Difference between revisions of "Part:BBa K2965021"
| Line 5: | Line 5: | ||
===Usage and Biology=== | ===Usage and Biology=== | ||
| − | + | AsCas12a RNPs bind to DNA and start to move along it by a hopping mechanism. When the AsCas12a RNPs encounter a potential target site containing a PAM sequence, they begin to initiate R-loop formation by forming base-pair hybrids between the crRNA and the target DNA strand. If sufficient matched nucleotides exist in the PAM-proximal region, the R-loop formation will be stable. After the stable R-loop conformation is formed, AsCas12a, via the activity of its RuvC domain, first cleaves the non-target DNA strand and then cleaves the opposite, target DNA strand, regardless of the presence of a PAM sequence. Finally, AsCas12a rapidly releases the PAM-distal DNA fragment but continues to hold the PAM-proximal portion of DNA[1]. | |
===Characterization=== | ===Characterization=== | ||
====Result==== | ====Result==== | ||
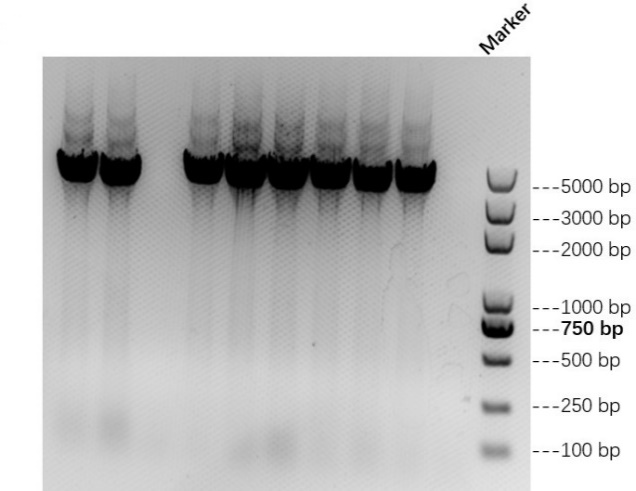
| − | + | We link AsCas12a fragment and pET-28a(+) plasmid to AbCas12a expression plasmid with His tag. Then we transfer recombinant plasmid into E. coli BL21 and test it by colony PCR. Result is shown in Figure 1 below. We also verify it by TSINGKE Biologocal Technology Institute sequencing. | |
[[File:BBa K2965021-Figure 1. AsCas12a colony PCR..png|center|500px|thumb|'''Figure 1. AsCas12a colony PCR.''']] | [[File:BBa K2965021-Figure 1. AsCas12a colony PCR..png|center|500px|thumb|'''Figure 1. AsCas12a colony PCR.''']] | ||
| − | + | We induce AsCas12a expression and purify it by gravity-flow column with Ni-NTA Sefinose(TM) Resin. SDS-PAGE and Western Blot is used to test protein purification as shown in Figure 2 and Figure 3. | |
[[File:BBa K2965021-Figure 2. AsCas12a SDS-PAGE..png|center|500px|thumb|'''Figure 2. AsCas12a SDS-PAGE.''']] | [[File:BBa K2965021-Figure 2. AsCas12a SDS-PAGE..png|center|500px|thumb|'''Figure 2. AsCas12a SDS-PAGE.''']] | ||
[[File:BBa K2965021-Figure 3. AsCas12a WB..png|center|500px|thumb|''' Figure 3. AsCas12a WB.''']] | [[File:BBa K2965021-Figure 3. AsCas12a WB..png|center|500px|thumb|''' Figure 3. AsCas12a WB.''']] | ||
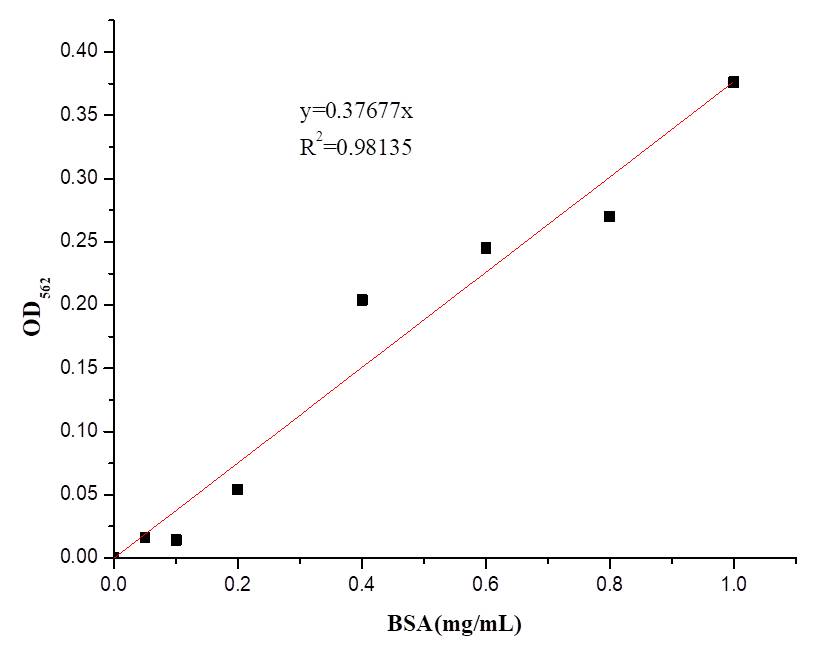
| − | + | We also determine the protein concentration with BCA kit. | |
[[File:BBa K2965021-Figure 4. BSA standard curve..png|center|500px|thumb|'''Figure 4. BSA standard curve.''']] | [[File:BBa K2965021-Figure 4. BSA standard curve..png|center|500px|thumb|'''Figure 4. BSA standard curve.''']] | ||
| − | + | Finally, we test the activity of AsCas12a by detecting the fluorescent. The extracted AsCas12a has ability to cut DNA compared with the control group and is significantly better than the bought AsCas12a. | |
[[File:BBa K2965021-Figure 5. AsCas12a activity assay..png|center|500px|thumb|''' Figure 5. AsCas12a activity assay.''']] | [[File:BBa K2965021-Figure 5. AsCas12a activity assay..png|center|500px|thumb|''' Figure 5. AsCas12a activity assay.''']] | ||
Revision as of 08:09, 21 October 2019
AsCas12a (AsCpf1)
