Part:BBa_K801090
phenylalanine ammonia lyase (PAL) + yeast consensus sequence
phenylalanine ammonia lyase (PAL) coding region from Rhodosporidium toruloides preceeded by the yeast consensus sequence for improved expression.
Background and priniciples
Usage and Biology
Text
Biosynthesis
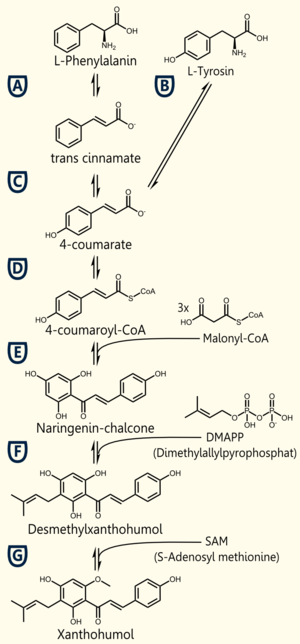
The biosynthetic pathway of 4-coumaroyl-coenzyme A starts with the conversion of L-Phenylalanine to cinnamate, being catalyzed by phenylalanin ammonia lyase (PAL) [A]. PAL also shows activity in converting tyrosine to p-coumarate, but with a lower efficiency [B]. The cinnamate 4-hydroxylase (C4H) catalyzes the synthesis of p-hydroxycinnamate from cinnamate and 4-coumarate [C]: CoA ligase (4CL) converts p-coumarate to its coenzyme-A ester, activating it for reaction with malonyl CoA [D] [Trantas et al., 2009].
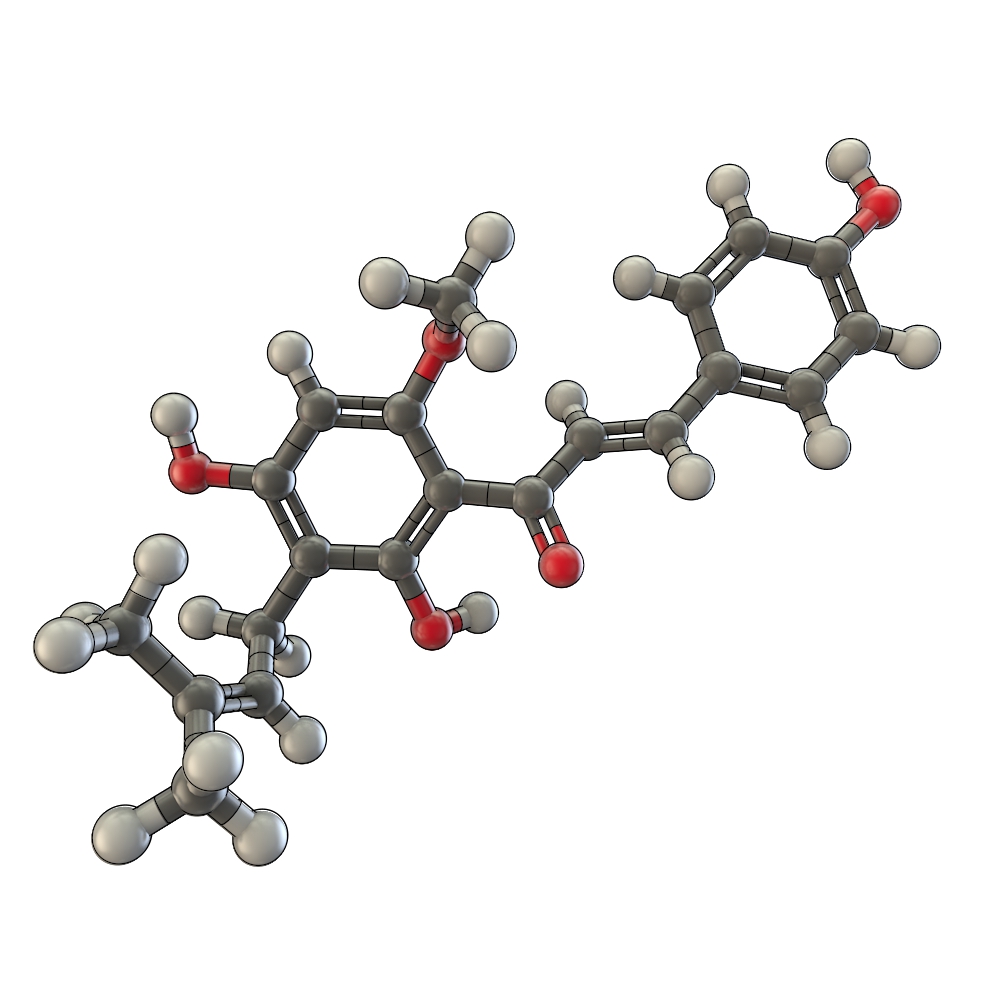
The flavonoid biosynthetic pathway starts with the condensation of one molecule of 4-coumaroyl-CoA and three molecules of malonyl-CoA, yielding naringenin chalcone. This reaction is carried out by the enzyme chalcone synthase (CHS) [E]. Chalcone is isomerised to a flavanone by the enzyme chalcone flavanone isomerase (CHI). From these central intermediates, the pathway diverges into several side branches, each resulting in a different class of flavonoids, such as xanthohumol.
