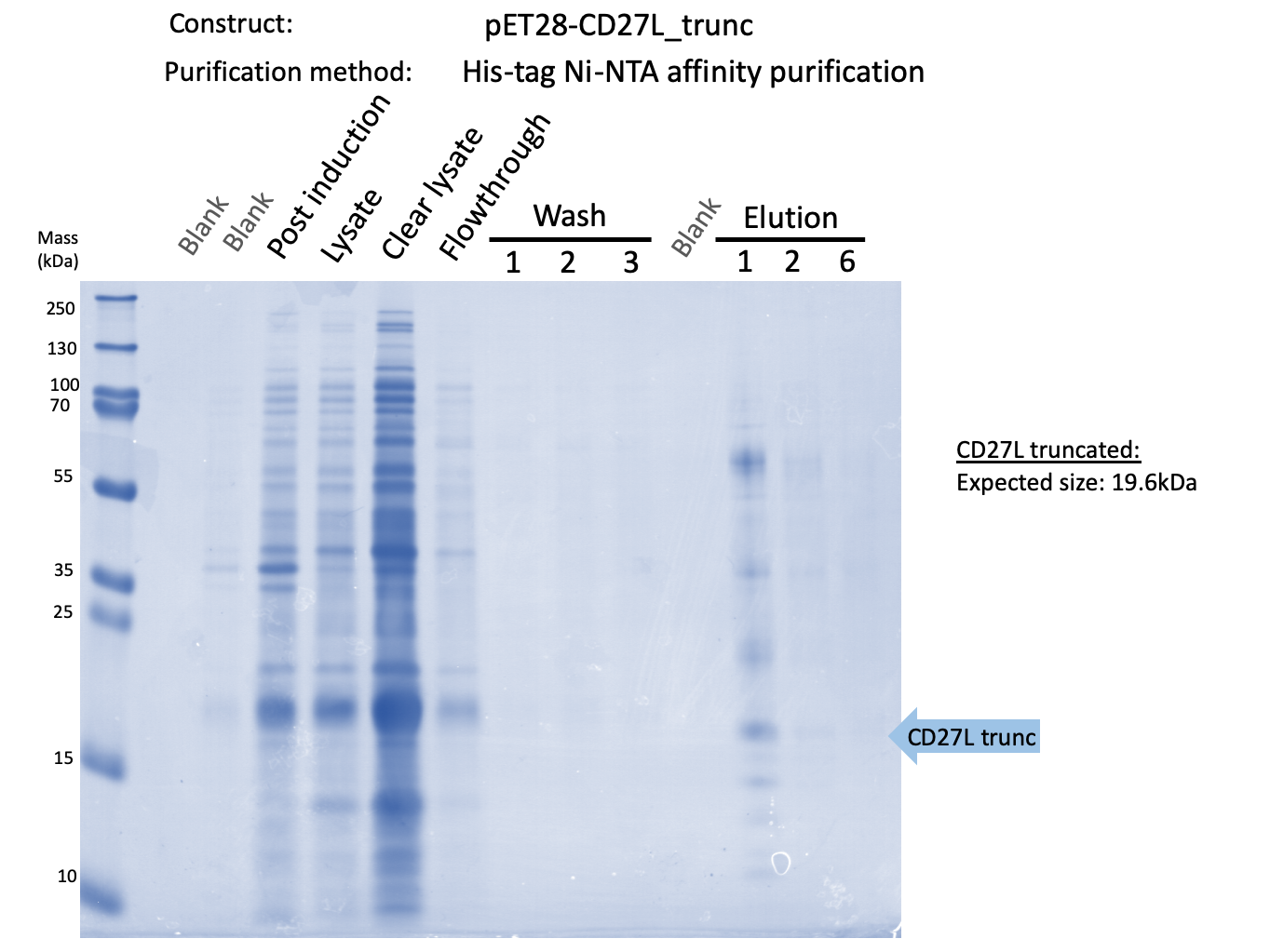
Thus, to solve this issue, we decided to attempt expression of only the truncated endolysin from the Dundee 2012 Biobrick, without its substantially large Type VI secretion tag. We added 6His and SpyTag to the N-terminus to allow for detection (via SpyCatcher gel-shift assays) and purification. This resulted in BBa_K31830201 (CD27L1-179). Upon analysis, there is evident expression of the truncated endolysin as seen in Figure 3.1. However, given identical conditions of growth and induction, there appears to be greater expression of the full-length endolysin (Fig. 3.2), which may perhaps indicate greater stability.
As a further improvement, we decided to measure the killing efficacy of the CD27L_Assay endolysin on B. subtilis (our surrogate target) as no killing data was provided for the Dundee 2012 BBa_K895005 part. As seen in Figure 4, there is a substantial decrease in OD600 over time relative to the negative controls.
