Part:BBa_K1761001
Outer Membrane Protein X (OmpX) with BamHI-linker and mNeonGreen
This part consists of three subparts, namely the uter membrane protein OmpX, a BamHI linker and the fluorophore mNeonGreen. OmpX is an outer membrane protein with the C- and N-termini in the intracellulair domain. To be able to use OmpX as a scaffold, a unnatural amino acid needs to be introduced. This can be done by implementing the amber stop codon TAG in one of the loops of OmpX via a mutation. With a specific tRNA an azide-functionalized amino acid can be built in, which can be used for the SPAAC click chemistry reaction with DBCO functionalized groups. The BamHI linker is a 213 bp long flexible GGSGGS linker. Using the restriction enzyme BamHI, the linker can become 45 bp shorter. mNeonGreen is a yellow-green fluorescent protein. For a fluorescene measurement, the mNeonGreen fluorophore can be excitated with a laser with a wavelenght of 480 nm and readed out at 517 nm.
Usage and Biology
OmpX, or Outer Membrane Protein X, belongs to a family of highly conserved bacterial proteins that promote bacterial adhesion to and entry into mammalian cells. It presents both C- and N-termini in the intracellular domain (see Figure 1). OmpX consists of an eight-stranded antiparallel all-next-neighbor β barrel. The core of the protein consists of an extended hydrogen-bonding network of highly conserved residues (see Figure 1). [1]
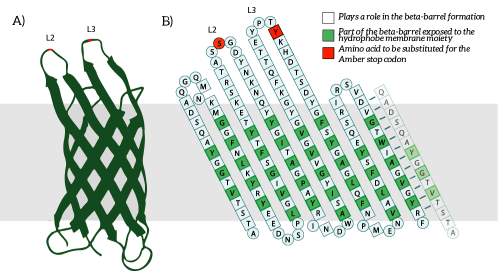
Figure 1: A) The OmpX protein structure has been elucidated through NMR and X-ray crystallography, B) The square residues are important for the secondary structure of OmpX. To keep the structure intact, we introduce an amber stop codon in one of the protruding loops. Figure 5B is adapted from [1].
mNeonGreen is a yellow-green fluorescent protein and is derived from a tetrameric fluorescent protein from cephalochordate Branchiostoma lanceolatum. mNeonGreen is the brightest monomeric green or yellow fluorescent protein yet described and is an excellent fluorescence resonance energy transfer (FRET) acceptor for the newest cyan fluorescent proteins. [2]
Gene Design
This construct consists of three parts, namely OmpX, a BamHI-linker, and mNeonGreen. OmpX is already described as a single part, and that same part, with the amber stop codon TAG incorporated for the integration of the unnatural amino acid, is used for this construct. For more information, see part BBa_K1761000 [1].
mNeonGreen is connected to OmpX with a BamHI-linker. This linker is a 213 bp long flexible GGSGGS linker and by using the restriction enzyme BamHI, the linker can become 45 bp shorter. mNeonGreen was characterized by Shaner et al [2]. The BamHI-linker is inspired by the article "Quantitative Understanding of the Energy Transfer between Fluorescent Proteins Connected via Flexible Peptide Liners" by T.H. Evers et al from 29 August 2006. [3]
These three parts together give a construct with OmpX (1) as transmembrane protein and mNeonGreen (2) as intracellular domain (see Figure 2).
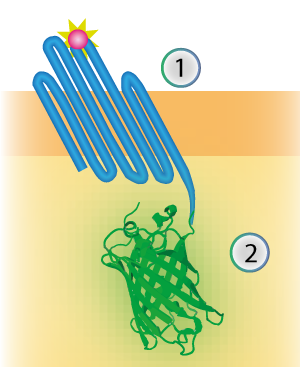
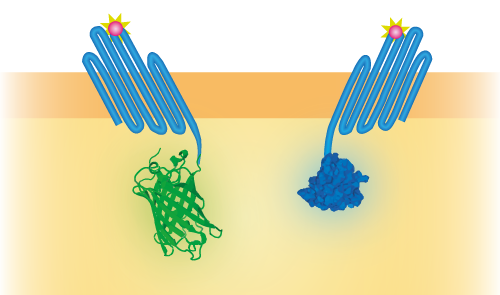
Figure 2: Schematical overview of the expressed OmpX (1) with mNeonGreen (2) in the outer membrane of E.coli.
