Part:BBa_K1471002
RBS with MerE.
This part contains a RBS for A. thaliana followed by a coding sequence named MerE from the bacterial operon mer. We did codon optimization of MerE gene, for it's expression in Arabidopsis thaliana.
RBS
The initiation of protein biosynthesis is a major determinant of the efficiency of gene expression at the translational level. It is known that the nucleotide sequences around the AUG translation initiation codon act as an important signal to trigger the initiation of the translation event. (Kozak, 1987) Understanding regulatory mechanisms of protein synthesis in eukaryotes is essential for the accurate annotation of genome sequences. Kozak reported that the nucleotide sequence GCCGCC(A/G)CCAUGG (AUG is the initiation codon) was frequently observed in vertebrate genes and that this 'consensus' sequence enhanced translation initiation. However, later studies using invertebrate, fungal and plant genes reported different 'consensus' sequences. (Nakagawa, 2007)
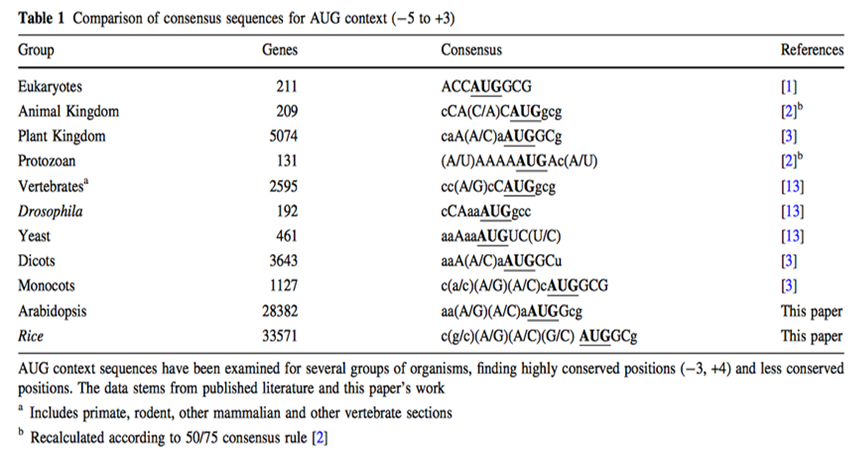
Although for any protein analysis it is crucial to know exactly which region of the mRNA is coding for protein, prediction of the translation initial site is still an unsolved problem. In eukaryotes, the scanning model postulates that the ribosome attaches first to the 5' end of the mRNA and scans along the 5'-3' direction until it encounters the first AUG. While translation initiation from the first AUG holds true in many cases, there are also a considerable number of exceptions. In these exceptions the main determining factor in AUG choice is the context of the respective codon. (Rangan, 2008) Two decades ago, a consensus sequence for the context of the AUG codon in higher plants was proposed on basis of very limited number of sequences. Joshi and colleagues got the generally assumption that the consensus sequence found (aaaaacaA(A/C)aAUGG) is valid for all plant clades, but Rangan found out that a considerable degree of variation between plants and major between the major eukaryotic groups along with some conserved features. However, the large variability and the periodicity suggest that general structural features rather than precise nucleotide sequence may play an important role in transcription initial site. (Rangan, 2008)
MerE
Brief description merE
Our part is a yeast RBS with an 8KDa CH3Hg or Hg+2 transmembrane bacterial transporter.
Biology
MerE is a gene is part of the mer operon, a collection of bacterial genes specialized on the tolerance to various compounds of mercury including methylmercury. It is naturally found in the transposon Tn21 from the plasmid NR1 Shigella flexneri or MB1 in the case of Bacillus megaterium. The general mechanism of the operon can be observed on the following representation (Das S., Dash H. R., 2012):
Schematic presentation of mer operon in narrow-spectrum Gram-negative mercury-resistant bacteria (Das S., Dash H. R., 2012)
Where organomercury compounds are transported inside the bacteria by merP, merT, mer E and merG, followed by the transformation of organic mercury by merB into its ionic form and the reduction from Hg+2 into volatile Hg0 by mer A with the help of NADPH(Das S., Dash H. R., 2012).
Although the transportation of methylmercury is barely understood, there is evidence that recombinant E. coli and other transformed GRAM negative bacteria are able to accumulate mercury thanks to the transformation of organic mercury into its ionic form(Das S., Dash H. R., 2012).
Behaviour
merE has been characterized previously on A. thaliana as a potential mercury accumulator and transporter. In Kyono,M., et al (2013)´s study shoot and root growth were observed to become more tolerant in transgenic Arabidopsis compared with controls. As it can be seen on the figure:
E represents transformed A. thalianaSusceptibility of transgenic plants to Hg(II) and CH3Hg(I). (Kiyono M. , et al, 2013)
MIT_MAHE 2020
Role of cysteine and histidine residues
In order to investigate the molecular function of merE in the transport of methylmercury, a study was conducted with six merE variants with specific site-directed mutations. By comparison methylmercury uptake by cell with intact merE and mutated MerE, it was demonstrated that the cysteine pairs (Cys28 and Cys30) within the first transmembrane domain of MerE and the histidine residue (His51) on the perisplasmic face located between the first and second transmembrane domains were required for MerE mediated CH3Hg(I) transport across the cell membrane. The cysteine pairs especially may play a critical role in transport of CH3Hg(I) and Hg(II). However, it is still unclear why the merE gene found in the T21 mer operon, confers resistance only to Hg(II) and not CH3Hg(I).
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
References
Rangan, et al. (2008). Analysis of Context Sequence Surrounding Translation Initiation Site from Complete Genome of Model Plants. New York University. [Online] Retrieved october 14th 2014 from: http://www.nyu.edu/projects/vogel/Reprints/Rangan_MolBt08.pdf Nakagawa, et al. (2007).
Diversity of preferred nucleotide sequences around the translation initiation codon in eukaryote genomes. Oxford University Press. [Online] Retrieved october 14th 2014 from: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2241899/ Liu Q, Xue Q. (2005).
Comparative studies on sequence characteristics around translation initiation codon in four eukaryotes. Zhejiang University. [Online] Retrieved october 14th 2014 from: http://www.ias.ac.in/jgenet/Vol84No3/317.pdf Kozak, M. (1989).
Circumstances and mechanisms of inhibition of translation by secondary structure in eucaryotic mRNAs. American Society for Microbiology (ASM). [Online] retrieved october 14th 2014 from: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC363665/
Das S., Dash H. R., (2012). Bioremediation of mercury and the importance of bacterial mer genes. National Institute of Technology.India: International Biodeterioration & Biodegradation. Volume 75. Pages 207-213
Kiyono M. , et al (2013) Increase methylmercury accumulation in Arabidopsis thaliana expressing bacterial broad-spectrum mercury transporter MerE. Springer. Issue 3; Pages 1-13
Kiyono M., Sone Y., Nakamura R., et al (2013) Role of MerC, MerE, MerF, MerT, and/or MerP in Resistance to Mercurials and the Transport of Mercurials in Escherichia coli. Biological and Pharmaceutical Bulletin. Volume 36; Issue 11; pages 1835-1841
Sone, Y., Uraguchi, S., Takanezawa, Y., Nakamura, R., Pan-Hou, H., & Kiyono, M. (2017). Cysteine and histidine residues are involved in Escherichia coli Tn21 MerE methylmercury transport. FEBS open bio, 7(12), 1994–1999. https://doi.org/10.1002/2211-5463.12341
| n/a | RBS with MerE. |


