Part:BBa_K1033000
Tyrosine ammonia-lyase (TAL) with RBS
Tyrosine ammonia-lyase (TAL) is an enzyme which catalyzes the formation of p-coumaric acid (aka p-hyroxycinnamic acid) from tyrosine. It belongs to the family of ammonia-lyases, enzymes that catalyze the deamination of amino acids [1]. P-coumaric acid is an important precursor in many metabolic pathways. TAL also has a secondary function as a phenylalanine ammonia-lyase (PAL) which catalyses the formation of cinnamic acid. This bacterial version have proved to have a hundred fold higher preference for tyrosine as a substrate, as opposite to many eukaryotic versions. [2].

Applications
This biobrick could have lots of applications. First it can by itself produce the antioxidant p-coumaric acid from tyrosine. The enzyme could also be mutated so that it catalyzes the formation of trans-cinnamic acid from phenylalanine.
P-coumaric acid is also needed as a precursor for many metabolites in the phenylpropanoid pathway. Examples of these are resveratrol that we have worked on. It can be used as a precursor for these biobricks we have made: BBa_K1033001, BBa_K1033002 , BBa_K1033003
Also many other compunds like flavanones, lignins and anthocyanins can be produced from p-coumaric acid. With this enzyme, we have opened up the phenylpropanoid pathway for bacteria. [4]
How to characterize it
P-coumaric acid shows no visible color, so one has to characterize it with the help of methods like spectrophotometry and chromatography.
Characterization data from Uppsala iGEM 2013
Summary
Tyrosine ammonia lyase (TAL) from rhodobacter sphaeroides was obtained from J.Conrado et al[4] We biobricked TAL with the ribosome binding site B0034 and overhangs in a single pcr. We also made a version with 6-HIS-tag for enzyme expression analysis. TAL was also mutagenized to remove illegal Not1 site. We verify all of our genetical constructs with sequencing.
We have expressed TAL in E. coli DH5alpha and E. coli nissle, a probiotic E.coli obtained from Trieste iGEM 2012. TAL will also be further characterized in lactobacillus by transforming the construct with our shuttle vector. This construct can be used to produce the precursor for example our resveratrol.
We managed to clone out and biobrick tyrosine ammonia lyase and verify the biobrick by sequencing. We have also done successful characterization of this part, showing that it works as expected. We managed to express our enzyme and detect it in a western blot, and also detected our metabolite in both spectrophotometry and HPLC. The biobrick was characterized in E. coli d5alpha and E. coli nissle. The cloning of our biobrick, Tyrosine ammonia lyase was done from rhodobacter sphaerides with the RBS B0034, that should work in various organisms like lactobacillus and E. coli. Sequencing was done at GATC biotech and Uppsala Genome center using sanger sequencing.
Western blot
As previously mentioned, we have succeeded in expressing the enzyme tyrosine ammonia-lyase (TAL) in e-coli. To enable the detection of this protein by anti-his antibodies, 6-histidine tags were incorporated in the
sequence. By doing this we would be able detect our enzyme with anti-his antibodies.
We expressed our protein with a promoter working in both lactobacillus and E. coli. We did this to later be able to transfer this TAL construct to lactobacilli strains.
The size of our protein was calculated using ProtParam to 54.9 kDA.

Figure 1:SDS-page and western blot. Expression of Tyrosine ammonia lyase with constitutive promoters in E. coli. The negative control has no bands, showing that there is no natural protein in wildtype E. coli with the same attributes. The positive control shows that the staining has worked. The two sample lanes 2 and 3 have clear bands around 55 kDA which suggests that these two recombinant E. coli strains produce the TAL protein. 1.Positive control, 2.TAL with CP8 promotor, 3.Tal with J23110 promotor, 4.Negative control
Spectrophotometry
As the next step, we have made a spectrophotometric assay of our metabolite p-coumaric acid produced by E. coli. By using n-octanol and a two-phase extraction, we were able to extract our metabolite from the LB medium incubated with TAL producing bacteria. After this purification step, we were able to characterize our biobrick on a spectrophotometer. The positive control we used had a final p-coumaric concentration of 500 µl. The spectrophotometry experiments show that there is a somewhat lower concentration in the sample extracts after 48 hours of stationary phase incubation compared to the positive control.[Protocol]



