Part:BBa_K2601040
Tet07-SUMO-yEGFP-HOTag3
Introduction
SUMOylation is a post-translational modification, which is involved in many cellular processes and provide a versatile way to regulate the dynamics of protein-protein interactions. Conjugation of the ubiquitin-related SUMO modifier to target proteins provides a platform for protein-protein interactions and ordered assembly of multi-protein complexes. In humans, three SUMO forms (SUMO1, SUMO2, and SUMO3) can be attached to lysine residues of target proteins. Modification of proteins by SUMO are recognized by SUMO-interacting motifs termed SIMs. In natural process, polySUMOylation recruits distinct interaction partners, such as E3 ubiquitin ligases, that bind to polySUMO chains through tandem SIMs. SIMs bind to a surface patch between the α-helix and a β-sheet of the SUMO protein and extend the β-sheet of SUMO by one additional strand. The SIM either attaches as a parallel or an antiparallel strand to the SUMO β-sheet. Binding is primarily mediated by a stretch of four residues containing 3–4 hydrophobic amino acids (I, V, or L). This core interaction motif is a common property of all SIMs.
Design
Some membrane-less organelles, such as stress granules and P bodies, have been discovered in recent years. Proteins condense into droplets and assemble these organelles through a process called phase separation. Physically, phase separation is the transformation of a one-phase thermodynamic system to a multi phase system, much like how oil and water demix from each other. According to thermodynamics, molecules will diffuse down the gradient of chemical potential instead of concentration. This is exactly why proteins will self organize into granules, diffusing from regions of low concentration to regions of high concentration. Here is an illustration of phase separation in cells.
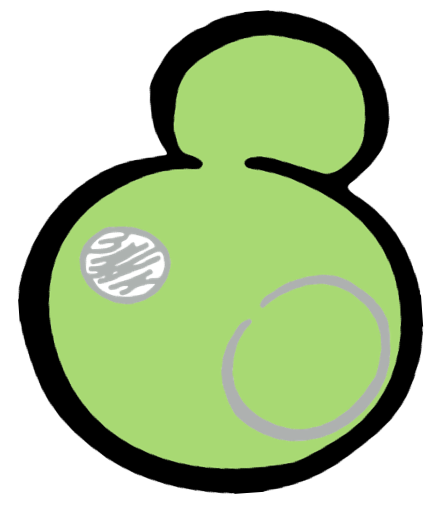 |  |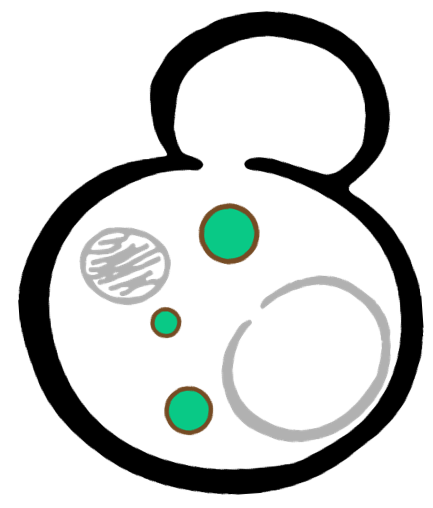  |
In order to rationally design a synthetic organelle based on protein phase separation, we needed a multivalent module and a protein-protein interaction module. The paired SUMO and SIM was one of the bioparts that we chose to introduce protein-protein interaction. SUMO and SIM can dimerize spontaneously. As for the multivalent module, we turned to de novo-designed homo-oligomeric short peptides. These short peptides are called HOTags (homo-oligomeric tags). HOTags contain approximately 30 amino acids. They have high stoichiometry, forming hexamer or tetrameric spontaneously.
The hexameric HOTag3, together with the tetrameric HOTag6, could robustly drive protein phase separation upon protein interaction (achieved by SUMO-SIM module). To verify the feasibility of the system, we fused two fluorescence proteins with the two components of synthetic organelles. Thus, we could observe the self-organization of components and the formation of organelles under fluorescence microscope. We named our system SPOT (Synthetic Phase separation-based Organelle Platform) because it could form granules (fluorescent spots) in yeast. Here is a demonstration of our overall design.
Properties
Previous studies revealed that the electrostatic interactions can be modulated through PTMs on either SIM or SUMO, thereby regulating the dynamics and specificity of their interactions. Reversible phosphorylation/dephosphorylation of the serine or threonine residues in phosphoSIMs dramatically enhances the affinity of SUMO binding. Biophysical measurements of selected interactions revealed a phospho-dependent shift of the dissociation constant Kd from around 50 to 1.5 μM. The specific SUMO and SIM in our design were previously characterized in Michael K. Rosen’s lab. In their studies, the apparent polySUMO-polySIM dissociation constant Kd is 70–10,000 nM, estimated from ITC measurements with (SUMO)m + (SIM)m (m = 1, 2 or 3).
Our results confirmed that SUMO/SIM module fused with HOTags can drive phase separation in yeast.
Furthermore, we used Tet07, an inducible promoter, to control the expression of the SIM component. Doxycycline is the inducer of the promoter. We have tested the strength of Tet07 using flow cytometry.
