Part:BBa_K2549013
TEV protease
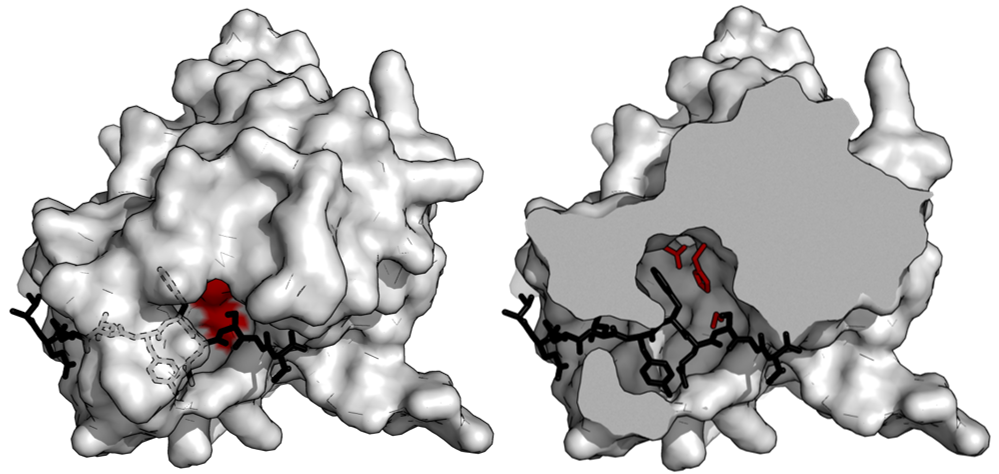
This part is a mutant TEV (tobacco etch virus) protease. The residue 219 of the wild-type TEV protease is mutated from serine to valine (S219V) to remove autoinactivation and perform a more efficient cleavage[1]. TEV protease is one of the best-characterized enzymes of the viral proteases which have more stringent sequence specificity. We used it to build our IMPLY logic gates. It can also be utilized by other iGEM teams to create their own engineered fusion proteins cleavage system.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal SapI.rc site found at 316
Applications
The main use of TEV protease[2] is for removing affinity tags from purified recombinant fusion proteins. The reason for that is due to the high sequence specificity. This specificity allows for the controlled cleavage of proteins when the preference sequence is inserted into flexible loops. It also makes it relatively non-toxic in vivo as the recognized sequence scarcely occurs in proteins.

Please refer to Media:Tev-faq-7pages.pdf for more details on TEV protease.
TEV in Tango Assay for GPCRs
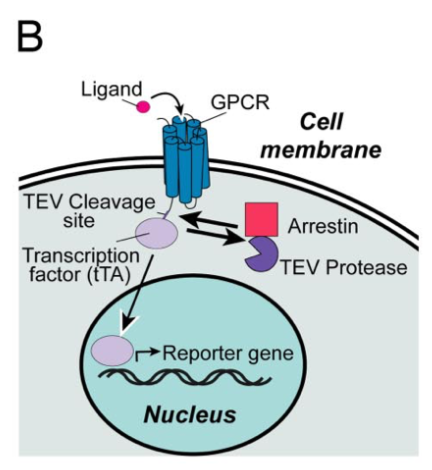
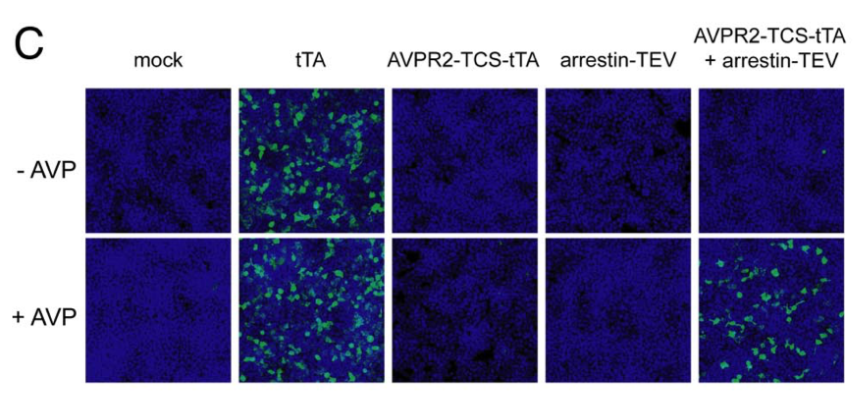
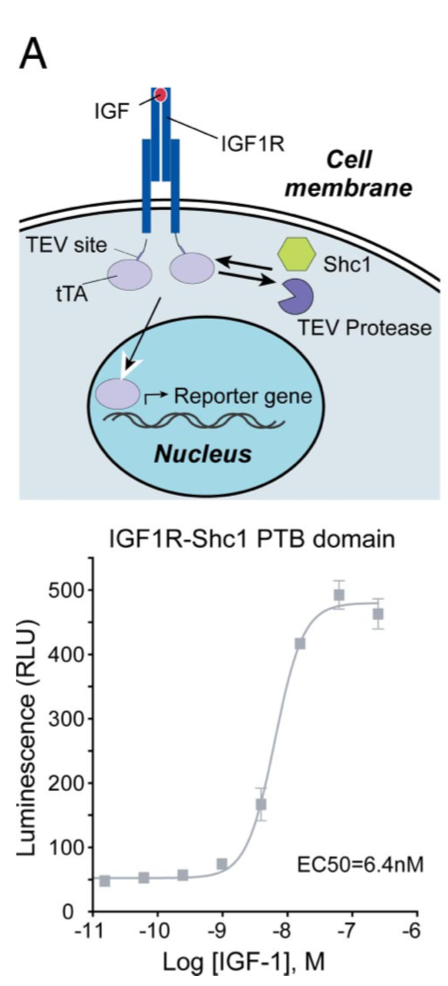
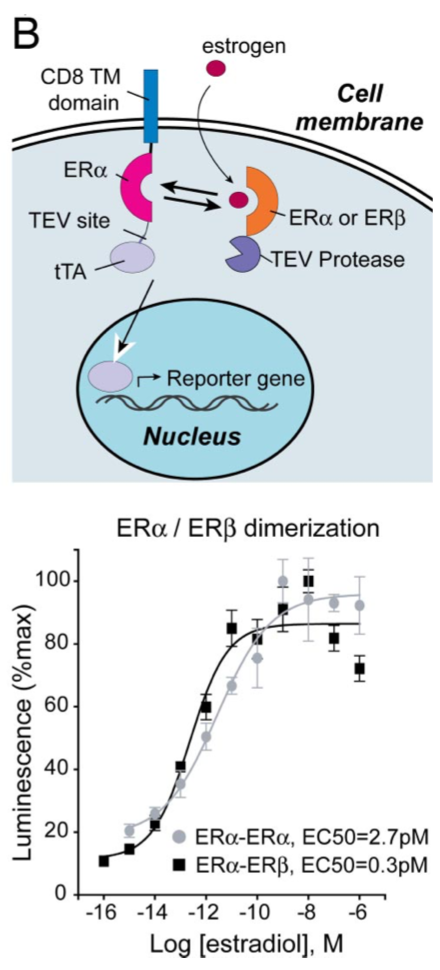