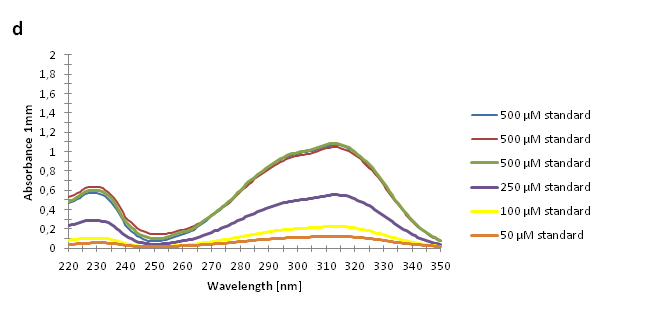
Figure 2. Absorbance spectra of n-octanol extracts collected from bacterial cultures. Samples were collected 21 h and 48 h after 30 °C incubation. The negative control is an extract from a strain with no TAL gene on the transformed plasmid. The positive control is an extract of a culture of the same strain as the negative control but with added p-coumaric acid to a concentration of 500 µM before extraction. P-coumaric acid absorbance spectra has two peaks. The one around 305 nm is preferable to detect because of background noise from components found in LB that do transfer to the octanol phase in the extraction. (a) Spectra from the strain with TAL CDS with promoter J23110. (b) Spectra from the strain with TAL CDS with promoter CP8. (c) Spectra from the strain with TAL CDS with promoter J23101.(d) Standards of pure p-coumaric acid solved in n-octanol. The standards show that p-coumaric acid has a broad peak in absorbance around 305 nm.
High pressure liquid chromatography
To further prove p-coumaric acid production in E. coli, we did a HPLC of our bacterial culture. We managaed to detect our metabolite at a retention time around 9 minutes. [Protocol]
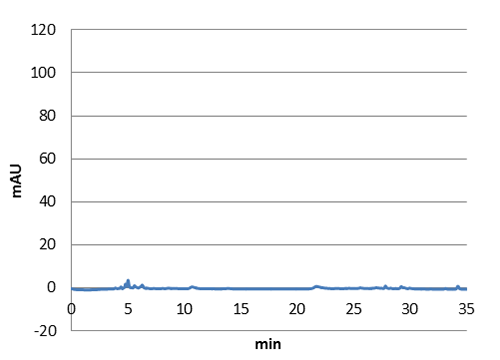
Figure 3: HPLC chromatogram of a n-octanol extract from a E. coli culture without our biobrick tyrosine ammonia lyase. No peak at 9 minutes can be detected in this negative control.

Figure 4. Graph showing the HPLC result of a standard sample with pure p-coumaric acid (50 µM) in n-octanol.

Figure 5. Graph showing the HPLC result of an octanol extract prepared from E. coli expressing tyrosine ammonia lyase. Reverse phase HPLC with a C18 matrix was used. A mix of acetate buffer and acetonitrile were used as mobile phase in this gradient elution experiment. The absorbance of the eluate was detected at 303 nm. The peak for p-coumaric acid can be seen ~9 min: the same retention time as the standard sample above.
References
1.Zhixiong Xue, Michael McCluskey, Keith Cantera, F. Sima Sariaslani, Lixuan Huang (2007) Identification, characterization and functional expression of a tyrosine ammonia-lyase and its mutants from the photosynthetic bacterium Rhodobacter sphaeroides. J Ind Microbiol Biotechnol 34:599-604
2. J.A. Kyndt, T.E. Meyer, M.A Cusanovich, J.J. Van Beeumen (2002) Characterization of a bacterial tyrosine ammonia lyase, a biosynthetic enzyme for the photoactive yellow protein. FEBS Letters 512 240-244
3. Robert J. Conrado et al, DNA guided assembly of biosynthetic pathways promotes improved catalytic effiency. Nucleic Acids Research , 2012, Vol 40 NO 4, 1879-1889
4. Joseph A Chemler1 and Mattheos AG Koffas2, Metabolic engineering for plant natural product biosynthesis in Joseph A Chemler1 and Mattheos AG Koffas2, Science Direct, elsevier
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 45
Illegal NgoMIV site found at 877
Illegal AgeI site found at 140
Illegal AgeI site found at 306 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 1335
//chassis/prokaryote/ecoli/nissle
//chassis/prokaryote/lactobacillus
//collections/probiotics
//collections/probiotics/production
| None |
