Part:BBa_K3815016
AANDENYAKWA. mutant SsrA degradation tag
Usage and Biology
This is an engineered derivative of wildtype ssrA tag from Escherichia coli, where three C-terminal amino acids LAA in WT are replaced with KWA. Refer to BBa_M0050 for the biological function of the tag.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Result
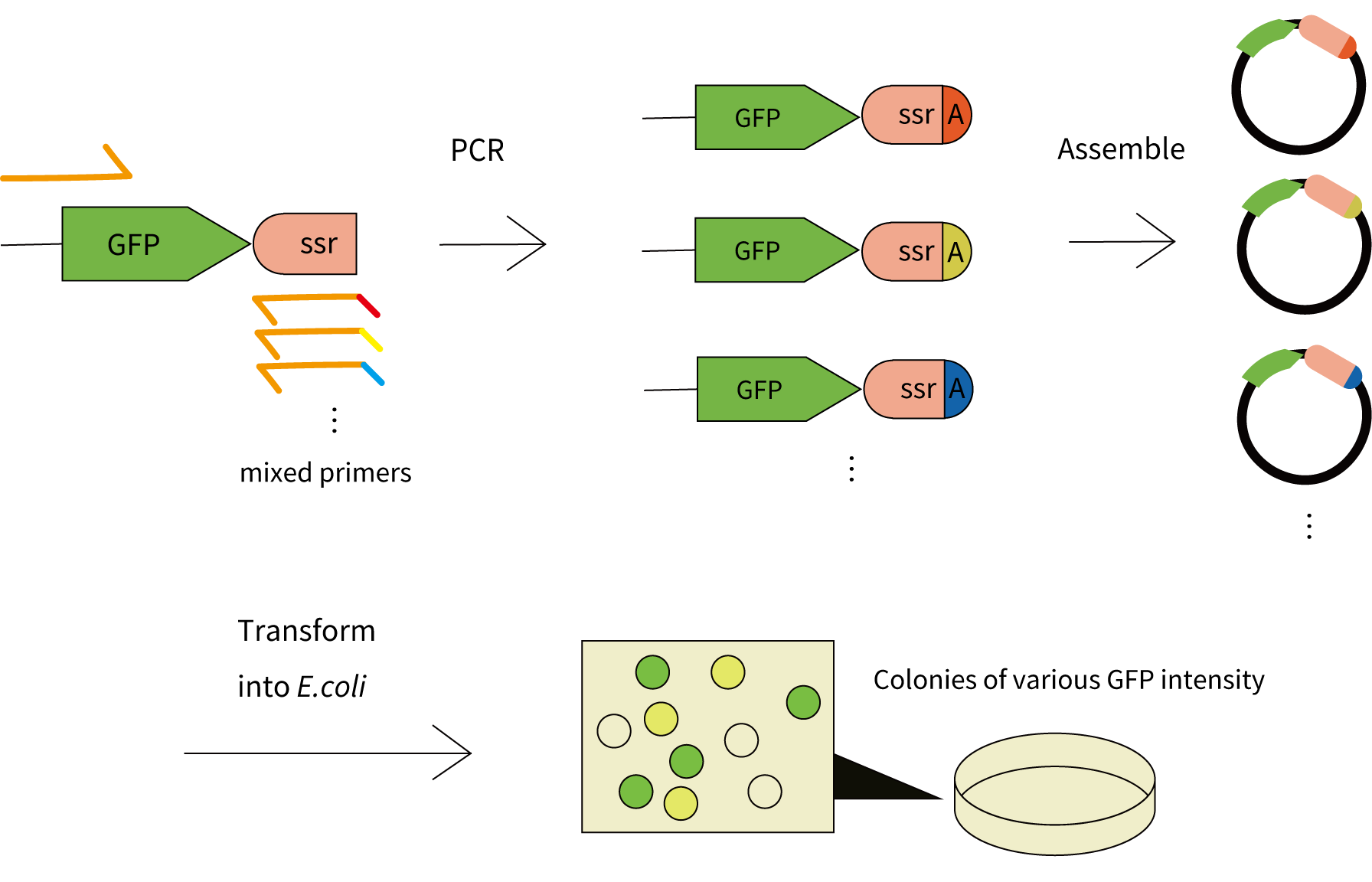
For a flexible control of protein half-life, we aimed to obtain a collection of mutant tags with various degradation efficiencies.
