Part:BBa_K525311
Fusion Protein of S-Layer SgsE and Firefly-Luciferase
Fusion protein of S-layer SgsE and firefly luciferase.
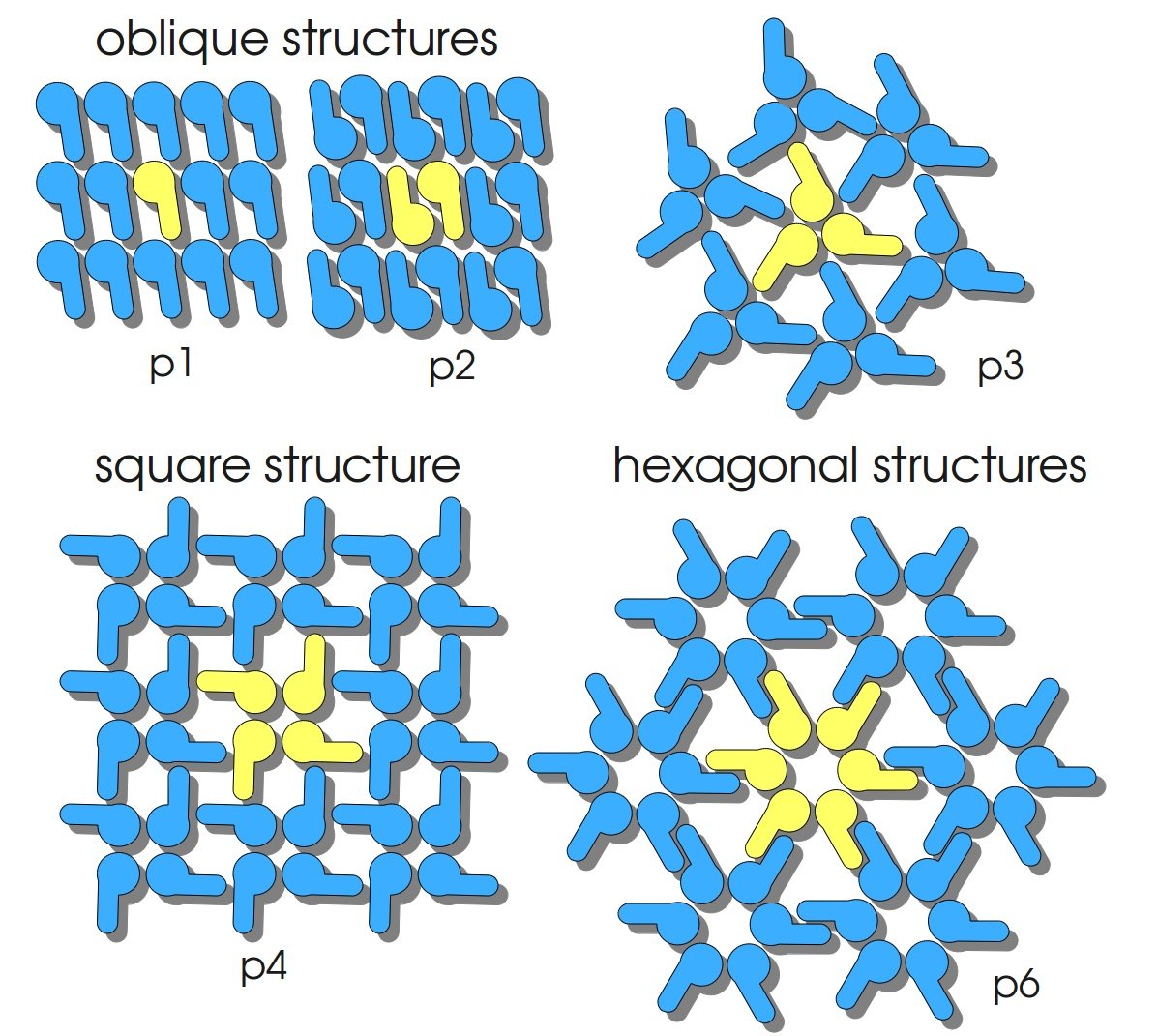
S-layers (crystalline bacterial surface layer) are crystal-like layers consisting of multiple protein monomers and can be found in various (archae-)bacteria. They constitute the outermost part of the cell wall. Especially their ability for self-assembly into distinct geometries is of scientific interest. At phase boundaries, in solutions and on a variety of surfaces they form different lattice structures. The geometry and arrangement is determined by the C-terminal self assembly-domain, which is specific for each S-layer protein. The most common lattice geometries are oblique, square and hexagonal. By modifying the characteristics of the S-layer through combination with functional groups and protein domains as well as their defined position and orientation to eachother (determined by the S-layer geometry) it is possible to realize various practical applications ([http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.2006.00573.x/full Sleytr et al., 2007]).
