Part:BBa_K3904404
tyrP-specific sgRNA
Introduction
Vilnius-Lithuania iGEM 2021 project AmeByelooks at amebiasis holistically and comprehensively, therefore target E. histolytica from several angles: prevention and diagnostics. Our team's preventive solution consists of probiotics engineered to produce naringenin - an antiprotozoal compound. Two strains of genetically modified microorganisms were chosen as the main chassis - world-renowned Lactobacillus casei BL23 (Lactobacillus paracasei) and Escherichia coli Nissle 1917. Furthermore, the team made specific gene deletions to enhance naringenin production and adapted a novel toxin-antitoxin system to prevent GMO spreads into the environment. The diagnostic part includes a rapid, point of care, user-friendly diagnostic test identifying extraintestinal amebiasis. The main components of this test are aptamers specific to the E. histolytica secreted proteins. These single-stranded DNA sequences fold into tertiary structures for particular fit with target proteins.
Contents
Usage and Biology
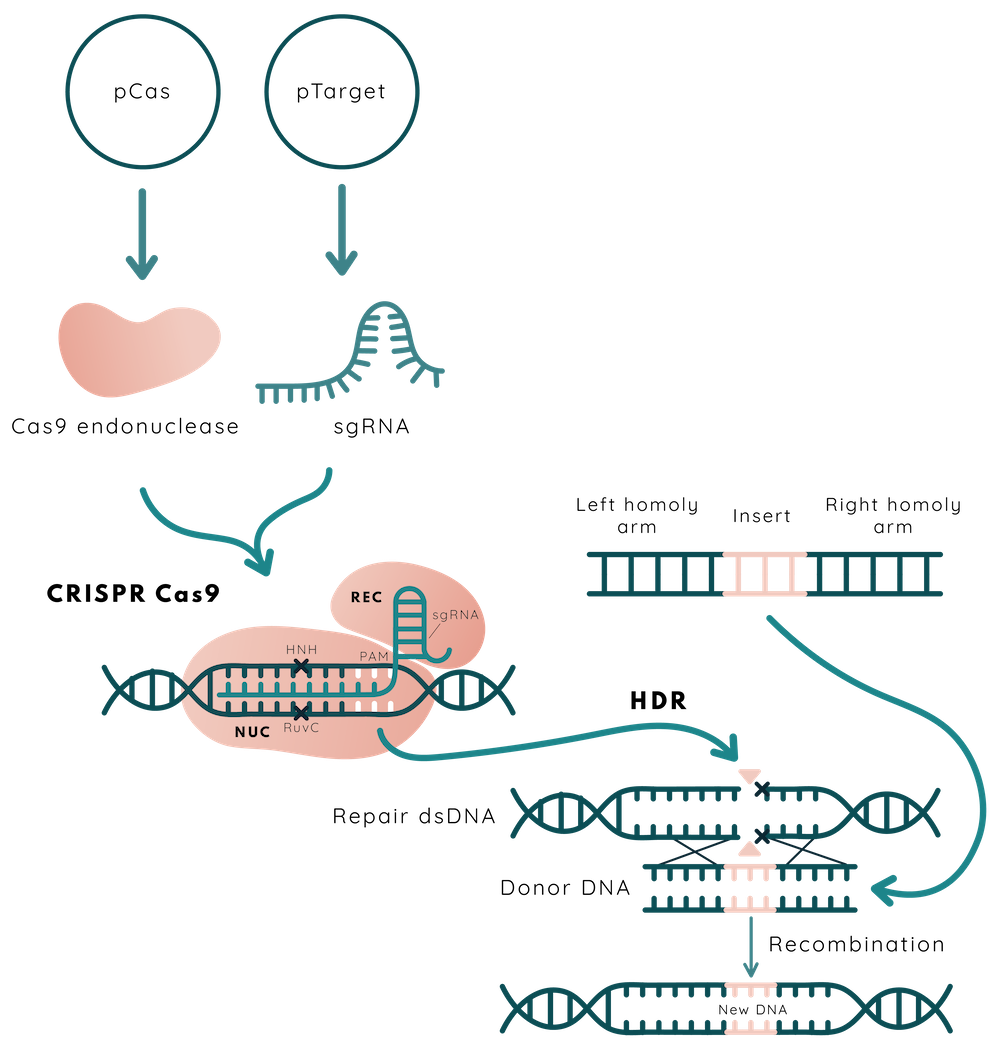
CRISPR-Cas9 is a versatile genome-editing technique. In our approach to editing E. coli Nissle 1917 genome, we have used two plasmid based system enabling to combine of Lambda Red recombination and CRISPR-Cas9 as counterselection tools encoded in pCas and pTarget.
