Part:BBa_K2013003
RBS and MHETase
This part contains the coding sequence of MHETase which can hydrolyzes MHET to TPA and EG. MHETase is second enzymes on the downstream of PETase in the bacterium Ideonella sakaiensis 201-F6 that Japanese scientists found.MHET is degraded into two kinds of natural environment harmless substances: terephthalic acid and ethylene glycol. In addition,we did codon optimization before synthesizing it to encode the target product successfully in E. coli.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 579
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 264
Illegal NgoMIV site found at 378
Illegal NgoMIV site found at 834
Illegal AgeI site found at 331
Illegal AgeI site found at 448 - 1000COMPATIBLE WITH RFC[1000]
Experimental Validation
This part is validated through four ways: amplification, PCR, Enzyme cutting, SDS-PAGE and Sequence.
Amplification
Enzyme:Q5
Primer-F:5′- GAATTCGCGGCCGCTTCTAGAGTACTAGAGTCACACAAAAGGGTACTAGATG-3′
Primer-R:5′- CGCTACTAGTATTATTACGGCGGAGCCGCGCAC-3′
Results

PCR
Enzyme:Taq
Primer-F:5′-CCACCTGACGTCTAAGAAAC-3′
Primer-R:5′-GTATTACCGCCTTTGAGTGA-3′
Results

Double digestion
After the assembly ,the plasmid was transferred into the Competent E. coli top10. After culturing overnight in LB,we minipreped the plasmid for double digestion .The first cutting procedure was performed with EcoRI and HindIII restriction endonuclease. The second cutting procedure was performed with PstI and BaHI restriction endonuclease.The plasmid was cutted in a 25μL system at 37 ℃ for 1 hours. The Electrophoresis was performed on a 1% Agarose glu.
Results

Expression of enzymes
SDS-PAGE
piGEM2016-module01* and piGEM2016-module01 are the vectors for the one of the purposes testing the expression of MHETase. The extracellular expression of MHETase was tested from the supernatant fractions collected from E. coli culture after 8h incubation by SDS-PAGE analyzing,which showed that MHETase could be encoded and secreted correctly and successfully. All the samples were obtained after cultivating E.coli BL21 (DE3) in M9 medium for 8h at 30℃, 150rpm. And the backbone is pUC57.

M: protein marker
Line 1: piGEM2016-module01*
Line 2: piGEM2016-module01
Line 3: pUC57
Characterization by 2021iGEM_WFLA_YK_PAO
Improvement of an existing part
Compared to the old part BBa_K2013003, expressing MHETase, we design a new part BBa_K3997005, which is expressing MHETase.
The BBa_K2013003 part is useful as an example. iGEM16_UESTC-China 2016 has cloned this part into pUC57 and did codon optimization before synthesizing it to encode the target product successfully in E. coli.
The difference is that our goal is to build an environmental-friendly method to achieve plastic degradation, and we set up a reaction system to test its property of hydrolysis of MHET. We chose MHETase, to construct our desired engineering strain.
First of all, we constructed a composite part BBa_K3997005, and transformed it into E.coli BL21(DE3). We successfully expressed and purified MHETase protein, and detected the enzyme activity.
Profile
Name: pET28a-MHETase-His
Base Pairs: 1979 bp
Origin: Ideonella sakaiensis 201-F6, synthetic
Properties: Polyethylene terephthalate degradation enzyme
Usage and Biology
Polyethylene terephthalate (PET) is the most widely produced polyester plastic and its accumulation in the environment has become a global concern. At the same time, the daily intake of microplastics by humans is gradually increasing, which damages human health. Therefore, researchers believe that it is important to develop an environmental-friendly plastic degradation method by using microorganisms. Recently, a novel bacterial strain called Ideonella sakaiensis 201-F6 has been discovered that produces a couple of unique enzymes, PETase and MHETase, enabling the bacteria to utilize PET as their sole carbon source.
Figure 1. Action and function of MHETase in MHET degradation.
The enzyme MHETase is a hydrolase, and it represents a key step in the process of microbial PET degradation in I. sakaiensis. It cleaves monoterephthalic acid, the PET degradation product by PETase, to ethylene glycol and terephthalic acid, and it is crucial for hydrolysis of
MHET. We attempt to express the MHETase in E. coli strain to express and purify the protein to test its activity of degradation the MHET (Figure 1). So as to set up a method of environmental-friendly plastic degradation.
