Part:BBa_K2243012
Bxb1 gp35
Bxb1 gp35 integrase comes from Mycobacterium Phage Bxb1 and can bind to specific attB/P sites to catalyze DNA recombination. It helps the phage to integrate its genome into bacterial genome naturally. By constructing the attB/P sites in different directions, Bxb1 gp35 can catalyze the recombination of DNA between their sites, leading to inversion when attB/P are in opposite directions and excision when attB/P are in the same directions. Bxb1 gp35 is widely used to construct combinational logic gate and performs well in changing DNA sequence
Contents
Usages
Many researchers have paid attention to recombinases because of their ability of changing genetic circuits. Integrase Bxb1-gp35 is one of the recombinases with outstanding performance. In existence of recombinase Bxb1 gp35, different orientation of attB and attP allows the sequence to be flipped, excised, or inserted between recognition sites, which makes it useful for gene editing. In our project, we selected Bxb1-gp35 to flip the sequence flanked by attB and attP site for Bio-Flip-Flop construction.
Biology
Integrase Bxb1-gp35 comes from Mycobacteriophage, which allows the phage to insert its DNA into the host’s genome. We got the sequence by de novo synthesis.
Design Notes
We got the coding sequence by de novo synthesis and introduced three point mutations to remove the BsmB1 restriction enzyme cutting sites(in convenience for adding RBS by Golden Gate Assenmbly later).
Characterization
Since the viability of a bio-flip-flop relies on the performance of two integrases and their corresponding excisionases. To select integrases for the bio-flip-flop, we constructed expression vectors for different recombinases and tested their performance individually.
To make sure that Bxb1 have an optimal performance. We used the standard testing system, consisting of the integrase expression plasmid and the recombination reporter plasmid (BBa_K2243010). By choosing the vector with different replication origins(a p15A origin with a pTac promoter, and a ColE1 origin with a pBAD promoter) and the RBS sequences upon the integrase, we measure the recombination efficiency under different conditions.The expression vector and reporter of a recombinase were used to co-transform E. coli Top10 and samples were prepared for testing. We picked out our optimal RBS with low leakage and high efficiency for both backbone.

Figure 2. The standard testing system used to characterize the recombinases.

Fig 3. Integrase expression vectors with different replication origins. The one shown on top has a re-laxed ColE1 replication origin and recombinase expression is induced by arabinose via a pBad induc-tion system. The one shown in the bottom picture has a relaxed p15A replication origin and recom-binase expression is induced by IPTG.
Flow cytometry were used to evaluate the recombination efficiency, the expression vector and reporter of a recombinase were used to co-transform E. coli Top10 and samples were prepared for flow cytometry reading. Our procedures are as follows:
1.Single colonies were picked and used to inoculate into 1ml of LB media with antibiotics in a V-bottom 96-well plate.
2.The cultures were grown at 37°C and 1000 RPM for 12h. Subsequently, an aliquot comprising 2 μL of the culture was transferred into 1ml of M9 glucose media with antibiotics and inducer (1mM IPTG or 10mM arabinose for RBS tuning, gradient concentration for transfer curve) in a V-bottom 96-well plate.
3.The cultures were grown at 37°C and 1000 RPM for 15h. An aliquot comprising 2μL of the cul-ture was transferred into 198 μL of phosphate buffered saline (PBS) containing 2 mg/mL kanamycin in a 96-well plate. This mixture was incubated for one hour at room temperature before testing. Two lasers were used to excite GFP and RFP simultaneously. Single-cell fluorescence distribution at both emission wavelengths was recorded.
The counted cells were gated to eliminate the population which showed no fluorescence. The remaining cells were divided into two subsets by a diagonal: RFP sub-set and GFP subset. The recombination efficiency was estimated from the proportion of the RFP subset in the total fluorescent population.

Fig 5.A example of Gating the RFP and GFP subsets. Change of fluorescence after induction was seen. Left: no inducer. Right: 10 mM Ara for 15h.
For the vector with ColE1 replication origin, we found proper RBS in a list of RBSs for Bxb1 gp35.

Fig 3. Bxb1 gp35 recombination efficiency with a variety of calculated RBS and RBS sequences from iGEM (B0030~B0035). T.I.R = Translation Initiation Rate
Transfer curves
We evaluated the performance of Bxb1 gp35 on ColE1 expression vectors under a series of inducer concentration gradients. The results enabled us to determine the appropriate inducer concentration.

