Part:BBa_K4096002
PKC-OP
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal XhoI site found at 1081
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Contents
- 1 Contribution by 2022 iGEM Team Shanghai_United
- 1.1 BBa_K4277006
- 1.2 Construct design
- 1.3 Experimental approach
- 1.4 Functional assay
- 1.5 Improvement of existing data
- 1.6 Construct design
- 1.7 Experimental approach
- 1.8 Proof of function
- 1.9 References
- 1.9.1 MUCK R,NADEAU E,MCALLISTER T,et al. Silage review: recent advances and future uses of silage additives [J]. J Dairy Sci,
- 1.9.2 刘海燕,张鹏举,王秀飞,等. 正交试验法优化玉米秸秆穰叶青贮发酵剂的研究 [J]. 中国饲料,2018 ( 11) : 74-79.
- 1.9.3 Coward-Kelly G., Aiello-Mazzari C., Kim S., Granda C., and Holtzapple M., 2003, Suggested improvements to the standard filter paper assay used to measure cellulase activity,Biotechnology & Bioengineering, 82(6): 745-749.
- 1.9.4 Ghose T.K., 1987, Measurement of cellulase activities, Pure & Appl Chem, 59(2): 257-268.
Contribution by 2022 iGEM Team Shanghai_United
BBa_K4277006
Name: pET28a-PKC
Base Pairs: 5515 bp
Origin: Pseudomonas aeruginosa, genome
Properties: Providing the method for feed degradation
Usage and Biology
The alkaline cellulase is from Pseudomonas aeruginosa (PKC), and used to degrade the insoluble cellulose to soluble glucose. Cellulase with high alkaline stability, active under low temperature is used to as additive.
Construct design
The alkali-cellulase process the biomass into glucose. The gene of alkali-cellulase referred as PKC was integrated into pET28a vector, shown as Figure 1.
Experimental approach
1.1 The colony PCR of pET28a-PKC in competent cells DH5α
PKC gene amplified by PCR, double-enzyme digested by NheI and HindIII, and inserted into the same cohesive site of pET28a (+) vector, to obtain the plasmid pET28a-PKC. Then, the pET28a-PKC transformed into DH5α. Colony PCR results showed that the correct band of PKC, and confirmed that the recombinant plasmid of pET28a-PKC is successful (Figure 2).
1.2 Sequencing result of plasmid pET28a-PKC
The recombinant plasmid further confirmed by sequencing. The sequence alignment result exhibited the no mutation or mismatch in the plasmid pET28a-PKC, shown in Figure 3.
1.3 The colony PCR of pET28a-PKC in competent cells BL21
The plasmid pET28a-PKC was extracted from E. coli DH5α, and transformed it into E. coli BL21(DE3). The colony PCR results showed that it was successful (Figure 4).
Functional assay
2.1 Protein expression of PKC
In order to obtain the protein PKC, we transformed the plasmid pET28a-PKC into E. coli BL21(DE3). The colony was inoculated and cultured in the LB medium, and added IPTG to induce protein expression when the OD600 reached 0.3-0.5. After overnight induction and cultivation, the cells were collected and lysed by ultrasonication to release the intracellular proteins. Next, we used nickel resin to purify the interested protein. The molecular weights of PKC were 48.27 KD.
SDS-page showed the protein PKC in lane S, indicating that it was successfully expressed in E. coli BL21 (DE3). The protein PKC was also found in lane P, possibly due to the inactivation of a small number of proteins.
2.2 Determination of PKC enzymatic activity
Determination of reducing the sugar by DNS method: The absorbance OD540 value of the purified enzyme solutions (PKC) was measured after color reaction with DNS. The activity of the enzyme can be converted by the amount of sugar consumed and the working time.
The enzymatic activity of PKC is about 0.26 U/mL according to DNS method (Table 1 and Figure). The results indicated that the protein PKC has active enzyme activity. It also proved that PKC had the function of degrading cellulose.