Construct design
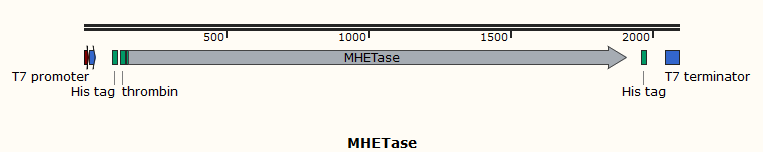
The enzyme MHETase is a hydrolase, and it is crucial for hydrolysis of MHET. To verify this property, we use E. coli as the starting strain and construct an engineered strain of MHETase to explore its biological activity of the hydrolysis of MHET. To purify the protein, we also transfer the plasmid expressing MHETase into BL21(DE3) with a 6×His tag at it’s N-terminal. The enzyme is under the regulation of T7 promoter and can be induced by adding IPTG.
The T7 promoter is often used for protein overexpression. It is powerful and specific. It is completely controlled by T7 RNAP. When T7 RNAP is present in the cell, the T7 expression system occupies an absolute advantage compared to the host expression system. Its expression The speed is 5 times that of the former.
BBa_K3997001
Name: 6×His
Base Pairs: 18bp
Origin: synthetic
Properties: Polyhistidine tag
Usage and Biology
It is an polyhistidine tag, which is used in the purification of recombinant proteins
BBa_K3997001
Name: MHETase
Base Pairs: 1752 bp
Origin: Ideonella sakaiensis 201-F6
Properties: hydrolysis of MHET.
Experimental approach
Production, purification, and activity analysis of MHETase
1. Construction of pET28a-MHETase
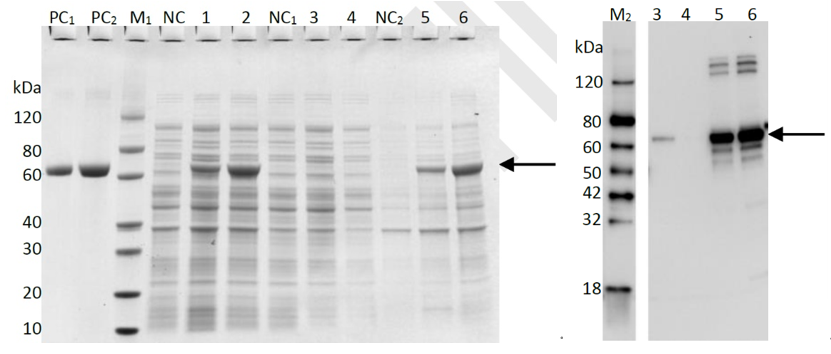
2. Purification of MHETase( ~65.17 kDa)_BL21(DE3) In order to present the function of the part, the MHETase gene was expressed in E. coli under the control of T7 promoter. Then the bacterial cells are collected and crushed, The samples of whole expression cell lysate, supernatant and pellet of cell lysate were analyzed using SDS-PAGE and Western blot (only for his-tag proteins from pET28a vector), which is found in the corresponding protein band of approximately 65kDa (Figure 3).
Lane M1: Protein marker
Lane M2: Western blot marker
Lane PC1: BSA (1μg)
Lane PC2: BSA (2μg)
Lane NC: Cell lysate without induction
Lane 1: Cell lysate with induction for 16h at 15oC
Lane 2: Cell lysate with induction for 4 h at 37oC
Lane NC1: Supernatant of cell lysate without induction
Lane 3: Supernatant of cell lysate with induction for 16h at 15oC
Lane 4: Supernatant of cell lysate with induction for 4 h at 37oC
Lane NC2: Pellet of cell lysate without induction
Lane 5: Pellet of cell lysate with induction for 16h at 15oC
Lane 6: Pellet of cell lysate with induction for 4 h at 37oC
The primary antibody for western blot is anti-His antibody
In addition, MHETase genes were also cloned to the expression vector pGEX-6P-1, which produce recombinant protein fusion with Glutathione-S-transferase (GST) tag.
Lane 1: MHETase Cell lysate without induction for 20 h at 16oC
Lane 2: MHETase Cell lysate with induction for 20 h at 16oC
Lane 3,4,5: GSH elution fractions of purification of lane 2 by GST-affinity chromatography
Proof of function
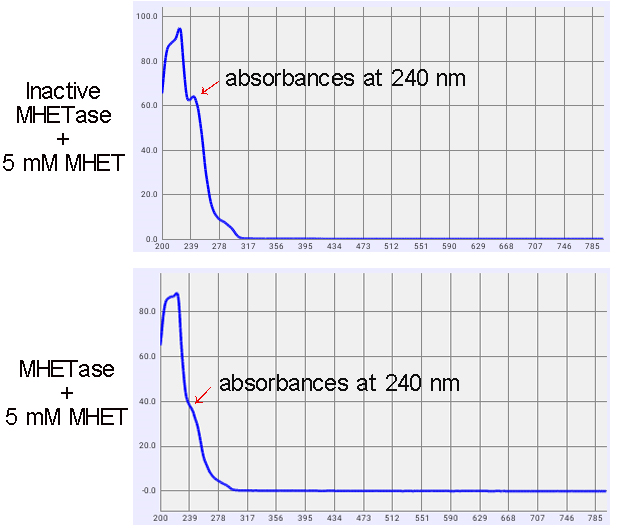
1. Enzyme Activity Tests
The activity of MHETase was indicated by the decline absorbances at a wavelength of 240 nm (Figure 4). The wavelength is the specific absorption of MHET. As shown in figure 2, after 1 d reaction, MHET concentration was dropped by 31.4%.