Fig 4. Bxb1 gp35 transfer curve with selected RBS, ColE1 replication origin, pBAD promoter

Fig 5. Bxb1 gp35 transfer curve with RBS B0032, ColE1 replication origin, pBAD promoter
Fusion Protein
Besides, we construct integrase-RDF fusion protein to recognize and bind attL and attR sequences,so the state transitions become reversible.click here to read more. https://parts.igem.org/Part:BBa_K2243013
Inversion Efficiency
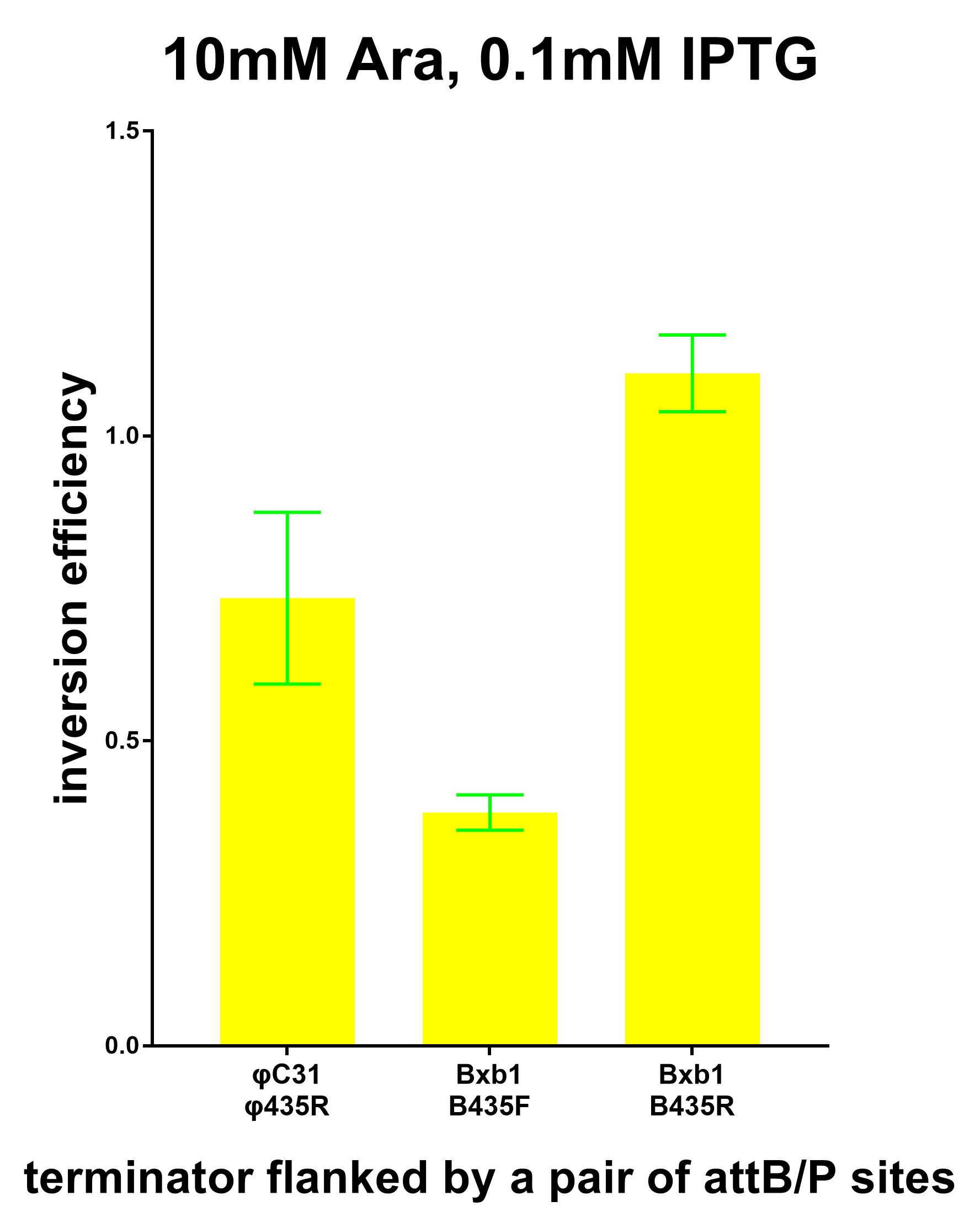
Furthermore, we co-transformed the system containing the terminator flanked by Bxb1 attB/P sites and the system expressing the Bxb1 recombinase to test the inversion efficiency using the plate reader. And the results are show in the graphs below.
Source
Mycobacterium Phage Bxb1
References
Roquet, N., Soleimany, A.P., Ferris, A.C., Aaronson, S. & Lu, T.K. Synthetic recombinase-based state machines in living cells. Science 353, aad8559 (2016).
This part was improved from Part:BBa_K907000.
https://parts.igem.org/Part:BBa_K907000
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 192
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 466
Illegal XhoI site found at 553 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 1105
Illegal NgoMIV site found at 1192
Illegal AgeI site found at 242 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 1300
| None |