We fused the ssrA tag sequence to GFP while introducing mutations in the tag by random-base primers, and cloned the mutant library of ssrA-tagged GFP into a plasmid vector, so that mutant tags of different activity can be identified by comparing GFP intensity of E.coli transformants.
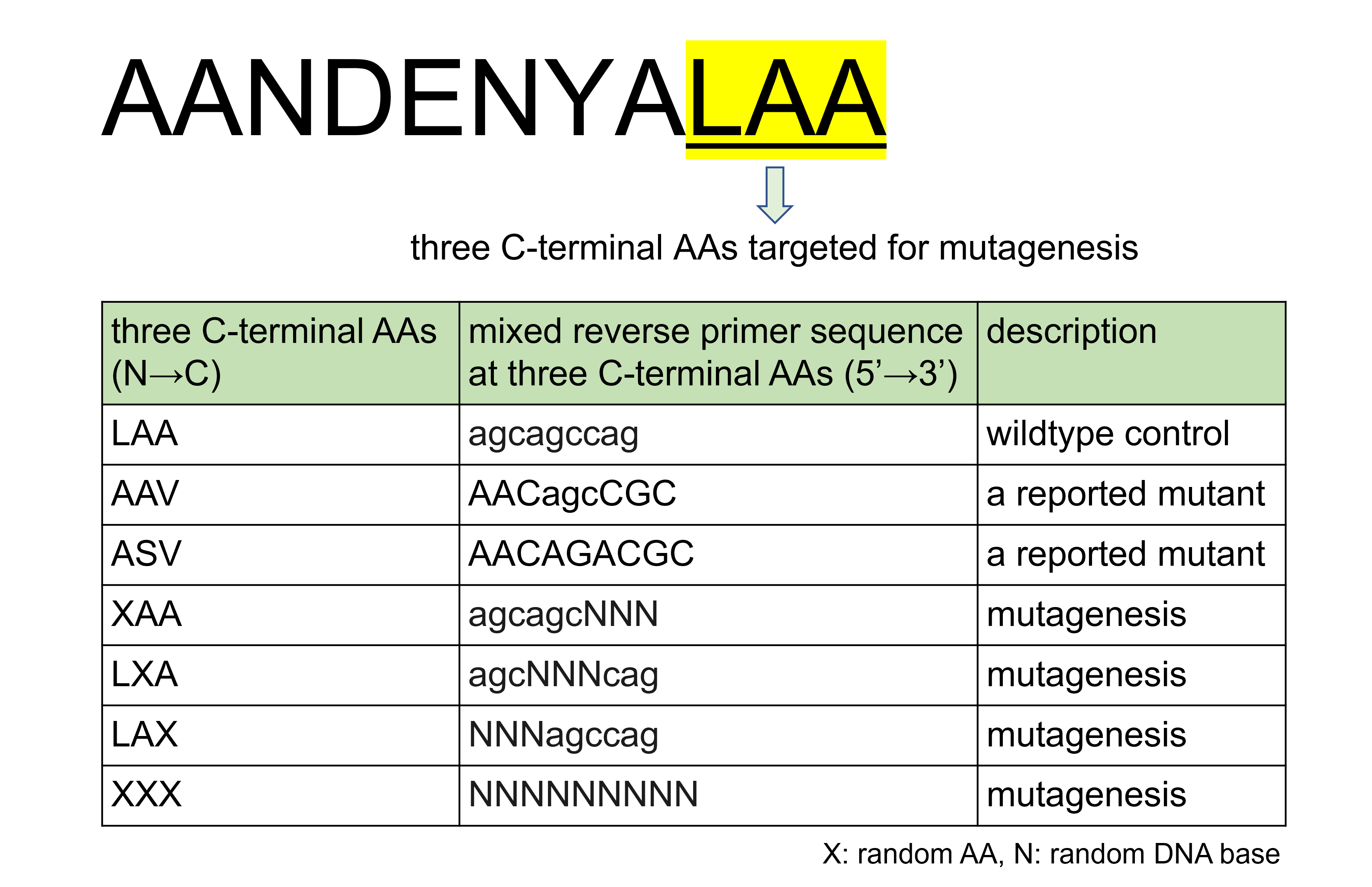
The wildtype ssrA tag amino acid sequence is AANDENYALAA. It was reported that the three amino acids at the C terminus of the tag (LAA in wildtype) have a great impact on the degradation rate of tagged protein [1]. Therefore, we chose these three amino acids as the targets for mutagenesis.