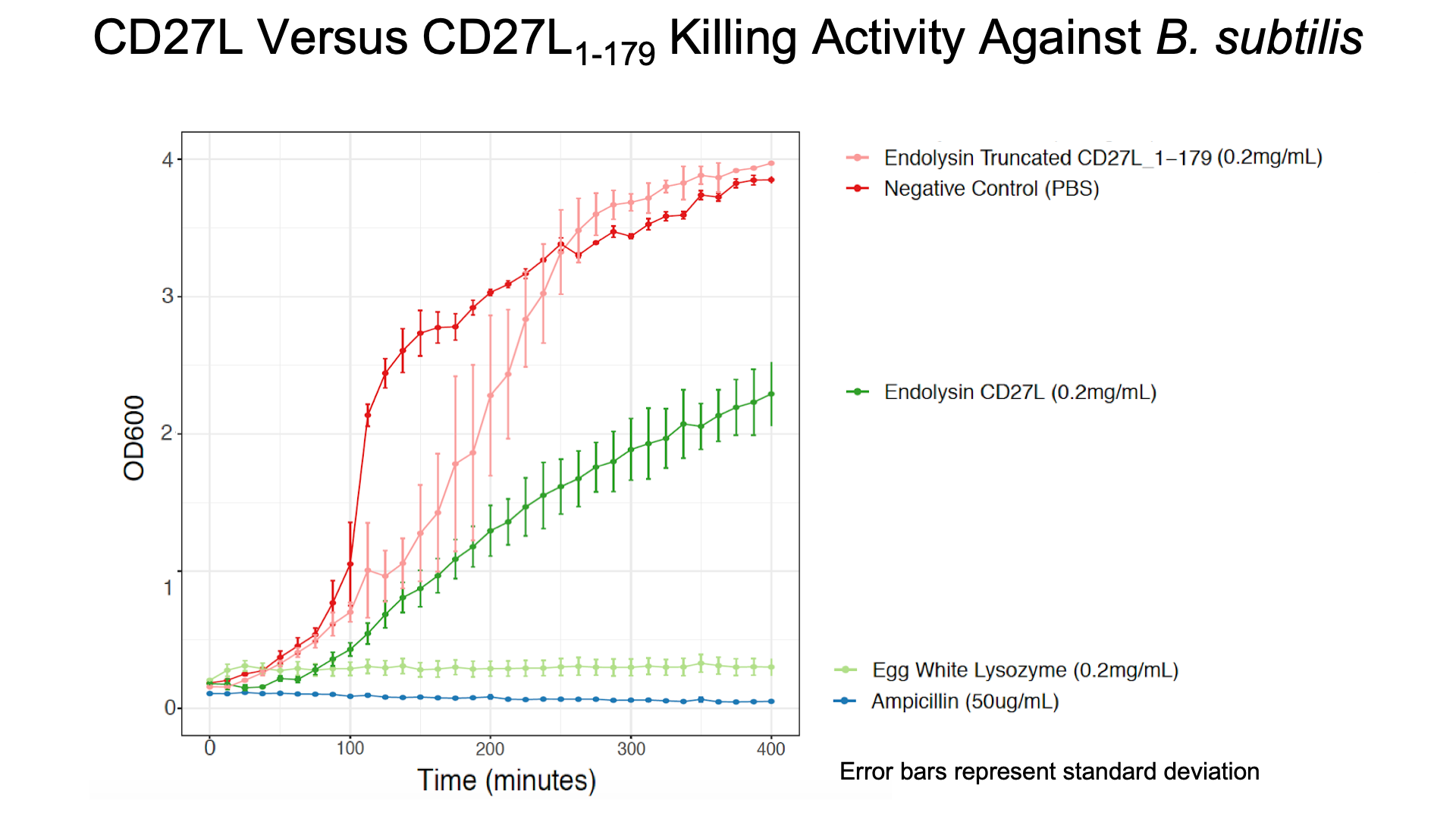
The truncated CD27L1-179 shows decreased growth relative to the negative control; however, log-phase growth resumes after 150 minutes. CD27L endolysin results in decreased growth during the first 100 minutes, and growth thereafter appears to be linear. This may point to inhibited log-phase growth in subsequent generations of B. subtilis following CD27L exposure.
References
1. Mayer, M. J., et al. “Molecular Characterization of a Clostridium Difficile Bacteriophage and Its Cloned Biologically Active Endolysin.” Journal of Bacteriology, vol. 190, no. 20, 2008, pp. 6734–6740., doi:10.1128/jb.00686-08.
2. Mayer, M. J et al. “Structure-based modification of a Clostridium difficile-targeting endolysin affects activity and host range.” Journal of bacteriology vol. 193,19 (2011): 5477-86. doi:10.1128/JB.00439-11
3. Dunne, Matthew et al. “The CD27L and CTP1L endolysins targeting Clostridia contain a built-in trigger and release factor.” PLoS pathogens vol. 10,7 e1004228. 24 Jul. 2014, doi:10.1371/journal.ppat.1004228
4. Twetman, Svante, et al. “Scanning Electron Microscopic Study of Streptococcus Mutans BHT Lysed by Lysozyme.” European Journal of Oral Sciences, vol. 93, no. 1, 1985, pp. 23–29., doi:10.1111/j.1600-0722.1985.tb01304.x.
| None |