Barnea G et al 2008[3] wrote: We have developed an experimental strategy to monitor protein interactions in a cell with a high degree of selectivity and sensitivity. A transcription factor is tethered to a membrane-bound receptor with a linker that contains a cleavage site for a specific protease. Activation of the receptor recruits a signaling protein fused to the protease that then cleaves and releases the transcription factor to activate reporter genes in the nucleus. This strategy converts a transient interaction into a stable and amplifiable reporter gene signal to record the activation of a receptor without interference from endogenous signaling pathways. We have developed this assay for three classes of receptors: G protein-coupled receptors, receptor tyrosine kinases, and steroid hormone receptors. Finally, we use the assay to identify a ligand for the orphan receptor GPR1, suggesting a role for this receptor in the regulation of inflammation.
TEV in ChaCha Design
Kipniss NH et al 2017[4] wrote: We found that the hM3D-CRISPR ChaCha exhibited better reporter gene activation (4.4 fold) compared to the hM3D- CRISPR Tango (1.2 fold) after 1-day treatment with CNO. The Tango design displayed higher leakiness than the ChaCha design with significant GFP expression without CNO treatment. The observation that ChaCha outperforms Tango may suggest that fusing the small TEVp (28 kDa) to the receptor likely preserves the conformational fidelity and activity of the GPCR molecule, while fusing a larger domain such as dCas9-VPR (220 kDa) to GPCR may compromise its conformation by exposing the C-terminal tail of a GPCR and permit leaky interactions with ARRB2.