As a result of transformation, colonies of various fluorescence intensity were obtained on the LB plates. 72 colonies in total were picked from plates, and cultured overnight in LB media. The images of E.coli cultures were taken under the blue light.
Plasmid was extracted from each bacterial culture by miniprep, and then Sanger-sequenced to identify ssrA tag sequence. Furthermore, to quantify protein degradation efficiency of each mutant, GFP Fluorescence of each bacterial overnight culture was imaged on Typhoon 9410 (GE healthcare).
The mutant ssrA tag sequence of each clone was sequenced, and biological triplicate of each bacterial overnight culture was imaged for GFP fluorescence, including controls (LB media only, bacterial culture not expressing GFP (no GFP), and bacterial culture expressing non ssrA-tagged GFP (no ssrA tag)).
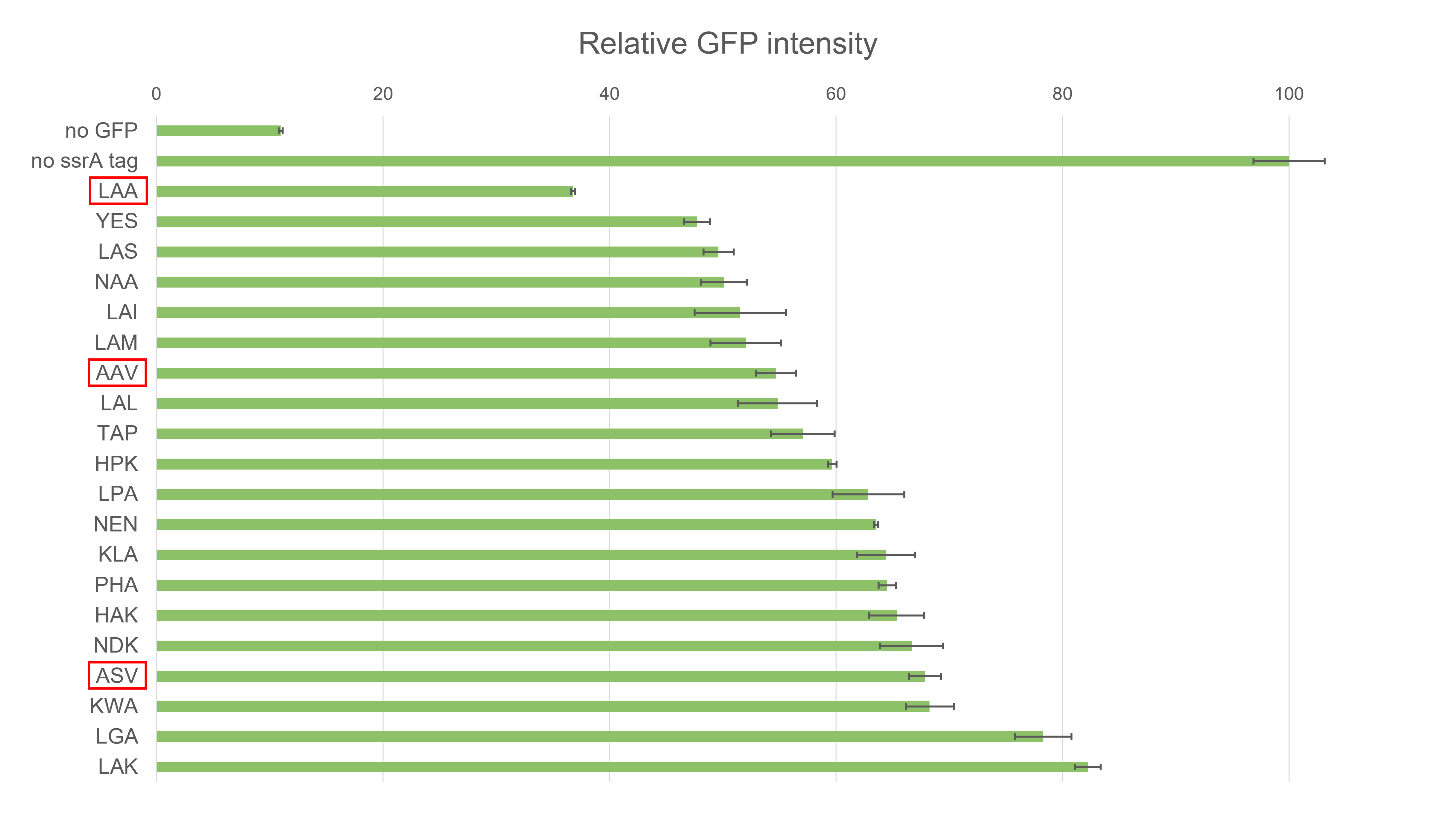
Conclusion
We mutagenized ssrA tag by PCR with mixed primers, and successfully obtained a mutant collection of engineered ssrA tag derivatives which show various protein degradation efficiencies.
Part collection of mutant ssrA tags
In our mutagenesis experiments, we identified 17 mutant tags which show various protein degradation efficiencies (refer to Result). The other mutant tags are listed below.
| Name | Type | Part Name | Designer |
|---|---|---|---|
| BBa_K3815015 | Tag | AANDENYALGA.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815016 | Tag | AANDENYAKWA.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815017 | Tag | AANDENYALAK.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815018 | Tag | AANDENYAHPK.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815019 | Tag | AANDENYALAL.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815020 | Tag | AANDENYANDK.