Improvement of existing data
The old part BBa_K4096002 of the iGEM2021_Shanghai_Metro_HS team attempted PKC protein expression, but the enzymatic activity was not detected. Compared with 2021, in our BBa_K4277006 the PKC was integrated into pET28a rather than pET-25b, and the functional assay contains not only protein expression but also successfully measured enzyme activity. pET28a enables the soluble protein over-expressing under strong bacteriophage T7 promotor. Thus, we chose the vector to express the PKC protein in order to measure the enzymatic activities. In addition, this part provides extra experimental data of PKC and a reference for the future iGEM team.
BBa_K4096002
Name: PKC-OP
Base Pairs: 1086bp
Origin: Pseudomonas aeruginosa, genome
Properties: A coding sequence of alkali cellulase.
Usage and Biology
BBa_K4096002 is a coding sequence of alkali cellulase (PKC-OP) from Pseudomonas aeruginosa. Cellulases break down the cellulose molecule into monosaccharides ("simple sugars") such as beta-glucose, or shorter polysaccharides and oligosaccharides. Several different kinds of cellulases are known, which differ structurally and mechanistically. Synonyms, derivatives, and specific enzymes associated with the name "cellulase" include endo-1,4-beta-D-glucanase (beta-1,4-glucanase, beta-1,4-endoglucan hydrolase, endoglucanase D, 1,4-(1,3,1,4)-beta-D-glucan 4-glucanohydrolase), carboxymethyl cellulase (CMCase), avicelase, celludextrinase, cellulase A, cellulosin AP, alkali cellulase, cellulase A 3, 9.5 cellulase, and pancellase SS.
Construct design
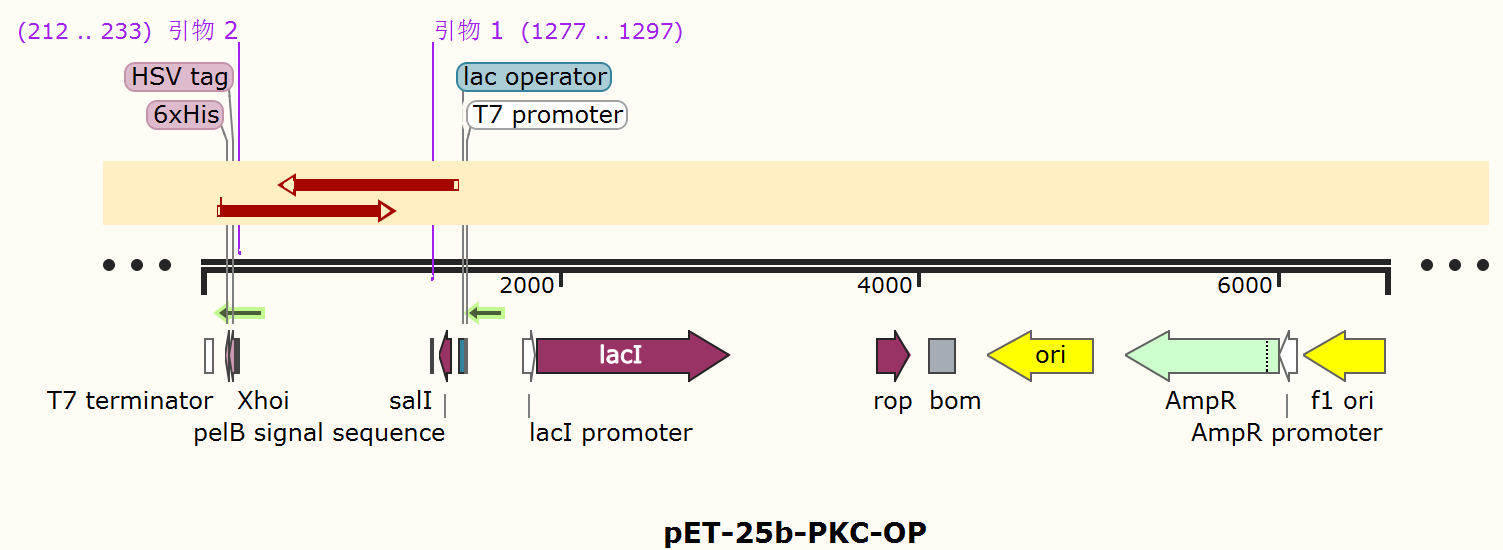
The alkaline cellulase gene from Pseudomonas aeruginosa PKC-001 was selected and we performed codon optimization on PKC-001 sequence to get PKC-OP, which was inserted into the pET-25b vector which contains a pelB signal peptide (Figure 1). The recombinant plasmid then was transformed it to E. coli BL21(DE3) strain to produce cellulase.
