Part:BBa_K1122674
Plac+fused PDC-ADH
This part encodes pLac promoter + LacZ reporter followed by ethanol production module (fused Pdc and AdhB)
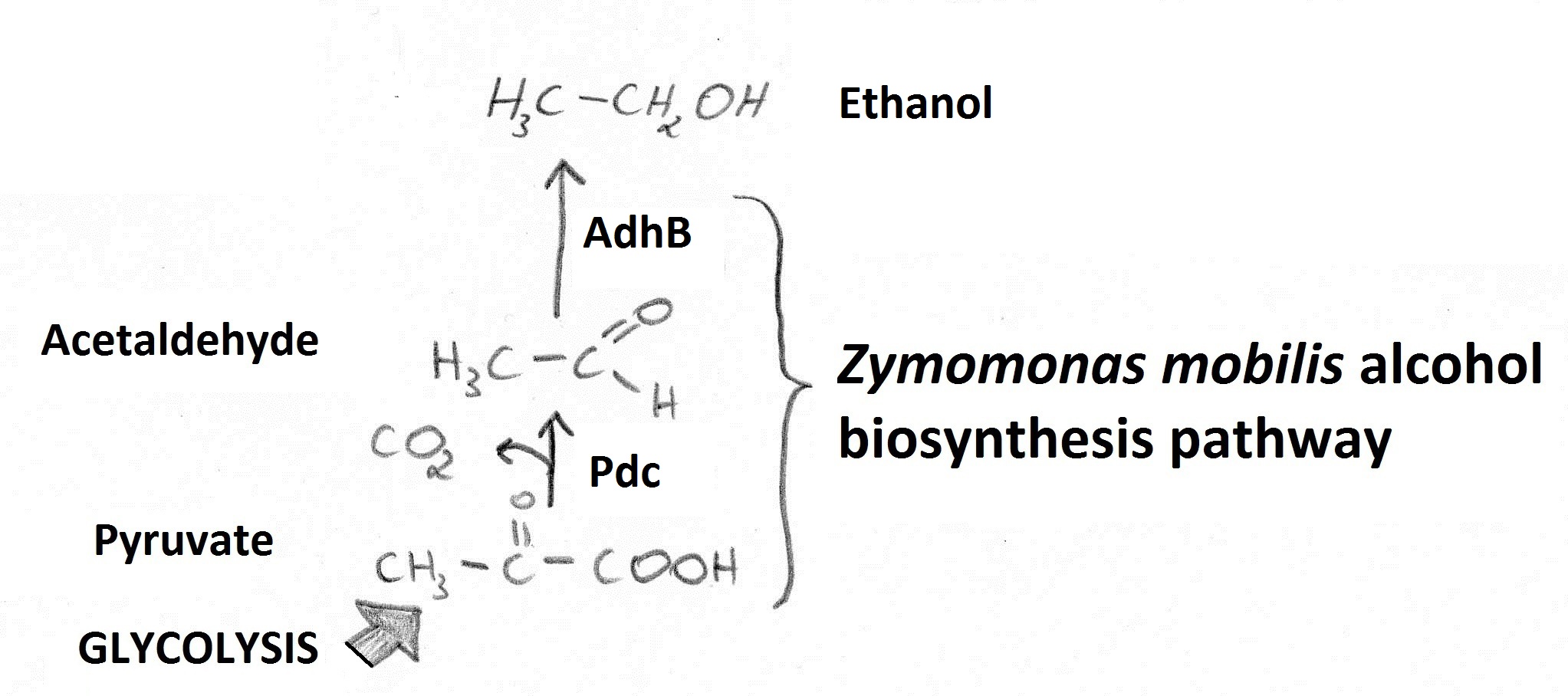
Microbial production of ethanol is of great importance due to its possible application as a biofuel. Increasing ethanol yields in bacteria is potentially beneficial as those are able to utilise a wider variety of renewable, biomass-derived carbon sources compared to standard ethanol producer: Sacharomyces cerevisiae. Our goal was to increase ethanol yields using a fusion of Zymomonas mobilis pyruvate decarboxylase (Pdc) and alcohol dehydrogenase B (AdhB). Reactions catalysed by those enzymes (Figure A) enable conversion of pyruvate to ethanol.
Figure A. Ethanol is generated from pyruvate (fed into the pathway from glycolysis) via an Acetaldehyde intermediate.
We based the work on the hypotheses that flow of substrate from one enzyme to another should be facilitated in the fusion protein, and that due to this less of the toxic aldehyde intermediate should be released into the cell. Other fusion proteins were already presented to increase amounts of final product produced (Wang et al., 2011) what further encouraged us to undertake this work.
