Part:BBa_K530000
CRTYB
Toulouse_INSA-UPS 2020contributed to the characterisation of this part by adding a new documentation learned form literature on the expression and stability of CrtI.
(--antonmykhailiuk 19:10, 08 October 2020 (UTC+2))
Enzyme in the pathway required for B-Carotene Synthesis. This enzyme is a combination, it is the full Phytoene Synthase enzyme spliced with a Lycopene B-Cyclase enzymatic domain. This sequence was taken from a WT strain of xanthophyllomyces dendrorhous. It catalyzes both the conversion of Geranylgeranyl diphosphate to Phytoene and Lycopene to B-Carotene.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 1068
Illegal BsaI.rc site found at 1399
Plasmid Map
Characterization of performance
Below we can see the beta-carotene production levels on a per cell basis for different combinations of parts involved in beta-carotene synthesis. These are compared with a wild type control. tHMG1 is used to funnel more initial substrate into the pathway and although useful, was not submitted as a part. 2I indicates that two copies of crtI were used.
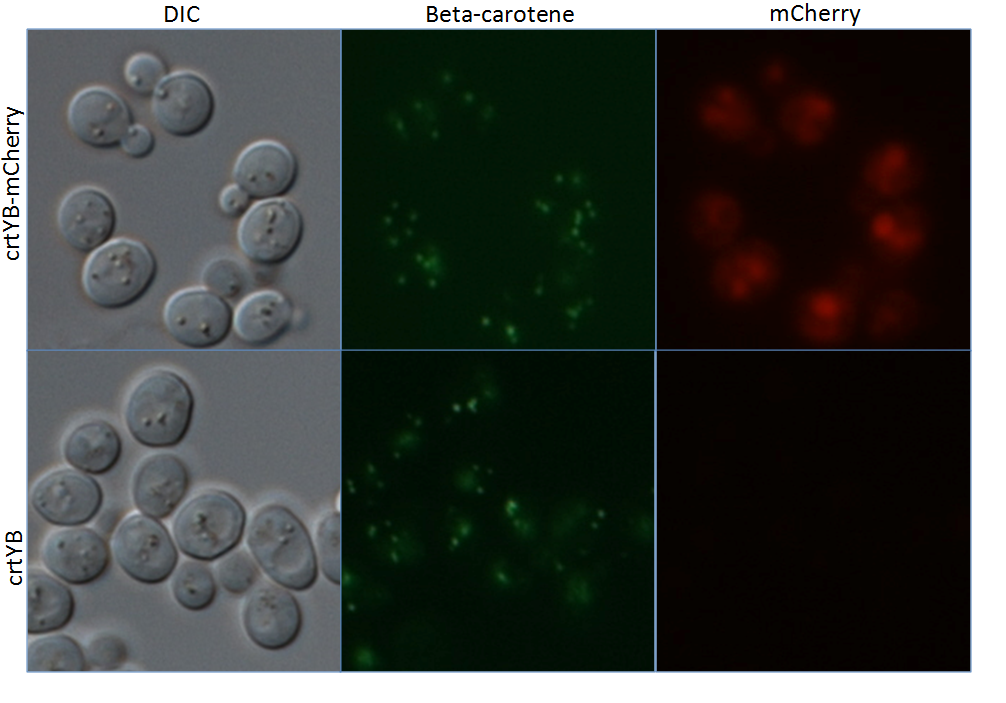
Localization of crtYB-mCherry and beta-carotene in yeast
Yeast cells, previously engineered to express all genes in the beta carotene biosynthetic pathway were subsequently transformed with a copy of crtYB C-terminally tagged with mCherry. The subcellular localization pattern of mCherry (crtYB-mCherry) was then assessed by fluorescence microscopy as compared to the parental, untagged control strain (crtYB) under identical conditions. Autofluorescence of beta carotene using a filter set for fluorescein fluorescence was also performed. Yeast cells are shown using differential interference contrast (DIC). Microscopy was performed at 1000X magnification.
Sequencing
This is the sequencing for colony 2.
Improvement by Team Foshan-GreatBay
Group: Foshan-GreatBay iGEM 2024
New Improved Parts: BBa_K5419002 (pX-2-XdCrtE), BBa_K5419007 (pXII-5-XdCrtI), and BBa_K5419009 (pXI-2-XdCrtYB)
Existing Parts: BBa_K2407309 (CrtI), BBa_K530000 (CrtYB)
Summary
To construct Saccharomyces cerevisiae (S. cerevisiae) strain with high β-carotene production, we added new composite parts (BBa_K5419002, BBa_K5419007, and BBa_K5419009). At the same time, experimental data were also added to the existing parts (BBa_K2407309 (CrtI), BBa_K530000 (CrtYB)) that included:
- Construction of the integration plasmids coding XdCrtE, XdCrtI, and XdCrtYB, respectively.
- Integration of the genes by CRISPR/Cas9 technology into S. cerevisiae.
- Analysis of the expression levels of the genes and the testing of the combination of these genes for the production of β-carotene.
Documentation
a. Usage and Biology
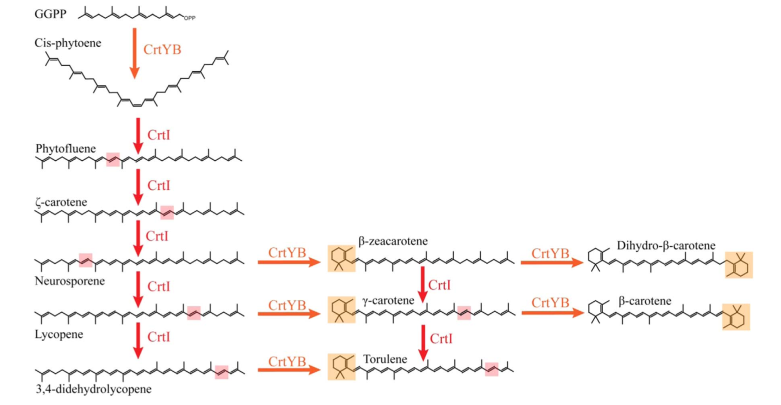
In the S. cerevisiae, CrtE gene encodes GGPP synthase, in the presence of which FPP forms GGPP. The two GGPP molecules then form octahydrodicarbons via the CrtB gene-encoded octahydrodicarbon synthase. Then, octahydro lycopene dehydrogenase encoded by CrtI gene converts octahydro lycopene to lycopene. Finally, the CrtY-encoded lycopene β-cyclase will catalyze lycopene and eventually form β-carotene [1]. CrtYB gene, on the other hand, encodes a bifunctional gene that functions as both CrtB and CrtY (Figure 1). For the species origin of the genes, we chose CrtE, CrtI, and CrtYB genes from Xanthophyllomyces dendrorhous (X. dendrorhous), which are more suitable for expression in S. cerevisiae [2].