SIM fused with HOTag6 couldn’t express in the absence of doxycycline. Before we added doxycycline, SUMO-HOtag3 was evenly distributed in the cells and can’t phase separate. But after we added it, phase separation gradually appears. Therefore, we confirmed that the dimerization modules was essential for phase separation and HOTags alone couldn’t induce the process.
 | 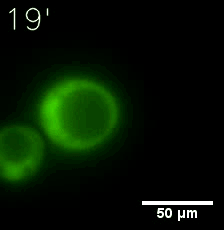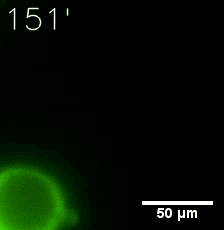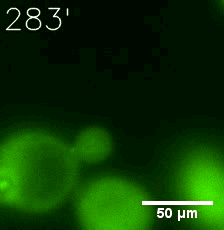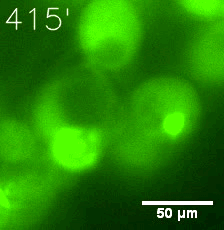 |
 | 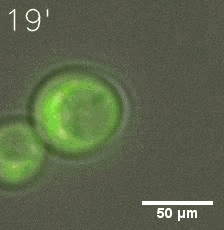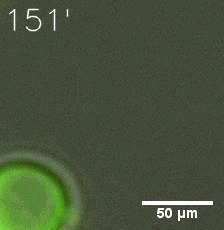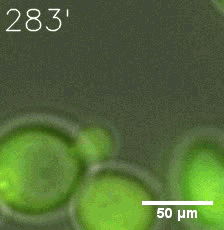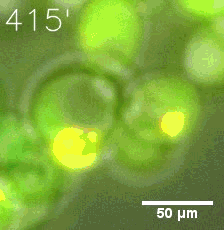 |
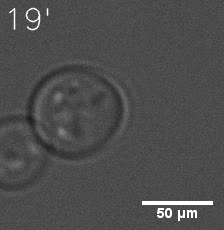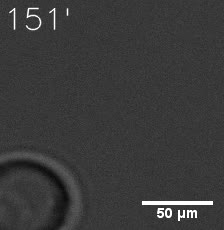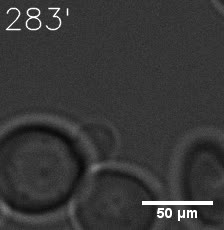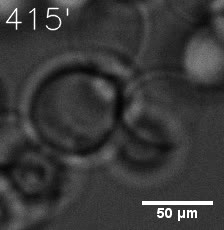
| Figure 7. GIF images of different channels. | |
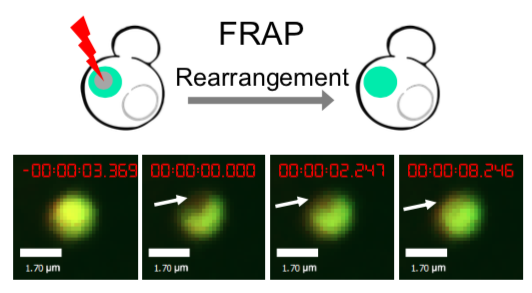
As for the biophysical properties of the system, we assumed the granules were liquid-like. A liquid-like granule is a dynamic system, exchanging mass with cytoplasm rapidly. There are three characteristics which define a liquid-like compartment. First, the compartments should be roughly spherical due to surface tension. It was proved by the 3D-rendered shape of the granules, taken by confocal microscope. Second, two droplets should fuse and coalesce into one droplet spontaneously. Third, the components should undergo rapid internal rearrangement. We used FRAP to verify this criteria. After photobleaching, the fluorescence of granules quickly recovered, which indicated that the granules had rapid mass exchange with cytoplasm.
 | 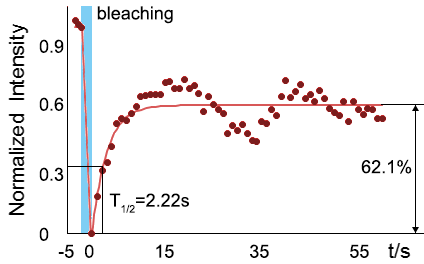 |
| Figure 11. The quantitation of FRAP was determined. The intensity of the fluorescence recovered to approximately 60 percent of the original state. | |
Reference
1. Husnjak, K., Keiten-Schmitz, J., & Müller, S. (2016). Identification and characterization of SUMO-SIM interactions. In SUMO (pp. 79-98). Humana Press, New York, NY.
2. Banani, S. F., Rice, A. M., Peeples, W. B., Lin, Y., Jain, S., Parker, R., & Rosen, M. K. (2016). Compositional control of phase-separated cellular bodies. Cell, 166(3), 651-663.
3. Zhang, Q., Huang, H., Zhang, L., Wu, R., Chung, C. I., Zhang, S. Q., ... & Shu, X. (2018). Visualizing Dynamics of Cell Signaling In Vivo with a Phase Separation-Based Kinase Reporter. Molecular cell, 69(2), 334-346.
4. Woolfson, D. N., Bartlett, G. J., Burton, A. J., Heal, J. W., Niitsu, A., Thomson, A. R., & Wood, C. W. (2015). De novo protein design: how do we expand into the universe of possible protein structures?. Current opinion in structural biology, 33, 16-26.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 1028
Illegal BamHI site found at 1049 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 1695
//chassis/eukaryote/yeast
//proteindomain/binding
| biology | Saccharomyces cerevisiae |