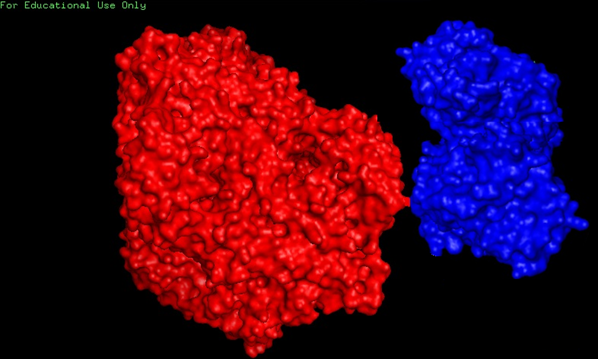
We have decided that we will perform the fusion of C terminus of Pdc to N terminus of AdhB. What could potentially generate a protein of a following structure: (figure B)
Figure B. Putative structure of a fused protein. Pdc tetramer (red) is an active state of this enzyme. C terminus of one of the members was linked to an N terminus of AdhB (blue). Presented AdhB enzyme is a dimer and just one of the monomers is linked to Pdc.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 1109
Illegal AgeI site found at 2315 - 1000COMPATIBLE WITH RFC[1000]
Generation of protein fusion
In order to generate fusion of pyruvate decarboxylase (pdc) and alcohol dehydrogense B (adhB) Mutagenesis with Blunt- Ended Ligation (MABEL) was used.
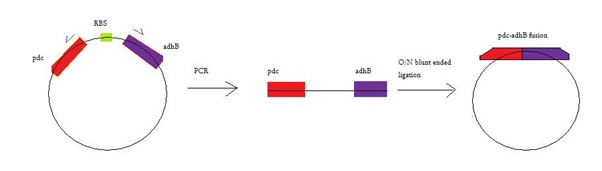
A pair of primers was designed complementary with 3' end of pdc and 5' end of adhB. Figure 1 represents the MABEL process.
Fig1. Represents MABEL process used for generation of fused pdc-adhB construct. Primer sequences used: Forward: GCATCAAGCACCTTTTATATCC; Reverse: CAGCAGTTTATTCACCGGTTTAC. See appendix of the Edinburgh University 2013 iGEM team figure 1 for full details on the part sequence, primer binding sites and deleted region.
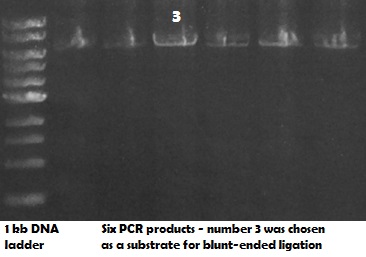
Generated PCR product (see Fig 1.) was analysed on an agarose gel (Fig 2.):
Fig 2.Presence of a single PCR product of correct size (app. 6000 bp) on a 0.8% agarose gel. 1kb NEB DNA ladder was used. Several replicates of the reaction were loaded on a gel.
Evidence for presence of protein fusion
DNA level evidence
Primers were designed to amplify the region of fusion (region with deleted RBS). Those primers were used for PCR on fused and non-fused pdc and adhB.
Figure 3. 2.5% agarose gel analysing PCR product created using primers amplifying the region of putative gene fusion. PCR products amplyfing fused protein region (first 3 lanes) are smaller than those amplyfing the same region in non-fused genes(last 3 lanes).
The same set of primers was used for sequencing of pdc-adhB fusion construct. Obtained results indicate that the fused state was present on a DNA level:
File:Bioethanol sequencing.zip Presence of following DNA sequence: GTAAACCGGTGAATAAACTGCTGGCATCAAGCACCTTTTATATCC corresponding to reverse complement of reverse MABEL primer followed by forward MABEL primer sequences indicates that gene fusion was obtained.
Protein level evidence
In order to express fused pdc-adhB the construct was placed under the control of J33207 – IPTG inducible promoter combined with LacZ reporter.
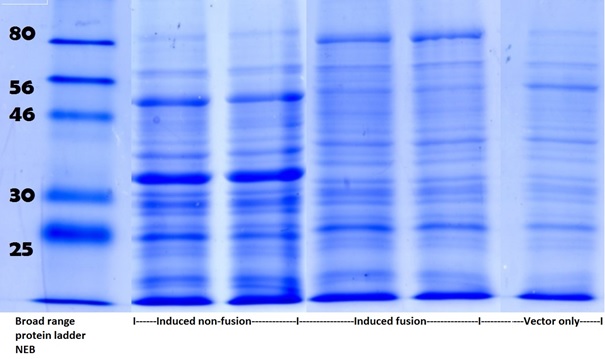
Full grown cultures expressing pdc-adhB fusion and non-fusion were lysed and analysed on SDS-PAGE:
Fig 4. Scan of SDS-PAGE following Coomassie Brilliant Blue G250 staining. Within induced non-fusion lanes two intense bands are present (with size corresponding to pdc and adhB). Within induced fusion lanes described bands are missing and an additional band of increased size is observed. In vector only lane none of above described bands is present. To see full SDS-PAGE go to Appendix figure 2 on Edinburgh iGEM 2013 wiki
In order to present that functionality of adhB is attributed to a peptide of a different mass in fusion state than in non-fusion state a native PAGE was performed.