For all our experiments we used pETDuet-1 as a vector. This vector has two multiple cloning sites, namely MCS-1 and MCS-2. To be able to use this part as a sensor, we inserted both OmpX – NanoLuc and OmpX – mNeonGreen into this vector, both in different multiple cloning sites. In this way, NanoLuc and mNeonGreen can function as a BRET pair (see Figure 3).
Figure 3: Schematical overview of the expressed OmpX with mNeonGreen and OmpX with NanoLuc in the outer membrane of E.coli. The mNeonGreen and NanoLuc domains form a BRET pair.
Sequence
The sequence of our OmpX - mNeonGreen part (see Figure 4) has been verified by sequencing at StarSeq. It contains the prefix and suffix with the correct restriction sites (EcoRI, XbaI, SpeI and PstI). OmpX is 513 bp long, the BamHI-Linker is 213 bp long and mNeonGreen 711 bp.

Figure 4: Snapgene plasmid overview of the BioBrick part BBa_K1761001. It shows the pSB1C3 vector with the prefix (containing the restriction sites EcoRI and XbaI), wildtype OmpX, the BamHI linker, mNeonGreen and the suffix (containting the restiction sites SpeI and PstI).
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 574
Illegal BamHI site found at 619 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Characterization
When mutated with the amber stop codon TAG, OmpX can covalently bind anything by using a bio-orthogonal “click-reaction". The presence of mNeonGreen in this construct is tested with a fluorescence assay.
For all the experiments we used the following vectors: pETDuet-1 with a construct inserted (OmpX + intracellular protein) and pEVOL-pAzF (tRNA + tRNA synthetase, see BBa_K1492002 [2]). Both vectors were transformed into BL21(DE3). The expression was introduced by adding arabinose, IPTG and the unnatural amino acid.
DBCO-PEG4-TAMRA Confirmination
To confirm whether OmpX is in the membrane and whether or not the unnatural amino acid is being incorporated into OmpX, DBCO-PEG4-TAMRA was used. TAMRA is a fluorescent dye that can be used to verify the “click-reaction". If the unnatural amino acid is present, DBCO-PEG4-TAMRA should “click” to the transmembrane protein OmpX and stay there. This can be analyzed with FACS. For more information about how to perform FACS experiments, see our Protocol Page [http://2015.igem.org/Team:TU_Eindhoven/Project/Protocols].
OmpX – mNeonGreen gave the following results after clicking with DBCO-PEG4-TAMRA (see Figure 5). From this it can be concluded that OmpX is in the membrane and that the “click-reaction" works.