b. Characterization/Measurement
(1) Construction Design
We constructed the plasmids by placing the genes under the regulation of a strong constitutive GAP promoter and a CYC terminator, respectively. Integration sequences were added upstream and downstream of the expression cassettes to integrate the target genes into the genome of S. cerevisiae using the CRISPR/Cas9 system (Figure 2).

(2) Construction of integration plasmids
Firstly, we obtained the target gene expression frames (GAP promotor-gene-CYC terminator) by PCR amplification, and agarose gel electrophoresis results showed that we succeeded in obtaining these fragments. Next, we double-digested the target fragment and the vector (containing the S. cerevisiae X-2, XI-2, and XII-5 integration site genes, respectively) and obtained the plasmid by enzymatic ligation. Finally, we transformed the enzyme-ligation product into E. coli DH5α competent cells, and the colony PCR and sequencing results showed that we successfully constructed three integration plasmids (Figure 3).

(3) Integration of target genes into the yeast genome
We reserved NotI digestion sites upstream and downstream of the integration fragment, respectively. After successfully obtaining the integration plasmids, we used NotI restriction endonuclease to obtain the complete destination fragment (containing the sequence upstream of the integration site, the GAP promoter, the target gene, the CYC terminator, and the sequence downstream of the integration site). Subsequently, we recovered these integration fragments in combination with the corresponding gRNA plasmids (X-2-XII-5-gRNA-HYG and XI-2-gRNA-HYG) and introduced them into the modified S. cerevisiae 1974 strain (which had pre-integrated the Cas9 gene) by a modified lithium acetate transformation method. After two rounds of integration, we used yeast colony PCR to verify that the target fragments were successfully integrated into the yeast genome (Figure 4).

(4) Measurement: Quantitative analysis
We used the quantitative PCR (qPCR) technique to determine the transcript levels of target genes integrated into the yeast genome. The ACT1 gene was selected as an internal reference gene and the 2-ΔΔCt method was used to calculate the relative expression of the target gene. The primer amplification efficiency standard curves showed that this qPCR amplification has great reproducibility and accuracy (Figure 5). Subsequently, we analyzed the expression of the target genes in the recombinant yeast strains. The results showed that the expression of the target gene was increased 0.843~1.796-fold in this group compared to the control. These results confirmed that the target genes had been successfully integrated into the yeast genome and could be efficiently transcribed (Table 1).


Finally, β-carotene production quantification
We quantified the β-carotene production of the three recombinant yeast strains. A standard curve was plotted using a series of β-carotene standards with concentration gradients, and a linear regression equation was established to calculate β-carotene concentration. After extraction and analysis, the recombinant yeast strains produced β-carotene at a concentration of 31.47 mg/L (Figure 6).