Usage and Biology
S-layer proteins can be used as scaffold for nanobiotechnological applications and devices by e.g. fusing the S-layer's self-assembly domain to other functional protein domains. It is possible to coat surfaces and liposomes with S-layers. A big advantage of S-layers: after expressing in E. coli and purification, the nanobiotechnological system is cell-free. This enhances the biological security of a device.
This S-layer fusion protein is used to characterize purification methods and immobilization behaviour of enzymes fused to an S-layer.
Important parameters
| Experiment | Characteristic | Result |
|---|---|---|
| Expression (E. coli) | Localisation | Cytoplasm, soluble |
| Compatibility | E. coli KRX and BL21(DE3) | |
| Induction of expression | expression of T7 polymerase + IPTG or lactose | |
| Purification | Molecular weight | 143.8 kDa |
| Theoretical pI | 6.02 | |
| Reporter | Luminescence | |
| Immobilization behaviour | Immobilization time | 4 h |
| Materials binding BBa_K525311 | Silicon dioxide, polyethylene sulfone |
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 167
Illegal BglII site found at 1022 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 76
Illegal AgeI site found at 4054 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 1657
Illegal SapI.rc site found at 3211
Expression in E. coli
The SgsE gene under the control of a T7 / lac promoter (BBa_K525303) was fused to firefly luciferase (BBa_K525999) using Freiburg BioBrick assembly for characterization experiments.
The SgsE|luciferase fusion protein was overexpressed in E. coli KRX after induction of T7 polymerase with 0.2 % L-rhamnose and induction of lac operator with 1 mM IPTG.
The following cultivation was carried out in a [http://www.gmi-inc.com/BioEngineering-KLF-Small-Laboratory-Fermenter.html#product_desc Bioengineering KLF] bioreactor with Bioengineering DCU and software. A sequencer which automatically pumped an inducer solution after 4 h cultivation time to start protein expression was implemented. Other parameters were:
- Medium: [http://2011.igem.org/Team:Bielefeld-Germany/Protocols/Materials#HSG_medium HSG medium] with 20 mg L-1 chloramphenicol
- Culture volume: 2.5 L
- Starting OD600: 0.4
- DO: 60 % airsaturation (controlled with stirrer cascade starting with 200 rpm)
- pH: 7.0 (controlled with 20 % phosphoric acid and 2 M NaOH)
- Antifoam: BASF pluronic XXX
- Induction solution: 0.2 % L-rhamnose and 1 mM IPTG
The following figure shows the expression of the SgsE | luciferase S-layer fusion protein BBa_K525311 in E. coli KRX in HSG medium with autoinduction sequencer as described above. Optical density, activity of the fused luciferase, dissolved oxygen and agitator speed are plotted against the cultivation time.
Purification
After the analysis of cultivations with expression of SgsE | luciferase fusion proteins, different cell fractions were analyzed. It could be seen that the proteins do not form inclusion bodies in E. coli and that they are soluble. This has the advantage that the proteins carrying an enzyme as fusion proteins do not have to be treated with denaturating agents like urea which could destroy the enzyme.
To capture the protein from the cell lysate an ion exchange chromatography (IEX) was carried out (binding with pH 7.0, 25 mM NaCl, quaternary amine beads, elution with 100 mM NaCl). A lot of protein was found in the flow-through. When concentrating and rebuffering the proteins for a hydrophobic interaction chromatography (HIC) afterwards with PES (polyethylene sulfone) dead-end 10 kDa membranes (sartorius Vivaspin 15) nearly all of the protein was lost. The S-layer proteins stuck to the membrane. Some could be removed again from the membrane after cutting out the filter and incubate it in ddH2O over-night. This problem has to be kept in mind when using this S-layer.
Stability
To test whether the S-layer protein enhances the half-life of the firefly luciferase, immobilization experiments were carried out. After the IEX described above, the elution fraction was immobilized on silicon dioxide beads or just diluted with HBSS buffer which is used for the immobilization / recrystallization of SgsE S-layer proteins. The immobilization is carried out at room temperature for 4 h. It could be seen that the luciferase activity nearly expired during this time (in the positive and the negative control). So the S-layer SgsE could not stabilize the luciferase at room temperature.
//function/reporter/light
//proteindomain/internal
| biology | S-Layer |
| chassis | E. coli |
| control | PT7 |
| origin | Geobacillus stearothermophilus |