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815021 | Tag | AANDENYANEN.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815022 | Tag | AANDENYAYES.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815023 | Tag | AANDENYAHAK.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815024 | Tag | AANDENYALAS.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815025 | Tag | AANDENYATAP.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815026 | Tag | AANDENYANAA.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815027 | Tag | AANDENYAKLA.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815028 | Tag | AANDENYALPA.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815029 | Tag | AANDENYALAM.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815030 | Tag | AANDENYALAI.mutant SsrA degradation tag | Alexander Liu |
| BBa_K3815031 | Tag | AANDENYAPHA.mutant SsrA degradation tag | Alexander Liu |
Improvement from an Existing Part
This is a part improved from BBa_K1051206. In this part, three C-terminal amino acids LAA are replaced with KWA, resulting in a reduced protein degradation efficiency. Therefore, this part show an increased half-life of fused proteins compared to WT. This part, together with the other mutants described in the part collection above, consists of a large repertoire of protein degradation tags of various efficiencies (refer to Result), which can be applied for fine-tuning of many kinds of synthetic systems that require a precise control of proteins.
Reference
- Flynn, J.M., Levchenko, I., Seidel, M., Wickner, S.H., Sauer, R.T., and Baker, T.A. (2001). Overlapping recognition determinants within the ssrA degradation tag allow modulation of proteolysis. Proc. Natl. Acad. Sci. U. S. A. 98, 10584–10589.
| None |