Part:BBa_K1825006
35s CaMV + nptII
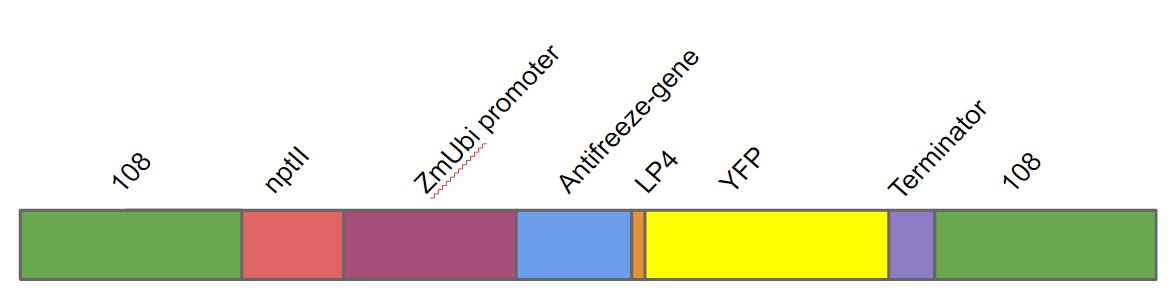
This is the neomycin phosphotransferase gene driven by the 35s cauliflower mosaic virus promoter. This promoter and gene was a part of a larger construct used for transforming Physcomitrella patens. We transformed this gene into Physcomitrella patens as a part of a larger DNA construct (fig. 1). This DNA construct contained in the following order. A homologous region to the 108 locus on the moss genome (to ensure stable integration into P. patens genome), the nptII-resistance gene driven by the 35s cauliflower mosaic virus, the Zea mays ubiquitin promoter driving the antifreeze protein (AFP) linked to yellow fluorescent protein (YFP) with the LP4 linker sequence, terminator and lastly another 108 region.
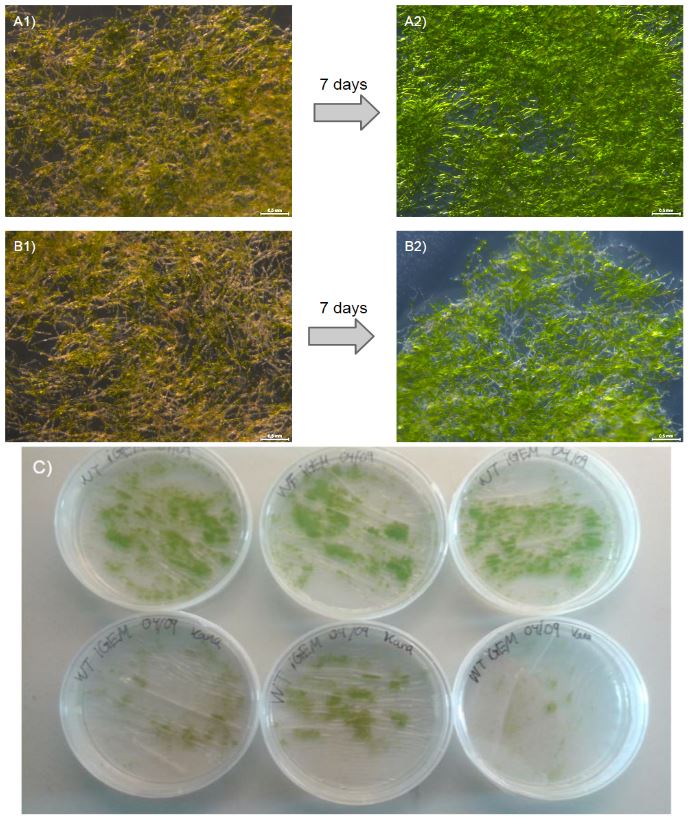
Our moss was able to grow from single transformed protoplasts to full clumps on media containing kanamycin, which suggests that the nptII-resistance cassette provides P.Patens with resistance to kanamycin (fig. 2).
To further validate this, we outlined an additional experiment where we grew wild type (WT) moss on either nonselective PhyB-media (3 plates) or PhyB-media containing kanamycin 50 mg/ml (3 plates). After 7 days, there was a visible difference in growth between WT on nonselective media and WT on kanamycin plates (fig. 3). The WT moss planted on kanamycin containing plates grew very little or not at all and is seen under the microscope as withering (fig. 3 B2). This is a stark contrast to our transformed moss that was able to grow from protoplasts to full clumps and the more vibrant WT without kanamycin (fig. 3 A2). This experiment validates the function of the nptII-cassette consisting of the neomycin phosphotransferase II gene driven by the 35s Cauliflower Mosaic virus promoter.

Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 1025
- 1000INCOMPATIBLE WITH RFC[1000]Illegal SapI.rc site found at 874
Illegal SapI.rc site found at 1084
//chassis/eukaryote/ppatens
| None |