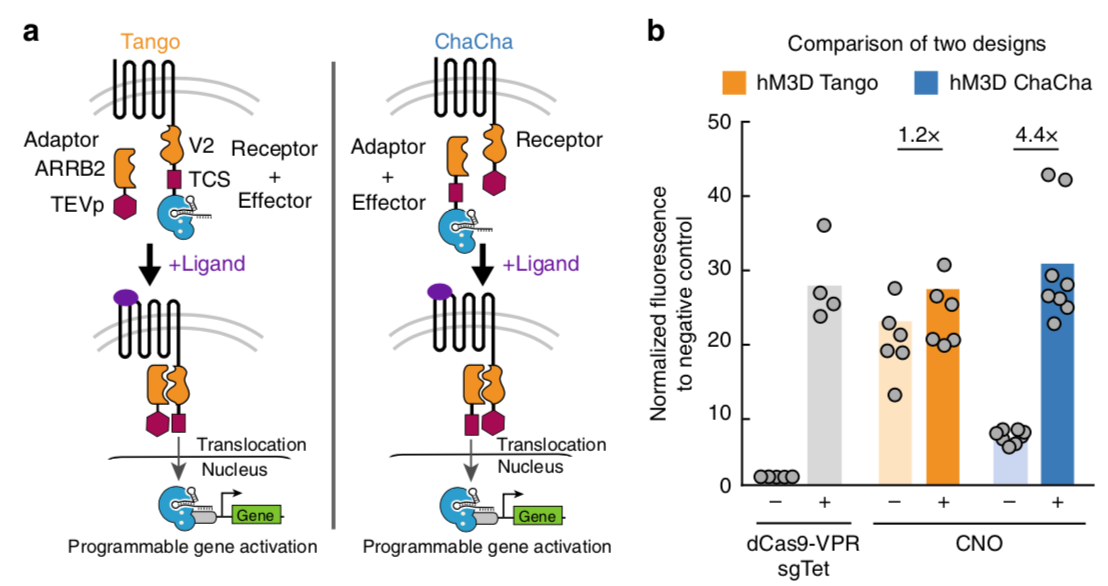
(b) Comparison of the performance of Tango (orange) and ChaCha (blue) designs using the synthetically evolved GPCR, hM3D in HEK293T cells with or without 20 μM clozapine-N-oxide (CNO), the ligand for hM3D (see Methods section for detailed experimental procedure). The free dCas9-VPR (gray) with and without a targeting sgRNA (sgTet) are used as positive and negative controls. The data were normalized to the free dCas9-VPR without a targeting sgRNA. The data represent two independent experiments with 4–8 technical replicates, and the bars represent the mean.
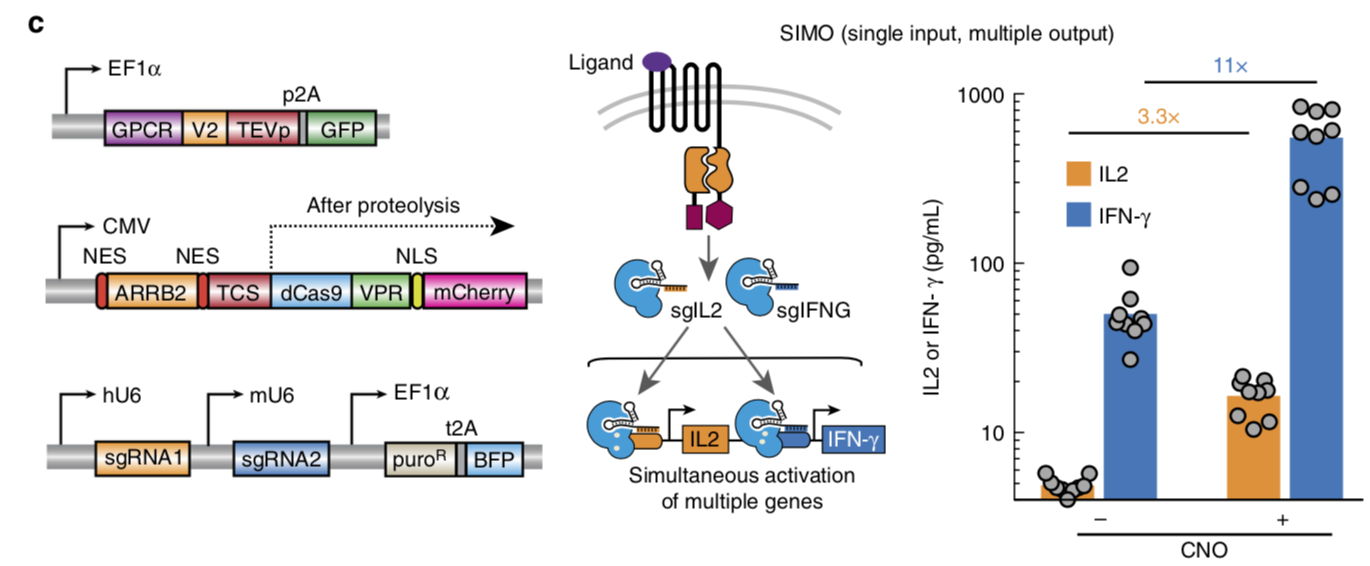
Note: Part:BBa_K2549013, Part:BBa_K2549014 and Part:BBa_K2549015 have the same Applications section.
References
- ↑ Tobacco etch virus protease: mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Kapust RB, Tözsér J, Fox JD, ..., Copeland TD, Waugh DS. Protein Eng, 2001 Dec;14(12):993-1000 PMID: 11809930
- ↑ https://en.wikipedia.org/wiki/TEV_protease
- ↑ The genetic design of signaling cascades to record receptor activation. Barnea G, Strapps W, Herrada G, ..., Axel R, Lee KJ. Proc Natl Acad Sci U S A, 2008 Jan;105(1):64-9 PMID: 18165312; DOI: 10.1073/pnas.0710487105
- ↑ Engineering cell sensing and responses using a GPCR-coupled CRISPR-Cas system. Kipniss NH, Dingal PCDP, Abbott TR, ..., Labanieh L, Qi LS. Nat Commun, 2017 Dec;8(1):2212 PMID: 29263378; DOI: 10.1038/s41467-017-02075-1
//function/degradation
| None |