Characterization
text
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 2096
Illegal BamHI site found at 358
Illegal XhoI site found at 409
Illegal XhoI site found at 472
Illegal XhoI site found at 490
Illegal XhoI site found at 568
Illegal XhoI site found at 769
Illegal XhoI site found at 1012
Illegal XhoI site found at 1759 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 928
Illegal NgoMIV site found at 1174
Illegal NgoMIV site found at 1342
Illegal NgoMIV site found at 1485
Illegal NgoMIV site found at 1819 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 469
Illegal BsaI site found at 799
Illegal BsaI site found at 805
Illegal BsaI.rc site found at 1930
Illegal SapI site found at 759
References
- http://www.ncbi.nlm.nih.gov/pubmed/12481418 Gerhauser et al., 2002 Gerhauser, C., Alt, A., Heiss, E., Gamal-Eldeen, A., Klimo, K., Knauft, J., Neu- mann, I., Scherf, H.-R., Frank, N., Bartsch, H., and Becker, H. (2002). Cancer chemopreventive activity of xanthohumol, a natural product derived from hop. Mol Cancer Ther, 1(11):959–69.
- http://www.ncbi.nlm.nih.gov/pubmed/10752639 Henderson et al., 2000 Henderson, M. C., Miranda, C. L., Stevens, J. F., Deinzer, M. L., and Buhler, D. R. (2000). In vitro inhibition of human p450 enzymes by prenylated flavonoids from hops, humulus lupulus. Xenobiotica, 30(3):235–51.
- http://www.ncbi.nlm.nih.gov/pubmed/14704995 Jiang and Morgan, 2004 Jiang, H. and Morgan, J. A. (2004). Optimization of an in vivo plant p450 monooxygenase system in Saccharomyces cerevisiae. Biotechnol Bioeng, 85(2):130–7.
- http://www.ncbi.nlm.nih.gov/pubmed/10737704 Miranda et al., 2000a Miranda, C. L., Aponso, G. L., Stevens, J. F., Deinzer, M. L., and Buhler, D. R. (2000a). Prenylated chalcones and flavanones as inducers of quinone reductase in mouse hepa 1c1c7 cells. Cancer Lett, 149(1-2):21–9.
- http://www.ncbi.nlm.nih.gov/pubmed/10418944 Miranda et al., 1999 Miranda, C. L., Stevens, J. F., Helmrich, A., Henderson, M. C., Rodriguez, R. J., Yang, Y. H., Deinzer, M. L., Barnes, D. W., and Buhler, D. R. (1999). Antiproliferative and cytotoxic effects of prenylated flavonoids from hops (Humulus lupulus) in human cancer cell lines. Food Chem Toxicol, 37(4):271–85.
- http://www.ncbi.nlm.nih.gov/pubmed/10995285 Miranda et al., 2000b Miranda, C. L., Stevens, J. F., Ivanov, V., McCall, M., Frei, B., Deinzer, M. L., and Buhler, D. R. (2000b). Antioxidant and prooxidant actions of prenylated and nonprenylated chalcones and flavanones in vitro. J Agric Food Chem, 48(9):3876–84.
- http://www.ncbi.nlm.nih.gov/pubmed/11038156 Miranda et al., 2000c Miranda, C. L., Yang, Y. H., Henderson, M. C., Stevens, J. F., Santana-Rios, G., Deinzer, M. L., and Buhler, D. R. (2000c). Prenylflavonoids from hops inhibit the metabolic activation of the carcinogenic heterocyclic amine 2-amino-3-methylimidazo[4, 5-f]quinoline, mediated by cdna-expressed human cyp1a2. Drug Metab Dispos, 28(11):1297–302.
- http://www.ncbi.nlm.nih.gov/pubmed/19631278 Trantas et al., 2009 Trantas, E., Panopoulos, N., and Ververidis, F. (2009). Metabolic engineering of the complete pathway leading to heterologous biosynthesis of various flavonoids and stilbenoids in Saccharomyces cerevisiae. Metab Eng, 11(6):355–66.
- http://www.ncbi.nlm.nih.gov/pubmed/11240137 Yilmazer et al., 2001a Yilmazer, M., Stevens, J. F., and Buhler, D. R. (2001a). In vitro glucuronidation of xanthohumol, a flavonoid in hop and beer, by rat and human liver microsomes. FEBS Lett, 491(3):252–6.
- http://www.ncbi.nlm.nih.gov/pubmed/11181488 Yilmazer et al., 2001b Yilmazer, M., Stevens, J. F., Deinzer, M. L., and Buhler, D. R. (2001b). In vitro biotransformation of xanthohumol, a flavonoid from hops (Humulus lupulus), by rat liver microsomes. Drug Metab Dispos, 29(3):223–31.
//cds/biosynthesis
//chassis/eukaryote/yeast
| None |