The profiles of every basic part are as follows:
Experimental approach
Construction of recombinant plasmid
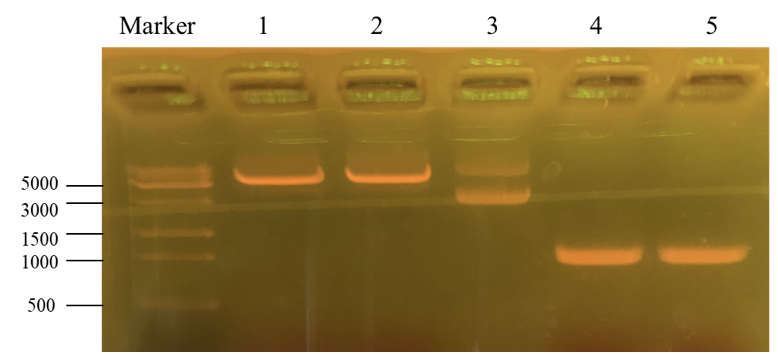
This picture is the nucleic acid electrophoresis result of enzyme cutting and PCR (By performing PCR, we can obtain the desirable PKC-OP’s gene segment.). Column “marker” is a column that is used to show the position of different lengths of genes. In this step, our aim is to verify whether these results are the desired ones. Number 1 and 2 is the result for pET-25b after enzyme digestion. Additionally, number 3 is the pET-25b without enzyme cutting. At last, number 4 and 5 is the PKC-OP after PCR. As our experiment moves on, the concentration of our PKC-OP and pET-25b are 233.2ng/μL and 17.8ng/μL. Lastly, we use the homologous combination to combine them.
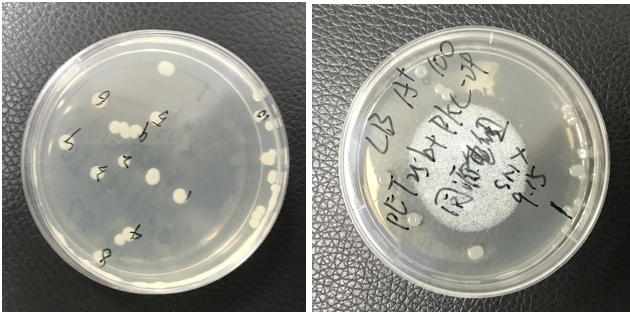
The pET25b-PKC-OP was constructed. The plate shows monoclonals of pET25b-PKC-OP constructs.
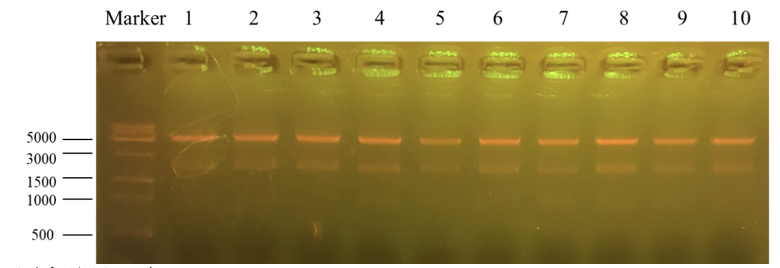
Column “marker” is a column that is used to show the position of different lengths of genes. Number 1 to 10 is the result for recombinant plasmid pET25b-PKC-OP after Apa1 enzyme digestion. We get two bands of 4712bp and 1894bp. It further indicates that the obtained monoclonals were positive monoclonals containing the recombinant plasmid. 1, 7 and 8 plasmids were sent to sequence.
