Part:BBa_K1856004
Anderson Strong-citrine
Anderson Strong promoter assembled to citrine and T7 terminator
Usage and Biology
The anderson medium promoter is a constitutive promoter that is part of the collection of Anderson Promoters recovered in a library screen by Chris Anderson. This promoter has been characterized in E. coli, and is available as the biobrick J23100. The Yale iGEM team has assembled this promoter to citrine (an improved version of YFP, with excitation peak at 514nm and emission peak at 527nm) and a T7-terminator to quantify the level of expression in E. coli and in non-model organism hosts.
This construct has been successfully cloned into E. coli using the broad-host range vector pKT230, a RSF1010 derived plasmid, as well as using the pPZP200 plasmid which can be transformed into agrobacterium and rhizobium. Leaky expression of citrine was observed.
RSF1010 plasmids belong to the IncQ group and can be transformed into a wide range of gram-negative bacteria as well as some gram-positive bacteria. These include cyanobacteria genera such as Synechocystis, Synechococcus and Anabaena, rhizobium genera such as Rhizobium and Sinorhizobium, and other genera such as Pseudomonas, Streptomyces and Mycobacterium. The plasmid is mobilizable by conjugation.
We have used a derivative of pKT230 that is compatible with ligation-independent cloning using BsaI. pKT230 has a kanamycin-resistance marker.
pPZP200 is a agrobacterium binary plasmid that can also be transformed into rhizobium genera. We also used a derivative that is LIC compatible for high-efficiency cloning of our constructs. pPZP200 has a spectinomycin-resistance marker.
Beyond the E. coli work, the construct has also been successfully cloned into S. meliloti and PCR-verified.
Characterization
Measured strength
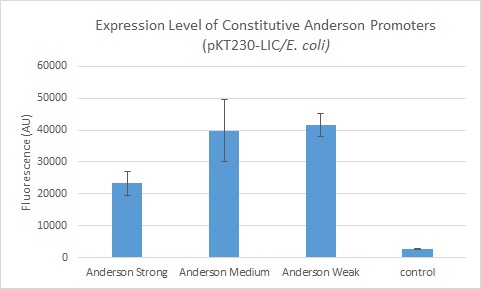
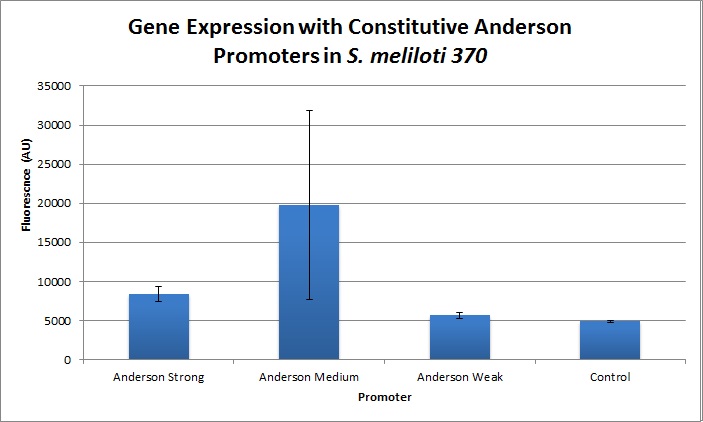
We found fluorescence levels nine times above the baseline level in E. coli cells transformed with the promoter-citrine construct. We also found this promoter to be effective in driving gene expression in S. meliloti, with fluorescence levels of over two times that of the baseline observed.
We further compared expression level of this promoter to two of the other promoters in the Anderson collection. The Anderson lab found relatively expression levels of 1, 0.33, and 0.10 respectively when comparing the promoters J23100 (Anderson Strong), J23110 (Anderson Medium) and J23114 (Anderson Weak). We found the relative expressions to be 1, 1.71 and 1.78 when we transformed our constructs into E.coli: K1856004 (Anderson Strong-Citrine), K1856003 (Anderson Medium-Citrine), K1856002 (Anderson Weak-Citrine). This highlights that even as all three Anderson promoters are effective in driving gene expression in E. coli, the relative expression level may vary depending on the gene fused to the promoter and on the E. coli strain used. The Yale iGEM team's readings were obtained in E. coli DH10B.
Comparing expression levels in S. meliloti (a rhizobium strain), the relative expression values were 1, 2.36, and no significant expression for the Anderson Strong, Anderson Medium, and Anderson Weak promoters respectively.
These fluorescence readings are part of a series of readings taken using different inducible and constitutive promoters to drive citrine expression. See parts K1856000, K1856001, K1856002, K1856003, K1856005, K1856006 for details of the full characterization set.
Obtaining the Anderson weak promoter-citrine construct
The sequences of construct can be found via the table below. The physical DNA can be obtained from:
Via request: The Yale iGEM team has the construct cloned into the broad-host range plasmid, pKT230, a RSF1010 derivative. This plasmid is known to work in cyanobacteria genera such as Synechocystis, Synechococcus and Anabaena, in rhizobium genera such as Rhizobium and Sinorhizobium. The team has also cloned the construct into pPZP200. Both are extremely versatile expression vectors. Please contact the Yale team if you would like to request for an aliquot of the constructs in these plasmids.
Via the Registry distribution: The constructs are included in the Registry distribution, cloned in pSB1C3 backbone.
Sequence and Features
- 10INCOMPATIBLE WITH RFC[10]Illegal EcoRI site found at 812
- 12INCOMPATIBLE WITH RFC[12]Illegal EcoRI site found at 812
Illegal NheI site found at 7
Illegal NheI site found at 30
Illegal NotI site found at 837 - 21INCOMPATIBLE WITH RFC[21]Illegal EcoRI site found at 812
Illegal BglII site found at 47
Illegal BamHI site found at 806
Illegal XhoI site found at 846 - 23INCOMPATIBLE WITH RFC[23]Illegal EcoRI site found at 812
- 25INCOMPATIBLE WITH RFC[25]Illegal EcoRI site found at 812
- 1000COMPATIBLE WITH RFC[1000]
| None |