Fig 5. Native PAGE stained for AdhB activity. Enzymatic activity can be attributed to a peptide of an increased mass in the fusion state (left) than in non-fusion state (right). The vector only sample was loaded in-between.
Moreover, it was presented that pdc and adhB activity can be attributed to same gel band showing that fusion state was indeed obtained:
Fig 6. Native pages with adhB and pdc staining. Both activities are present in a band just below the well. Due to different acrylamide concentration used the fused protein was unable to migrate into the gel as opposed to PAGE present on figure 5.
Functional Parameters
Pdc and adhB catalyse conversion of pyruvate to ethanol via acetaldehyde intermediate (link to K173003).
It was hypothesized that generation of pdc-adhB fusion will increase ethanol yields. This suggestion was based on two observations:
1.Flow of substrate from one enzyme to another should be facilitated in fused protein state;
2.Due to better flow of intermediate product less of it should be released into the cell. As this intermediate is a toxic acetaldehyde moiety the fusion state could decrease toxicity of ethanol production to bacterial cell;
It was established that highest ethanol yields can be achieved in glucose as a carbon source. LB+ 4% glucose was used for ethanol production. Following ethanol levels were achieved in the experiment:
Fig 7. Ethanol production graph. Cultures were first incubated for 24h in aerobic environment. Following that they were switched into fermentative conditions for 48 hours. For statistical analysis considering multiple runs of the experiment see figure 3 and 4 in Appendix to the Results of Edinburgh iGEM 2013
Activity of adhB as a fusion member
It was observed that the adhB activity seems to be lower in the fusion state than in non-fused state (see fig. 5). To quantify this an NAD+ reduction assay was performed.
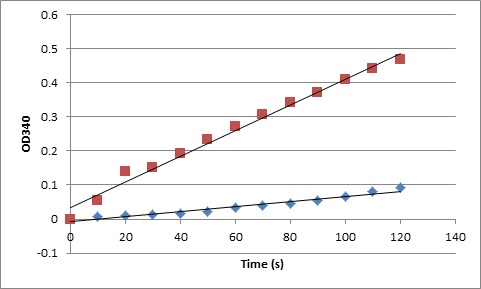
Fig 8. OD340 readings over time correspond to ctivity of adhB in fused (blue) and non-fused (red) state.
From those measurements and protein concentration data obtained using Pierce assay specific activity was calculated: AdhB – fusion: 0.091 U/mg. AdhB – non-fusion: 1.79 U/mg
Those indicate that the adhB has lower activity in the fusion state. Despite this an increased ethanol concentration is obtained. Thus this observation provides an evidence that the fusion itself is the reason for increased ethanol production, not i.e. highier activity of one enzyme in fusion state.
Microscopy of cells expressing fused protein
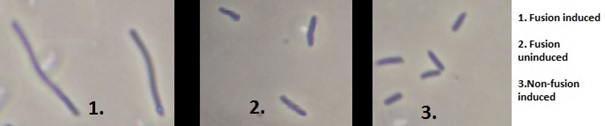
Phase contrast microscopy of cell cultures (fig 9.) indicates that cells expressing fused pdc-adhB constructs are more likelly to be elongated.
Figure 9. Phase contrast microscopy images (x100) obtained for selected cultures. To see full microscopy slides go to Appendix figures 5 to 7 on Edinburgh iGEM 2013 wiki.
This might be explained by possible formation of long clusters of fused proteins. This peomenon would be achieved due to dimeric nature of active adhB and tetrameric nature of active pdc.
References
INGRAM, L. O., CONWAY, T., CLARK, D. P., SEWELL, G. W. & PRESTON, J. F. 1987. GENETIC-ENGINEERING OF ETHANOL-PRODUCTION IN ESCHERICHIA-COLI. Applied and Environmental Microbiology, 53, 2420-2425.
WANG, C., YOON, S.-H., JANG, H.-J., CHUNG, Y.-R., KIM, J.-Y., CHOI, E.-S. & KIM, S.-W. 2011. Metabolic engineering of Escherichia coli for alpha-farnesene production. Metabolic Engineering, 13, 648-655.
| None |