Sequencing feedback shows we have obtained the correct plasmids which is consistent with their DNA profiles.
Proof of function
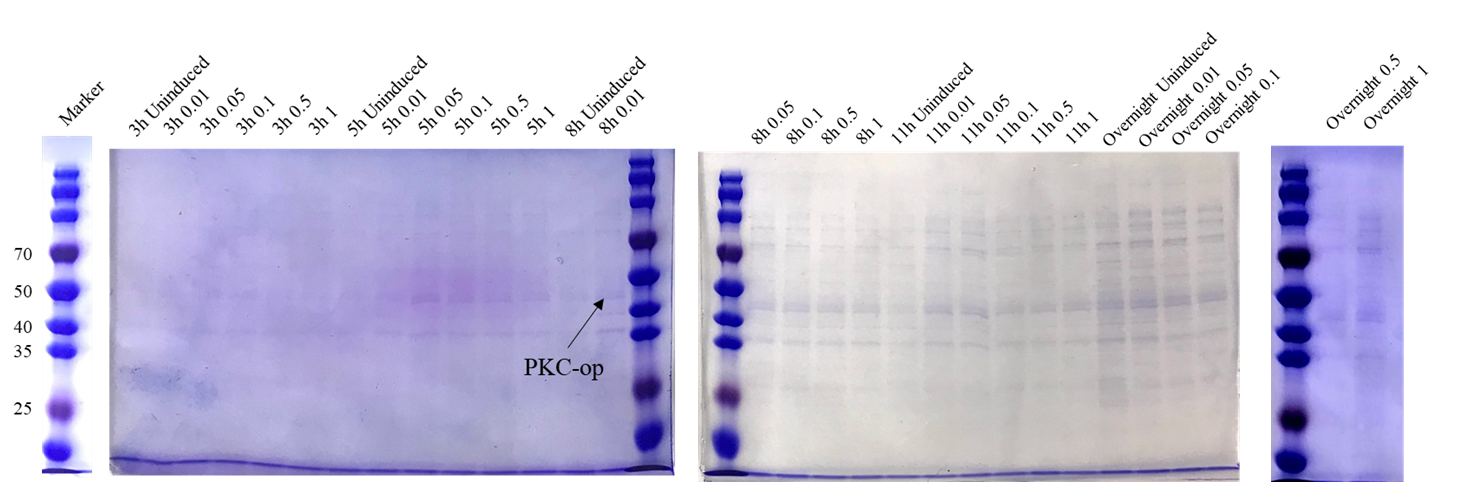
Different induction conditions were tested for protein expression. As the pET-25b vector contains a pelB signal peptide, the engineered strain would secret the protein into the medium. Therefore, we collected the culture supernatant after induction and ran SDS-PAGE for verification (Figure 6).
The theoretical molecular weight of the PKC cellulase is 45.6 kDa. As seen from the SDS-PAGE (Figure 6), there is a wide band just above 40 kDa and it indicates that we have obtained the PKC cellulase.
References
MUCK R,NADEAU E,MCALLISTER T,et al. Silage review: recent advances and future uses of silage additives [J]. J Dairy Sci,
刘海燕,张鹏举,王秀飞,等. 正交试验法优化玉米秸秆穰叶青贮发酵剂的研究 [J]. 中国饲料,2018 ( 11) : 74-79.
Coward-Kelly G., Aiello-Mazzari C., Kim S., Granda C., and Holtzapple M., 2003, Suggested improvements to the standard filter paper assay used to measure cellulase activity,Biotechnology & Bioengineering, 82(6): 745-749.
Ghose T.K., 1987, Measurement of cellulase activities, Pure & Appl Chem, 59(2): 257-268.
| None |