References
1. Shosuke Yoshida et al. A bacterium that degrades and assimilates poly(ethylene terephthalate), Science (2016).
2. Harry P Austin. et al. Characterization and engineering of a plastic-degrading aromatic polyesterase, PNAS(2018)
3. Chun-Chi Chen et al. General features to enhance enzymatic activity of poly(ethylene terephthalate) hydrolysis, Nature Catalysis(2021).
DUT_China 2021 Contribution
New data collected from laboratory experiments by Ailurus vecTM
Ailurus vecTM can provide the best expression vector for a specific protein in certain throughput. With the sponsorship of Ailurus vecTM, we tried the throughput of N = 1,536 to optimize the output of MHETase in our project. The results are as follows.
In this result, it is not difficult to see that different promoters, RBS, and terminator have obvious differences in the expression of MHETase. Through the service of Ailurus vecTM, we have obtained the following combination of plasmid expression system which is most conducive to the large-scale expression of MHETase.
Backbone: pSB1C3
The results of the expression vector optimization service we obtained at Ailurus vecTM will provide convenience for the iGEM team who will follow-up research on MHETase.
More details can be learnt in this PDF and excel.
text to copy easily
Annotation
- Promoter:
CCTTGATTGCTGTTATTTTTTCCTGGCCCAGCTTTCTTTATGCTGTATCGA
- * Name:747026/-
- 5' UTR/RBS:
GAGTTCCCCGCGCCAGCGGGGATAAACCGTCTTAATCATGCCTAGGAAGTTTTCTA
- * Name: 5UTR_phoA-psiF
- N-terminal tag:
GTTTTTATTTTTTAATGTATTTGTACATGGAGAAAATAAAG
- * Name: 6xHis-tag
- Gene of interest: MHETase
- 3' UTR/Terminator:
CCAATTATTGAAGGCCTCCCTAACGGGGGGCCTTTTTTTGTTTCTGGTCTCCC
- * Name: L3S3P21
New information learned from literature
According to a study by Brandon C. Knott et al., Ideonella sakaiensis can secrete a two-enzyme system to deconstruct polyethylene terephthalate (PET) to its constituent monomers. Specifically, the I. sakaiensis PETase depolymerizes PET, liberating soluble products, including mono(2-hydroxyethyl) terephthalate (MHET), which is cleaved to terephthalic acid and ethylene glycol by MHETase. The research reported a 1.6 Å resolution MHETase structure, illustrating that the MHETase core domain is similar to PETase, capped by a lid domain. Bioinformatics analysis suggests that MHETase evolved from ferulic acid esterases, and two homologous enzymes are shown to exhibit MHET turnover. Analysis of the two homologous enzymes and the MHETase S131G mutant demonstrates the importance of this residue for accommodation of MHET in the active site. Researchers also demonstrated that the MHETase lid is crucial for hydrolysis of MHET and, furthermore, that MHETase does not turnover mono(2-hydroxyethyl)-furanoate or mono(2-hydroxyethyl)-isophthalate. A highly synergistic relationship between PETase and MHETase was observed for thebconversion of amorphous PET film to monomers across all nonzero MHETase concentrations tested. Finally, they compared the performance of MHETase:PETase chimeric proteins of varying linker lengths, which all exhibit improved PET and MHET turnover relative to the free enzymes. These results offered insights into the two-enzyme PET depolymerization system and will inform future efforts in the biological deconstruction and upcycling of mixed plastics.
[1] KNOTT B C, ERICKSON E, ALLEN M D, et al. Characterization and engineering of a two-enzyme system for plastics depolymerization [J]. Proceedings of the National Academy of Sciences, 2020, 117(41):
[2] YOSHIDA S, HIRAGA K, TAKEHANA T, et al. Response to Comment on "A bacterium that degrades and assimilates poly(ethylene terephthalate)" [J]. Science, 2016, 353(6301):
[3] RAUWERDINK A, KAZLAUSKAS R J. How the Same Core Catalytic Machinery Catalyzes 17 Different Reactions: the Serine-Histidine-Aspartate Catalytic Triad of alpha/beta-Hydrolase Fold Enzymes [J]. Acs Catalysis, 2015, 5(10): 6153-76.
[4] AUSTIN H P, ALLEN M D, DONOHOE B S, et al. Characterization and engineering of a plastic-degrading aromatic polyesterase [J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(19): E4350-E7.
| None |