AsCas12a (AsCpf1) comes from Acidaminococcus sp. BV3L6, it is a programmable DNA endonuclease guided by a single CRISPR RNA (crRNA). Targeting requires a crRNA complementary to the target site as well as a 5' TTTN protospacer adjacent motif (PAM) on the DNA strand opposite the target sequence. Once it recognizes its target, it will be activated and exhibits a “collateral effects” of promiscuous DNase activity.
Usage and Biology
AsCas12a RNPs bind to DNA and start to move along it by a hopping mechanism. When the AsCas12a RNPs encounter a potential target site containing a PAM sequence, they begin to initiate R-loop formation by forming base-pair hybrids between the crRNA and the target DNA strand. If sufficient matched nucleotides exist in the PAM-proximal region, the R-loop formation will be stable. After the stable R-loop conformation is formed, AsCas12a, via the activity of its RuvC domain, first cleaves the non-target DNA strand and then cleaves the opposite, target DNA strand, regardless of the presence of a PAM sequence. Finally, AsCas12a rapidly releases the PAM-distal DNA fragment but continues to hold the PAM-proximal portion of DNA[1].
Characterization
Result
We link AsCas12a fragment and pET-28a(+) plasmid to AbCas12a expression plasmid with His tag. Then we transfer recombinant plasmid into E. coli BL21 and test it by colony PCR. Result is shown in Figure 1 below. We also verify it by TSINGKE Biologocal Technology Institute sequencing.
We induce AsCas12a expression and purify it by gravity-flow column with Ni-NTA Sefinose(TM) Resin. SDS-PAGE and Western Blot is used to test protein purification as shown in Figure 2 and Figure 3.
We also determine the protein concentration with BCA kit.
Finally, we test the activity of AsCas12a by detecting the fluorescent. The extracted AsCas12a has ability to cut DNA compared with the control group and is significantly better than the bought AsCas12a.
Reference
[1] Jeon Y, Choi YH, Jang Y, Yu J, Goo J, Lee G, et al. Direct observation of DNA target searching and cleavage by CRISPR-Cas12a. Nature Communications. 2018;9(1):2777-11. [2] Mohanraju, P., Oost, J. v., Jinek, M. and Swarts, D. C. (2018). Heterologous Expression and Purification of the CRISPR-Cas12a/Cpf1 Protein. Bio-protocol 8(9): e2842. DOI: 10.21769/BioProtoc.2842.
Sequence and Features
- 10INCOMPATIBLE WITH RFC[10]Illegal PstI site found at 196
Illegal PstI site found at 1258
Illegal PstI site found at 3667
Illegal PstI site found at 3865 - 12INCOMPATIBLE WITH RFC[12]Illegal PstI site found at 196
Illegal PstI site found at 1258
Illegal PstI site found at 3667
Illegal PstI site found at 3865 - 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 380
Illegal BglII site found at 1922
Illegal BglII site found at 2231 - 23INCOMPATIBLE WITH RFC[23]Illegal PstI site found at 196
Illegal PstI site found at 1258
Illegal PstI site found at 3667
Illegal PstI site found at 3865 - 25INCOMPATIBLE WITH RFC[25]Illegal PstI site found at 196
Illegal PstI site found at 1258
Illegal PstI site found at 3667
Illegal PstI site found at 3865
Illegal NgoMIV site found at 3466 - 1000COMPATIBLE WITH RFC[1000]