c. Learn
Based on our experimental results and literature research, it is advisable to refer to the following two recommendations when producing β-carotene in S. cerevisiae strains:
- Adaptation of gene origin: Genes from X. dendrorhous may be more suitable for expression in S. cerevisiae. This host specificity may result from differences in codon usage preferences, post-translational modification mechanisms, or protein folding environments [2].
- Functional properties of the enzyme: XdCrtBY is a bifunctional enzyme that may be spatially more favorable for successive catalytic reactions. This bifunctional property could allow the product of the first reaction to bind faster to the second catalytic site, thus accelerating the whole reaction process and increasing the efficiency of carotenoid production [3].
References
[1] WANG Rui-zhao, PAN Cai-hui, WANG Ying, XIAO Wen-hai, YUAN Ying-jin. Design and Construction of high β-carotene Producing Saccharomyces cerevisiae[J]. China Biotechnology, 2016, 36(7): 83-91.
[2] Puigbo P, Bravo IG, Garcia-Vallve S. CAIcal: a combined set of tools to assess codon usage adaptation. Biol Direct, 2008, 3: 38.
[3] Tokuhiro K, Muramatsu M, Ohto C, et al. Overproduction of geranylgeraniol by metabolically engineered Saccharomyces cerevisiae. Appl Environ Microbiol, 2009, 75(17): 5536–5543.
html
Contribution from other teams
Toulouse_INSA-UPS 2020's contribution
Characterisation
Since the CrtI (phytoene desaturase) is a part of the biosynthesis pathway of carotenoids (fig. 1), it is often co-expressed with the other enzymes of the pathway: such as CrtB or CrtY. Rabeharindranto et al. analyzed the expression of single domain CrtI, CrtY, and CrtB (fig.2). CrtB is unambiguously detected as an intense bands in all strains B, B/I or Y/B/I. On the other hand, CrtI can be observed at the expected size (67kDa) only when coexpressed with CrtY domain (strain Y/B/I). The explanation could be that CrtI protein needs to be co-expressed together with CrtY to have a normal production/stability. A faint band with 50kDa migration pattern could be observed in B/I strain which could further support the idea of low production or stability of CrtI in absence of CrtY.
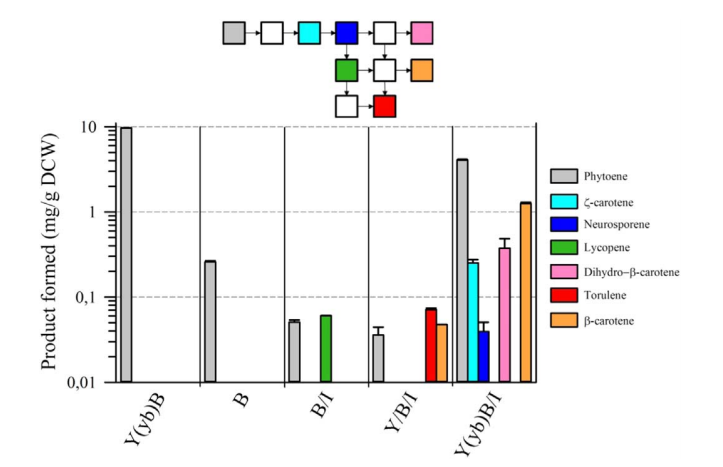
The second point concerns the presence of a natural fusion of CrtY (lycopene cyclase) and CrtB(phytoene synthase). Although, in eucaryotes, CrtB enzyme is predicted to be cytosolic and CrtY enzyme is predicted to be transmembrane [2], surprisingly, there is a natural fusion between CrtY and CrtB which gives CrtYB enzyme[3]. There is a strong opinion on the importance of this natural fusion on the activity of phytoene synthase [4,5]. Rabeharindranto et al. confirmed the impact of CrtY and CrtB splitting on the phytoene production as it has significantly decreased (40 times) in the B strain compared to the Y(yb)B strain (fig. 3).
References for Toulouse_INSA-UPS 2020's contribution
- [1]Rabeharindranto, H., Castaño-Cerezo, S., Lautier, T., Garcia-Alles, L. F., Treitz, C., Tholey, A., & Truan, G. (2019). Enzyme-fusion strategies for redirecting and improving carotenoid synthesis in S. cerevisiae. Metabolic Engineering Communications, 8, e00086. https://doi.org/10.1016/j.mec.2019.e00086
- [2]Schaub, P., Yu, Q., Gemmecker, S., Poussin-Courmontagne, P., Mailliot, J., McEwen, A.G., et al., 2012. On the structure and function of the phytoene desaturase CRTI from Pantoea ananatis, a membrane-peripheral and FAD-dependent oxidase/isomerase. PLoS One 7, e39550. http://dx.doi.org/10.1371/journal.pone.0039550
- [3]Verdoes, J.C., Krubasik, P., Sandmann, G., Van Ooyen, A.J.J., 1999. Isolation and functional characterisation of a novel type of carotenoid biosynthetic gene from Xanthophyllomyces dendrorhous. Mol. Gen. Genet. MGG 262, 453–461.
- [4]Niklitschek, M., Alcaíno, J., Barahona, S., Sepúlveda, D., Lozano, C., Carmona, M., et al., 2008. Genomic organization of the structural genes controlling the astaxanthin biosynthesis pathway of Xanthophyllomyces dendrorhous. Biol. Res. 41, 93–108. http://dx.doi.org/10.4067/S0716-97602008000100011.
- [5]Xie, W., Lv, X., Ye, L., Zhou, P., Yu, H., 2015a. Construction of lycopene-overproducing
| None |