Mechanism of genome editing
pCas plasmid is used for Cas9, Lambda Red system expression, and plasmid curing of pTarget. Cas9 - the RNA-guided endonuclease - is expressed constitutively, while the expression of Lambda Red genes (Gam, Exo, Beta) is under the control of arabinose inducible promoter araBp. pTarget plasmid caries constitutively expressed single-guide RNA (sgRNA). This RNA molecule, as and in nature, is composed of two central parts: CRISPR RNA (crRNA) and tracrRNA. crRNA is 17-20 nt length RNA sequence complementary to the targeted DNA adjacent to the protospacer adjacent motif (PAM) and tracrRNA is the scaffold for the Cas (in this case Cas9) nuclease binding to guide RNA and forming the ribonucleoprotein complex (1). In nature those two parts exist as two separate RNA molecules, however, in laboratory experiments they are usually combined into one single-guide RNA (sgRNA). As both pCas and pTarget plasmids are in a cell, Cas9 nuclease and sgRNA are able to form ribonucleoprotein complex and perform a double-strand break in the chosen part of the DNA. However, if arabinose has been added to the cell culture and a double-stranded DNA repair template is present in the cell, the Lambda Red system performs homologous recombination. If this process is unsuccessful, the Cas9-sgRNA complex will cause a double-strand break and will cause cell death (2). This is employed as a counterselection in order to avoid the additional antibiotic as selection marker usage.
Table 1. sgRNA collection for E. coli Nissle 1917 genome editing.
| ackA-specific sgRNA | BBa_K3904401 |
| pta-specific sgRNA | BBa_K3904402 |
| colicin-specific sgRNA | BBa_K3904405 |
| nupG-specific sgRNA | BBa_K3904426 |
tyrP gene knockout importance to AmeBye project
This 20 nt length RNA sequence is designed to target tyrosine-specific transport protein-coding gene tyrP. tyrP is known to be involved in transporting tyrosine across the cytoplasmic membrane, therefore tyrP knockout mutants are potentially able to produce 10 % higher amounts of L-tyrosine than that of the original strains (3). This L-tyrosine then can be converted to p-coumaric acid by tyrosine ammonia-lyase (TAL), which is further utilized to make p-coumaroyl-CoA by 4-coumarate: CoA ligase. The sequential condensation of one p-coumaroyl-CoA and three malonyl-CoA leads to the formation of naringenin chalcone by chalcone synthase (CHS), which is then converted to naringenin by chalcone isomerase (CHI) (4).
Genome editing efficiency
tyrP knockout generation with this sgRNA have achieved 80 % efficiency (fig. 1).
Fig. 1. Restriction of cPCR product representing tyrP knockout generation. tyrP gene have been amplified from genomic DNA and restricted by BcuI. 1212 bp fragments represent wild type genotype, 1014 bp and 184 bp - knockouts. 1 - wild type (negative control), 2 - tyrP knockout (1), 3 - tyrP knockout (2), 4 - tyrP knockout (3), 5 - tyrP knockout (4), 6 - tyrP knockout (5), 7 - tyrP knockout (6), 8 - tyrP knockout (7), 9 - tyrP knockout (8), 10 - tyrP knockout (9), 11 - tyrP knockout (10).
However, we do not used these mutants in our experiments as they performed unknown genomic reorganization for unknown reasons. Only genome sequencing might help to show real changes.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 9
- 1000COMPATIBLE WITH RFC[1000]
References
- Doudna, J. A., & Charpentier, E. (2014). The new frontier of genome engineering with CRISPR-Cas9. Science, 346(6213).
- Jiang, Y., Chen, B., Duan, C., Sun, B., Yang, J., & Yang, S. (2015). Multigene editing in the Escherichia coli genome via the CRISPR-Cas9 system. Applied and environmental microbiology, 81(7), 2506-2514.
- Wang, Q., Zeng, W., & Zhou, J. (2019). Effect of gene knockout of L-tyrosine transport system on L-tyrosine production in Escherichia coli. Sheng wu gong cheng xue bao= Chinese journal of biotechnology, 35(7), 1247-1255.
- Ganesan, V., Li, Z., Wang, X., & Zhang, H. (2017). Heterologous biosynthesis of natural product naringenin by co-culture engineering. Synthetic and systems biotechnology, 2(3), 236-242.
| None |