Figure 5: FACS result of OmpX - mNeonGreen
Fluorescence Confirmation
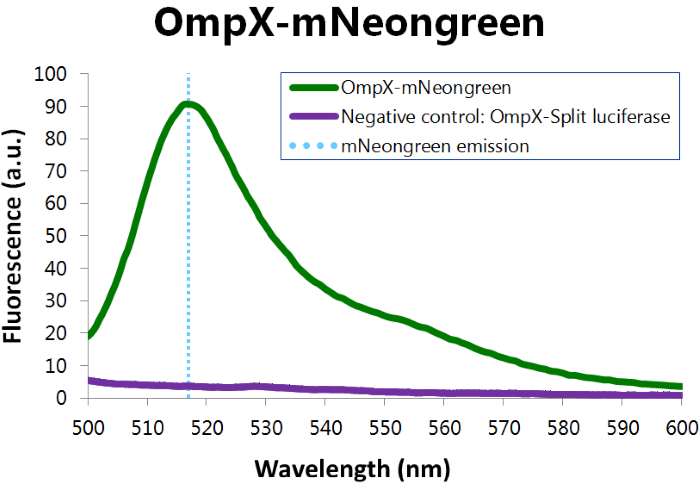
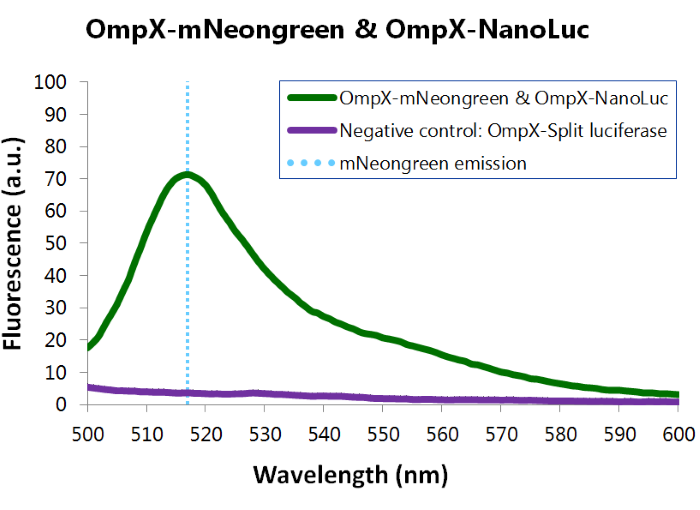
To confirm whether mNeonGreen is present in the bacteria, a fluorescence assay was performed. Excitation took place at a wavelength of 480 nm with a laser. Emission was read out at 517 nm. From this experiment, it can be concluded that mNeonGreen is present and works (see Figure 6). This was also tested with both OmpX – mNeonGreen and OmpX – NanoLuc inserted, which gave the same results (see Figure 7).
For more information about how to perform a fluorescence assay, see our Protocol Page [http://2015.igem.org/Team:TU_Eindhoven/Project/Protocols].
Figure 6: Fluorescence results of OmpX - mNeonGreen
Figure 7: Fluorescence results of OmpX -mNeonGreen and OmpX - NanoLuc
Bioluminescence Confirmation
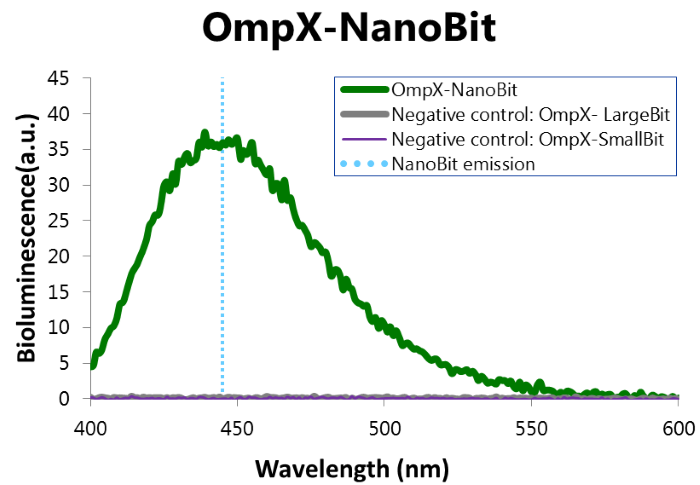
To confirm whether the membrane proteins can compe in close proximity, the OmpX constructs with LargeBit and SmallBit were expressed and a bioluminescence measurement was performed. The results of this experiment are shown in Figure 8.
Figure 8: Bioluminescence of results of the NanoBit construct.
References
[1] J. Vogt and G. E. Schulz, “The structure of the outer membrane protein OmpX from Escherichia coli reveals possible mechanisms of virulence.,” Structure, vol. 7, no. 10, pp. 1301–9, Oct. 1999.
[2] N.C. Shaner et al, “A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum.,” Nature Methods, vol. 10, no. 5, pp. 407-409, Mar. 2013.
[3] T.H. Evers et al, “Quantitative Understanding of the Energy Transfer between Fluorescent Proteins Connected via Flexible Peptide Linkers.,” Biochemistry, vol. 45, no. 44, pp. 13183-92, Nov. 2006.
//function/fret
| emission | 517 nm |
| excitation | 480 nm |